diff --git a/DESCRIPTION b/DESCRIPTION
index f42fc33f..ad023168 100644
--- a/DESCRIPTION
+++ b/DESCRIPTION
@@ -1,14 +1,14 @@
Package: manynet
-Title: Many Ways to Make, Modify, Map, Mark, and Measure Myriad Networks
-Version: 1.5.1
-Date: 2025-06-23
-Description: Many tools for making, modifying, mapping, marking, measuring,
+Title: Many Ways to Make, Modify, Mark, and Measure Myriad Networks
+Version: 1.6.0
+Date: 2025-08-22
+Description: Many tools for making, modifying, marking, measuring,
and motifs and memberships of many different types of networks.
All functions operate with matrices, edge lists, and 'igraph', 'network', and 'tidygraph' objects,
- and on one-mode, two-mode (bipartite), and sometimes three-mode networks.
+ on directed, multiplex, multimodal, signed, and other networks.
The package includes functions for importing and exporting, creating and generating networks,
modifying networks and node and tie attributes,
- and describing and visualizing networks with sensible defaults.
+ and describing networks with sensible defaults.
URL: https://stocnet.github.io/manynet/
BugReports: https://github.com/stocnet/manynet/issues
License: MIT + file LICENSE
@@ -20,25 +20,16 @@ Depends: R (>= 3.6.0)
Imports:
cli,
dplyr (>= 1.1.0),
- ggplot2,
- ggraph,
igraph (>= 2.1.0),
network,
pillar,
tidygraph
Suggests:
- BiocManager,
- concaveman,
- gganimate,
- ggdendro,
- ggforce,
- gifski,
- graphlayouts,
+ autograph,
knitr,
learnr,
methods,
netdiffuseR,
- patchwork,
readxl,
rmarkdown,
RSiena,
@@ -47,8 +38,6 @@ Suggests:
tibble,
tidyr,
xml2
-Enhances:
- Rgraphviz
Authors@R:
c(person(given = "James",
family = "Hollway",
diff --git a/NAMESPACE b/NAMESPACE
index d70d1f4e..738e8b7c 100644
--- a/NAMESPACE
+++ b/NAMESPACE
@@ -1,6 +1,5 @@
# Generated by roxygen2: do not edit by hand
-S3method("+",ggplot)
S3method(add_nodes,igraph)
S3method(add_nodes,network)
S3method(add_nodes,tbl_graph)
@@ -140,6 +139,7 @@ S3method(net_dims,igraph)
S3method(net_dims,matrix)
S3method(net_dims,network)
S3method(pillar_shaft,logi)
+S3method(pillar_shaft,mdate)
S3method(print,diff_model)
S3method(print,learn_model)
S3method(print,mnet)
@@ -315,7 +315,6 @@ export(add_node_attribute)
export(add_nodes)
export(add_tie_attribute)
export(add_ties)
-export(aes)
export(apply_changes)
export(arrange_ties)
export(as_changelist)
@@ -329,9 +328,6 @@ export(as_network)
export(as_nodelist)
export(as_siena)
export(as_tidygraph)
-export(autographd)
-export(autographr)
-export(autographs)
export(bind_node_attributes)
export(bind_ties)
export(clear_glossary)
@@ -372,14 +368,7 @@ export(generate_random)
export(generate_scalefree)
export(generate_smallworld)
export(generate_utilities)
-export(ggplot)
-export(ggsave)
-export(ggtitle)
export(gloss)
-export(graphr)
-export(graphs)
-export(grapht)
-export(guides)
export(is.tbl_graph)
export(is_acyclic)
export(is_aperiodic)
@@ -408,22 +397,6 @@ export(join_ties)
export(k_elbow)
export(k_silhouette)
export(k_strict)
-export(labs)
-export(layout_tbl_graph_alluvial)
-export(layout_tbl_graph_concentric)
-export(layout_tbl_graph_configuration)
-export(layout_tbl_graph_dyad)
-export(layout_tbl_graph_hexad)
-export(layout_tbl_graph_hierarchy)
-export(layout_tbl_graph_ladder)
-export(layout_tbl_graph_lineage)
-export(layout_tbl_graph_multilevel)
-export(layout_tbl_graph_pentad)
-export(layout_tbl_graph_quad)
-export(layout_tbl_graph_railway)
-export(layout_tbl_graph_tetrad)
-export(layout_tbl_graph_triad)
-export(many_palettes)
export(mutate)
export(mutate_changes)
export(mutate_net)
@@ -632,6 +605,7 @@ export(node_is_pendant)
export(node_is_random)
export(node_is_recovered)
export(node_is_universal)
+export(node_kcoreness)
export(node_kernighanlin)
export(node_leading_eigen)
export(node_leiden)
@@ -685,42 +659,10 @@ export(rename)
export(rename_nodes)
export(rename_ties)
export(run_tute)
-export(scale_color_centres)
-export(scale_color_ethz)
-export(scale_color_iheid)
-export(scale_color_rug)
-export(scale_color_sdgs)
-export(scale_color_uzh)
-export(scale_colour_centres)
-export(scale_colour_ethz)
-export(scale_colour_iheid)
-export(scale_colour_rug)
-export(scale_colour_sdgs)
-export(scale_colour_uzh)
-export(scale_edge_color_centres)
-export(scale_edge_color_ethz)
-export(scale_edge_color_iheid)
-export(scale_edge_color_rug)
-export(scale_edge_color_sdgs)
-export(scale_edge_color_uzh)
-export(scale_edge_colour_centres)
-export(scale_edge_colour_ethz)
-export(scale_edge_colour_iheid)
-export(scale_edge_colour_rug)
-export(scale_edge_colour_sdgs)
-export(scale_edge_colour_uzh)
-export(scale_fill_centres)
-export(scale_fill_ethz)
-export(scale_fill_iheid)
-export(scale_fill_rug)
-export(scale_fill_sdgs)
-export(scale_fill_uzh)
-export(scale_size)
export(select)
export(select_changes)
export(select_nodes)
export(select_ties)
-export(set_manynet_theme)
export(snet_abort)
export(snet_info)
export(snet_minor_info)
@@ -734,10 +676,6 @@ export(snet_unavailable)
export(snet_warn)
export(summarise_ties)
export(table_data)
-export(theme_ethz)
-export(theme_iheid)
-export(theme_rug)
-export(theme_uzh)
export(tie_attribute)
export(tie_betweenness)
export(tie_closeness)
@@ -809,8 +747,6 @@ export(write_matrix)
export(write_nodelist)
export(write_pajek)
export(write_ucinet)
-export(xlab)
-export(ylab)
importFrom(cli,cli_div)
importFrom(cli,cli_end)
importFrom(cli,cli_inform)
@@ -825,39 +761,12 @@ importFrom(dplyr,distinct)
importFrom(dplyr,everything)
importFrom(dplyr,filter)
importFrom(dplyr,group_by)
-importFrom(dplyr,left_join)
importFrom(dplyr,mutate)
importFrom(dplyr,rename)
importFrom(dplyr,select)
importFrom(dplyr,summarise)
importFrom(dplyr,tibble)
importFrom(dplyr,ungroup)
-importFrom(ggplot2,aes)
-importFrom(ggplot2,arrow)
-importFrom(ggplot2,geom_point)
-importFrom(ggplot2,geom_segment)
-importFrom(ggplot2,geom_text)
-importFrom(ggplot2,ggplot)
-importFrom(ggplot2,ggsave)
-importFrom(ggplot2,ggtitle)
-importFrom(ggplot2,guides)
-importFrom(ggplot2,labs)
-importFrom(ggplot2,scale_alpha_manual)
-importFrom(ggplot2,scale_color_brewer)
-importFrom(ggplot2,scale_fill_brewer)
-importFrom(ggplot2,scale_size)
-importFrom(ggplot2,theme_void)
-importFrom(ggplot2,unit)
-importFrom(ggplot2,xlab)
-importFrom(ggplot2,ylab)
-importFrom(ggraph,create_layout)
-importFrom(ggraph,geom_conn_bundle)
-importFrom(ggraph,geom_edge_link)
-importFrom(ggraph,geom_node_label)
-importFrom(ggraph,geom_node_point)
-importFrom(ggraph,geom_node_text)
-importFrom(ggraph,get_con)
-importFrom(ggraph,scale_edge_width_continuous)
importFrom(igraph,E)
importFrom(igraph,V)
importFrom(igraph,add_edges)
@@ -894,12 +803,10 @@ importFrom(igraph,edge_density)
importFrom(igraph,eulerian_path)
importFrom(igraph,feedback_arc_set)
importFrom(igraph,fit_power_law)
-importFrom(igraph,get.edgelist)
importFrom(igraph,graph_from_adjacency_matrix)
importFrom(igraph,graph_from_biadjacency_matrix)
importFrom(igraph,graph_from_data_frame)
importFrom(igraph,graph_from_incidence_matrix)
-importFrom(igraph,gsize)
importFrom(igraph,has_eulerian_path)
importFrom(igraph,induced_subgraph)
importFrom(igraph,is_bipartite)
diff --git a/NEWS.md b/NEWS.md
index 8e51f320..347e9816 100644
--- a/NEWS.md
+++ b/NEWS.md
@@ -1,3 +1,45 @@
+# manynet 1.6.0
+
+## Package
+
+- Dropped viz-related dependencies: `{ggplot2}`, `{ggraph}`, `{patchwork}`,
+`{ggdendro}`, `{concaveman}`, `{gifski}`, `{graphlayouts}`, `{ggforce}`,
+and `{BiocManager}`
+
+## Classes
+
+- Added mdate pillar shaft for pretty printing of NAs in messydate variables
+
+## Modifying
+
+- Improved `to_no_missing()` to record missing removal
+- Improved `to_no_isolates()` to record isolate removal
+- Improved `to_giant()` to record giant component scoping
+- Improved `to_matching()` to record matching
+- Fixed `to_matching()` triggering warnings from handling NAs (thanks @schochastics for fixing #109)
+- Improved `to_mentoring()` to record mentoring
+- Improved `to_eulerian()` to record Eulerian pathing
+
+## Measuring
+
+- Improved `net_core()` with method options for calculating correlation, distance, ndiff, and diff
+
+## Membership
+
+- Improved `node_coreness()` to implement Borgatti and Everett's continuous coreness algorithm
+- Moved previous functionality of `node_coreness()` to `node_kcoreness()`
+
+## Mapping
+
+- Moved `graphr()`, `graphs()`, and `grapht()` to `{autograph}`
+- Moved palette functionality to `{autograph}`
+- Moved remaining layouts to `{autograph}`
+- Moved theme functionality to `{autograph}`
+- Dropped remaining plot functions
+- Dropped or avoided using `{autograph}`-related functions in examples to
+make package lighter
+- Please see `{autograph}` for continued development of these features
+
# manynet 1.5.1
## Package
diff --git a/R/make_generate.R b/R/make_generate.R
index 30925f9c..83eeb222 100644
--- a/R/make_generate.R
+++ b/R/make_generate.R
@@ -53,8 +53,8 @@ NULL
#' _Publicationes Mathematicae_. 6: 290–297.
#' @importFrom igraph sample_bipartite sample_gnp sample_gnm
#' @examples
-#' graphr(generate_random(12, 0.4))
-#' # graphr(generate_random(c(6, 6), 0.4))
+#' generate_random(12, 0.4)
+#' # generate_random(c(6, 6), 0.4)
#' @export
generate_random <- function(n, p = 0.5, directed = FALSE, with_attr = TRUE) {
if(is_manynet(n)){
@@ -243,8 +243,8 @@ NULL
#' \doi{10.1038/30918}.
#' @importFrom igraph sample_smallworld
#' @examples
-#' graphr(generate_smallworld(12, 0.025))
-#' graphr(generate_smallworld(12, 0.25))
+#' generate_smallworld(12, 0.025)
+#' generate_smallworld(12, 0.25)
#' @export
generate_smallworld <- function(n, p = 0.05, directed = FALSE, width = 2) {
directed <- infer_directed(n, directed)
@@ -270,8 +270,8 @@ generate_smallworld <- function(n, p = 0.05, directed = FALSE, width = 2) {
#' _Science_ 286(5439):509–12.
#' \doi{10.1126/science.286.5439.509}.
#' @examples
-#' graphr(generate_scalefree(12, 0.25))
-#' graphr(generate_scalefree(12, 1.25))
+#' generate_scalefree(12, 0.25)
+#' generate_scalefree(12, 1.25)
#' @export
generate_scalefree <- function(n, p = 1, directed = FALSE) {
directed <- infer_directed(n, directed)
diff --git a/R/make_play.R b/R/make_play.R
index 060d50da..431961c1 100644
--- a/R/make_play.R
+++ b/R/make_play.R
@@ -146,8 +146,6 @@ NULL
#' original network.
#' @examples
#' smeg <- generate_smallworld(15, 0.025)
-#' plot(play_diffusion(smeg, recovery = 0.4))
-#' #graphr(play_diffusion(ison_karateka))
#' @export
play_diffusion <- function(.data,
seeds = 1,
@@ -406,9 +404,6 @@ play_learning <- function(.data,
#' latticeEg <- add_node_attribute(latticeEg, "startValues", startValues)
#' latticeEg
#' play_segregation(latticeEg, "startValues", 0.5)
-#' # graphr(latticeEg, node_color = "startValues", node_size = 5) +
-#' # graphr(play_segregation(latticeEg, "startValues", 0.2),
-#' # node_color = "startValues", node_size = 5)
#' @export
play_segregation <- function(.data,
attribute,
diff --git a/R/make_read.R b/R/make_read.R
index ec60a1c3..9b8868c9 100644
--- a/R/make_read.R
+++ b/R/make_read.R
@@ -707,8 +707,6 @@ write_graphml <- function(.data,
#' @examples
#' # mnet <- read_cran()
#' # mnet <- to_ego(mnet, "manynet", max_dist = 2)
-#' # graphr(mnet, layout = "hierarchy",
-#' # edge_color = "type", node_color = "Compilation")
#' @export
read_cran <- function(pkg = "all"){
snet_progress_step("Downloading data about available packages from CRAN")
diff --git a/R/manip_correlation.R b/R/manip_correlation.R
index 6ae65023..1dd39551 100644
--- a/R/manip_correlation.R
+++ b/R/manip_correlation.R
@@ -74,9 +74,6 @@ NULL
#' @param with_attr Logical whether any attributes of the object
#' should be retained.
#' By default TRUE.
-#' @examples
-#' graphr(ison_adolescents, node_size = 4)
-#' graphr(to_permuted(ison_adolescents), node_size = 4)
#' @export
to_permuted <- function(.data, with_attr = TRUE) {
out <- as_matrix(.data)
diff --git a/R/manip_format.R b/R/manip_format.R
index 2cf76024..bf173600 100644
--- a/R/manip_format.R
+++ b/R/manip_format.R
@@ -378,7 +378,6 @@ to_acyclic.network <- function(.data) {
#' @importFrom igraph complementer
#' @examples
#' to_anti(ison_southern_women)
-#' #graphr(to_anti(ison_southern_women))
#' @export
to_anti <- function(.data) UseMethod("to_anti")
diff --git a/R/manip_reformed.R b/R/manip_reformed.R
index e3144dc8..b8fa8ebc 100644
--- a/R/manip_reformed.R
+++ b/R/manip_reformed.R
@@ -55,8 +55,6 @@ NULL
#' @examples
#' to_mode1(ison_southern_women)
#' to_mode2(ison_southern_women)
-#' #graphr(to_mode1(ison_southern_women))
-#' #graphr(to_mode2(ison_southern_women))
#' @export
to_mode1 <- function(.data, similarity = c("count","jaccard","rand","pearson","yule")) UseMethod("to_mode1")
@@ -161,7 +159,6 @@ to_mode2.data.frame <- function(.data, similarity = c("count","jaccard","rand","
#' @importFrom igraph make_line_graph E
#' @examples
#' to_ties(ison_adolescents)
-#' #graphr(to_ties(ison_adolescents))
#' @export
to_ties <- function(.data) UseMethod("to_ties")
@@ -252,7 +249,8 @@ to_no_missing <- function(.data) UseMethod("to_no_missing")
#' @export
to_no_missing.tbl_graph <- function(.data){
- delete_nodes(.data, !stats::complete.cases(as_nodelist(.data)))
+ delete_nodes(.data, !stats::complete.cases(as_nodelist(.data))) %>%
+ add_info(name = paste(net_name(.data), "without nodes with missing data"))
}
@@ -346,7 +344,8 @@ to_giant.network <- function(.data) {
#' @export
to_giant.tbl_graph <- function(.data) {
- as_tidygraph(to_giant(as_igraph(.data)))
+ as_tidygraph(to_giant(as_igraph(.data))) %>%
+ add_info(name = paste(net_name(.data, prefix = "Giant component of")))
}
#' @export
@@ -374,7 +373,9 @@ to_no_isolates <- function(.data) UseMethod("to_no_isolates")
to_no_isolates.tbl_graph <- function(.data) {
nodes <- NULL
# Delete edges not present vertices
- .data %>% tidygraph::activate(nodes) %>% dplyr::filter(!tidygraph::node_is_isolated())
+ .data %>% tidygraph::activate(nodes) %>%
+ dplyr::filter(!tidygraph::node_is_isolated()) %>%
+ add_info(name = paste(net_name(.data), "without isolates"))
}
#' @export
@@ -571,7 +572,7 @@ NULL
#'
#' Goldberg, Andrew V., and Robert E. Tarjan. 1986.
#' "A new approach to the maximum flow problem".
-#' _Proceedings of the eighteenth annual ACM symposium on Theory of computing – STOC '86_.
+#' _Proceedings of the 18th Annual ACM Symposium on Theory of Computing_.
#' 136-146.
#' \doi{10.1145/12130.12144}
#' @param mark A logical vector marking two types or modes.
@@ -582,9 +583,9 @@ NULL
#' @importFrom igraph max_bipartite_match
#' @examples
#' to_matching(ison_southern_women)
-#' #graphr(to_matching(ison_southern_women))
#' @export
-to_matching <- function(.data, mark = "type", capacities = NULL) UseMethod("to_matching")
+to_matching <- function(.data, mark = "type",
+ capacities = NULL) UseMethod("to_matching")
#' @export
to_matching.igraph <- function(.data, mark = "type", capacities = NULL){
@@ -594,8 +595,10 @@ to_matching.igraph <- function(.data, mark = "type", capacities = NULL){
el <- igraph::max_bipartite_match(.data,
types = node_attribute(.data, mark))$matching
el <- data.frame(from = names(el), to = el)
- out <- suppressWarnings(as_igraph(el, twomode = TRUE))
- out <- igraph::delete_vertices(out, "NA")
+ el$from[is.na(el$from)] <- "dummy"
+ el$to[is.na(el$to)] <- "dummy"
+ out <- as_igraph(el, twomode = TRUE)
+ out <- igraph::delete_vertices(out, "dummy")
out <- to_twomode(out, node_attribute(.data, mark))
} else {
if(length(capacities) == 1)
@@ -646,7 +649,8 @@ to_matching.igraph <- function(.data, mark = "type", capacities = NULL){
#' @export
to_matching.tbl_graph <- function(.data, mark = "type", capacities = NULL){
- as_tidygraph(to_matching.igraph(.data, mark, capacities = capacities))
+ as_tidygraph(to_matching.igraph(.data, mark, capacities = capacities)) %>%
+ add_info(name = paste(net_name(.data, prefix = "Stable matching of")))
}
#' @export
@@ -684,14 +688,13 @@ to_matching.matrix <- function(.data, mark = "type", capacities = NULL){
#' "Accelerating the Diffusion of Innovations Using Opinion Leaders",
#' _Annals of the American Academy of Political and Social Science_ 566: 56-67.
#' \doi{10.1177/000271629956600105}
-#' @examples
-#' graphr(to_mentoring(ison_adolescents))
#' @export
to_mentoring <- function(.data, elites = 0.1) UseMethod("to_mentoring")
#' @export
to_mentoring.tbl_graph <- function(.data, elites = 0.1){
- as_tidygraph(to_mentoring.igraph(.data, elites = elites))
+ as_tidygraph(to_mentoring.igraph(.data, elites = elites)) %>%
+ add_info(name = paste(net_name(.data), "mentorship"))
}
#' @export
@@ -744,7 +747,6 @@ to_mentoring.igraph <- function(.data, elites = 0.1){
#' \doi{10.1007/BF01442866}
#' @examples
#' to_eulerian(delete_nodes(ison_koenigsberg, "Lomse"))
-#' #graphr(to_eulerian(delete_nodes(ison_koenigsberg, "Lomse")))
#' @export
to_eulerian <- function(.data) UseMethod("to_eulerian")
@@ -765,7 +767,8 @@ to_eulerian.tbl_graph <- function(.data){
out <- paste(attr(igraph::eulerian_path(.data)$vpath, "names"),
collapse = "-+")
out <- create_explicit(out)
- out
+ out %>%
+ add_info(name = paste(net_name(.data, prefix = "Eulerian path of")))
}
#' @rdname manip_paths
diff --git a/R/manip_ties.R b/R/manip_ties.R
index 0201a1cd..0d1bf1b4 100644
--- a/R/manip_ties.R
+++ b/R/manip_ties.R
@@ -30,7 +30,7 @@ NULL
#' @param ties The number of ties to be added or an even list of ties.
#' @importFrom igraph add_edges
#' @examples
-#' ison_adolescents %>% add_ties(c("Betty","Tina")) %>% graphr()
+#' ison_adolescents %>% add_ties(c("Betty","Tina"))
#' @export
add_ties <- function(.data, ties, attribute = NULL) UseMethod("add_ties")
diff --git a/R/manynet-defunct.R b/R/manynet-defunct.R
index 7073c13a..da5fd5b2 100644
--- a/R/manynet-defunct.R
+++ b/R/manynet-defunct.R
@@ -27,37 +27,37 @@ node_mode <- function(.data) {
node_is_mode(.data)
}
-#' @describeIn defunct Deprecated on 2024-06-17.
-#' @export
-autographr <- function(.data, layout, labels = TRUE,
- node_color, node_shape, node_size, node_group,
- edge_color, edge_size, ...) {
- .Deprecated("graphr", package = "migraph",
- old = "autographr")
- graphr(.data, layout, labels,
- node_color, node_shape, node_size, node_group,
- edge_color, edge_size, ...)
-}
-
-#' @describeIn defunct Deprecated on 2024-06-17.
-#' @export
-autographs <- function(netlist, waves, based_on = c("first", "last", "both"), ...) {
- .Deprecated("graphs", package = "migraph",
- old = "autographs")
- graphs(netlist, waves, based_on, ...)
-}
-
-#' @describeIn defunct Deprecated on 2024-06-17.
-#' @export
-autographd <- function(tlist, layout, labels = TRUE,
- node_color, node_shape, node_size, edge_color, edge_size,
- keep_isolates = TRUE, ...) {
- .Deprecated("grapht", package = "migraph",
- old = "autographd")
- grapht(tlist, layout, labels,
- node_color, node_shape, node_size, edge_color, edge_size,
- keep_isolates, ...)
-}
+#' #' @describeIn defunct Deprecated on 2024-06-17.
+#' #' @export
+#' autographr <- function(.data, layout, labels = TRUE,
+#' node_color, node_shape, node_size, node_group,
+#' edge_color, edge_size, ...) {
+#' .Deprecated("graphr", package = "migraph",
+#' old = "autographr")
+#' graphr(.data, layout, labels,
+#' node_color, node_shape, node_size, node_group,
+#' edge_color, edge_size, ...)
+#' }
+
+#' #' @describeIn defunct Deprecated on 2024-06-17.
+#' #' @export
+#' autographs <- function(netlist, waves, based_on = c("first", "last", "both"), ...) {
+#' .Deprecated("graphs", package = "migraph",
+#' old = "autographs")
+#' graphs(netlist, waves, based_on, ...)
+#' }
+
+#' #' @describeIn defunct Deprecated on 2024-06-17.
+#' #' @export
+#' autographd <- function(tlist, layout, labels = TRUE,
+#' node_color, node_shape, node_size, edge_color, edge_size,
+#' keep_isolates = TRUE, ...) {
+#' .Deprecated("grapht", package = "migraph",
+#' old = "autographd")
+#' grapht(tlist, layout, labels,
+#' node_color, node_shape, node_size, edge_color, edge_size,
+#' keep_isolates, ...)
+#' }
#' @describeIn defunct Deprecated on 2024-06-14.
#' @export
@@ -727,13 +727,13 @@ net_by_quad <- function(.data) {
net_by_tetrad(.data)
}
-#' @describeIn defunct Deprecated on 2024-10-10.
-#' @export
-layout_tbl_graph_quad <- function(.data, circular = FALSE, times = 1000) {
- .Deprecated("layout_tbl_graph_tetrad", package = "manynet",
- old = "layout_tbl_graph_quad")
- layout_tbl_graph_tetrad(.data, circular = circular, times = times)
-}
+#' #' @describeIn defunct Deprecated on 2024-10-10.
+#' #' @export
+#' layout_tbl_graph_quad <- function(.data, circular = FALSE, times = 1000) {
+#' .Deprecated("layout_tbl_graph_tetrad", package = "manynet",
+#' old = "layout_tbl_graph_quad")
+#' layout_tbl_graph_tetrad(.data, circular = circular, times = times)
+#' }
#' @describeIn defunct Deprecated on 2024-12-30.
#' @export
diff --git a/R/manynet-tutorials.R b/R/manynet-tutorials.R
index 7b654f98..69bf030d 100644
--- a/R/manynet-tutorials.R
+++ b/R/manynet-tutorials.R
@@ -186,10 +186,14 @@ pillar_shaft.logi <- function(x, ...) {
pillar::style_na(x)), align = "left")
}
+#' @noRd
+#' @export
+pillar_shaft.mdate <- function(x, ...) {
+ pillar::pillar_shaft(as.character(x), width = 11)
+}
# Glossary ####
-
#' Adding network glossary items
#'
#' @description
diff --git a/R/manynet-utils.R b/R/manynet-utils.R
index 36aa8688..218669da 100644
--- a/R/manynet-utils.R
+++ b/R/manynet-utils.R
@@ -31,21 +31,6 @@ thisRequires <- function(pkgname){
}
}
-thisRequiresBio <- function(pkgname) {
- if (!requireNamespace(pkgname, quietly = TRUE) & interactive()) {
- if(utils::askYesNo(msg = paste("The", pkgname,
- "package is required to run this function. Would you like to install", pkgname, "from BioConductor?"))) {
- thisRequires("BiocManager")
- BiocManager::install(pkgname)
- }}
-}
-
-#' @export
-`+.ggplot` <- function(e1, e2, ...) {
- thisRequires("patchwork")
- patchwork::wrap_plots(e1, e2, ...)
-}
-
seq_nodes <- function(.data){
seq.int(net_nodes(.data))
}
diff --git a/R/map_autograph.R b/R/map_autograph.R
deleted file mode 100644
index 0d4c3ab2..00000000
--- a/R/map_autograph.R
+++ /dev/null
@@ -1,1114 +0,0 @@
-# Single graphs ####
-
-#' Easily graph networks with sensible defaults
-#'
-#' @description
-#' This function provides users with an easy way to graph
-#' (m)any network data for exploration, investigation, inspiration,
-#' and communication.
-#'
-#' It builds upon `{ggplot2}` and `{ggraph}` to offer
-#' pretty and extensible graphing solutions.
-#' However, compared to those solutions,
-#' `graphr()` contains various algorithms to provide better looking
-#' graphs by default.
-#' This means that just passing the function some network data
-#' will often be sufficient to return a reasonable-looking graph.
-#'
-#' The function also makes it easy to modify many of the most
-#' commonly adapted aspects of a graph,
-#' including node and edge size, colour, and shape,
-#' as arguments rather than additional functions that you need to remember.
-#' These can be defined outright, e.g. `node_size = 8`, or
-#' in reference to an attribute of the network, e.g. `node_size = "wealth"`.
-#'
-#' Lastly, `graphr()` uses `{ggplot2}`-related theme information, so
-#' it is easy to make colour palette and fonts institution-specific and consistent.
-#' See e.g. `theme_iheid()` for more.
-#'
-#' To learn more about what can be done visually,
-#' try `run_tute("Visualisation")`.
-#' @name map_graphr
-#' @family mapping
-#' @param .data A manynet-consistent object.
-#' @param layout An igraph, ggraph, or manynet layout algorithm.
-#' If not declared, defaults to "triad" for networks with 3 nodes,
-#' "quad" for networks with 4 nodes,
-#' "stress" for all other one mode networks,
-#' or "hierarchy" for two mode networks.
-#' For "hierarchy" layout, one can further split graph by
-#' declaring the "center" argument as the "events", "actors",
-#' or by declaring a node name.
-#' For "concentric" layout algorithm please declare the "membership" as an
-#' extra argument.
-#' The "membership" argument expects either a quoted node attribute present
-#' in data or vector with the same length as nodes to draw concentric circles.
-#' For "multilevel" layout algorithm please declare the "level"
-#' as extra argument.
-#' The "level" argument expects either a quoted node attribute present
-#' in data or vector with the same length as nodes to hierarchically
-#' order categories.
-#' If "level" is missing, function will look for 'lvl' node attribute in data.
-#' The "lineage" layout ranks nodes in Y axis according to values.
-#' For "lineage" layout algorithm please declare the "rank"
-#' as extra argument.
-#' The "rank" argument expects either a quoted node attribute present
-#' in data or vector with the same length as nodes.
-#' @param labels Logical, whether to print node names
-#' as labels if present.
-#' @param node_shape Node variable to be used for shaping the nodes.
-#' It is easiest if this is added as a node attribute to
-#' the graph before plotting.
-#' Nodes can also be shaped by declaring a shape instead.
-#' @param node_size Node variable to be used for sizing the nodes.
-#' This can be any continuous variable on the nodes of the network.
-#' Since this function expects this to be an existing variable,
-#' it is recommended to calculate all node-related statistics prior
-#' to using this function.
-#' Nodes can also be sized by declaring a numeric size or vector instead.
-#' @param node_color,node_colour Node variable to be used for coloring the nodes.
-#' It is easiest if this is added as a node attribute to
-#' the graph before plotting.
-#' Nodes can also be colored by declaring a color instead.
-#' @param node_group Node variable to be used for grouping the nodes.
-#' It is easiest if this is added as a hull over
-#' groups before plotting.
-#' Group variables should have a minimum of 3 nodes,
-#' if less, number groups will be reduced by
-#' merging categories with lower counts into one called "other".
-#' @param edge_color,edge_colour Tie variable to be used for coloring the nodes.
-#' It is easiest if this is added as an edge or tie attribute
-#' to the graph before plotting.
-#' Edges can also be colored by declaring a color instead.
-#' @param edge_size Tie variable to be used for sizing the edges.
-#' This can be any continuous variable on the nodes of the network.
-#' Since this function expects this to be an existing variable,
-#' it is recommended to calculate all edge-related statistics prior
-#' to using this function.
-#' Edges can also be sized by declaring a numeric size or vector instead.
-#' @param snap Logical scalar, whether the layout should be snapped to a grid.
-#' @param ... Extra arguments to pass on to the layout algorithm, if necessary.
-#' @return A `ggplot2::ggplot()` object.
-#' The last plot can be saved to the file system using `ggplot2::ggsave()`.
-#' @importFrom ggraph geom_edge_link geom_node_text geom_conn_bundle
-#' get_con geom_node_point scale_edge_width_continuous geom_node_label
-#' @importFrom ggplot2 aes arrow unit scale_color_brewer scale_fill_brewer
-#' @examples
-#' graphr(ison_adolescents)
-#' ison_adolescents %>%
-#' mutate(color = rep(c("introvert","extrovert"), times = 4),
-#' size = ifelse(node_is_cutpoint(ison_adolescents), 6, 3)) %>%
-#' mutate_ties(ecolor = rep(c("friends", "acquaintances"), times = 5)) %>%
-#' graphr(node_color = "color", node_size = "size",
-#' edge_size = 1.5, edge_color = "ecolor")
-#' @export
-graphr <- function(.data, layout, labels = TRUE,
- node_color, node_shape, node_size, node_group,
- edge_color, edge_size, snap = FALSE, ...,
- node_colour, edge_colour) {
- g <- as_tidygraph(.data)
- if (missing(layout)) {
- if (net_nodes(g) <= 6) {
- layout <- "configuration"
- } else if (is_twomode(g)) {
- layout <- "hierarchy"
- } else layout <- "stress"
- }
- if (missing(node_color) && missing(node_colour)) {
- node_color <- NULL
- } else if (missing(node_color)) {
- node_color <- as.character(substitute(node_colour))
- } else {
- node_color <- as.character(substitute(node_color))
- }
- if (missing(node_shape)) node_shape <- NULL else
- node_shape <- as.character(substitute(node_shape))
- if (missing(node_size)) node_size <- NULL else if (!is.numeric(node_size)) {
- node_size <- as.character(substitute(node_size))
- }
- if (missing(node_group)) node_group <- NULL else {
- node_group <- as.character(substitute(node_group))
- g <- tidygraph::activate(g, "nodes") %>%
- tidygraph::mutate(node_group = reduce_categories(g, node_group))
- }
- if (missing(edge_color) && missing(edge_colour)) {
- edge_color <- NULL
- } else if (missing(edge_color)) {
- edge_color <- as.character(substitute(edge_colour))
- } else {
- edge_color <- as.character(substitute(edge_color))
- }
- if (missing(edge_size)) edge_size <- NULL else if (!is.numeric(edge_size)) {
- edge_size <- as.character(substitute(edge_size))
- }
- # Add layout ----
- p <- .graph_layout(g, layout, labels, node_group, snap, ...)
- # Add background ----
- if(getOption("mnet_background", default = "#FFFFFF")!="#FFFFFF")
- p <- p + ggplot2::theme(panel.background = ggplot2::element_rect(fill = getOption("mnet_background", default = "#FFFFFF")))
- # Add edges ----
- p <- .graph_edges(p, g, edge_color, edge_size, node_size)
- # Add nodes ----
- p <- .graph_nodes(p, g, node_color, node_shape, node_size)
- # Add labels ----
- if (isTRUE(labels) & is_labelled(g)) {
- p <- .graph_labels(p, g, layout)
- }
- # assign("last.warning", NULL, envir = baseenv()) # to avoid persistent ggrepel
- p
-}
-
-.graph_layout <- function(g, layout, labels, node_group, snap, ...) {
- name <- NULL
- dots <- list(...)
- if ("x" %in% names(dots) & "y" %in% names(dots)) {
- lo <- ggraph::create_layout(g, layout = "manual",
- x = dots[["x"]], y = dots[["y"]])
- } else lo <- suppressWarnings(ggraph::create_layout(g, layout, ...))
- if ("graph" %in% names(attributes(lo))) {
- if (!setequal(names(as.data.frame(attr(lo, "graph"))), names(lo))) {
- for (n in setdiff(names(as.data.frame(attr(lo, "graph"))), names(lo))) {
- lo[n] <- igraph::vertex_attr(g, n)
- }
- }
- }
- p <- ggraph::ggraph(lo) + ggplot2::theme_void()
- if (!is.null(node_group)) {
- x <- y <- NULL
- thisRequires("ggforce")
- thisRequires("concaveman")
- p <- p +
- ggforce::geom_mark_hull(ggplot2::aes(x, y, fill = node_group,
- label = node_group), data = lo) +
- ggplot2::scale_fill_manual(values = colorsafe_palette,
- guide = ggplot2::guide_legend("Group"))
- }
- if(snap){
- snet_info("Snapping layout coordinates to grid.")
- if(grepl("lattice",
- igraph::graph_attr(attr(p$data, "graph"), "grand")$name,
- ignore.case = TRUE))
- p$data[,c("x","y")] <- round(p$data[,c("x","y")])
- else p$data[,c("x","y")] <- depth_first_recursive_search(p)
- }
- p
-}
-
-.graph_edges <- function(p, g, edge_color, edge_size, node_size) {
- if (is_directed(g)) {
- out <- .infer_directed_edge_mapping(g, edge_color, edge_size, node_size)
- p <- map_directed_edges(p, g, out)
- } else {
- out <- .infer_edge_mapping(g, edge_color, edge_size)
- p <- map_edges(p, g, out)
- }
- if (is_complex(g)) {
- p <- p + ggraph::geom_edge_loop0(edge_alpha = 0.4)
- }
- # Check legends
- if (length(unique(out[["esize"]])) == 1) {
- p <- p + ggplot2::guides(edge_width = "none")
- } else p <- p + ggraph::scale_edge_width_continuous(range = c(0.3, 3),
- guide = ggplot2::guide_legend(
- ifelse(is.null(edge_size) &
- is_weighted(g),
- "Edge Weight", "Edge Size")))
- if (length(unique(out[["ecolor"]])) == 1) {
- p <- p + ggplot2::guides(edge_colour = "none")
- } else if (length(unique(out[["ecolor"]])) == 2){
- p <- p + ggraph::scale_edge_colour_manual(values = getOption("mnet_highlight", default = c("grey","black")),
- guide = ggplot2::guide_legend(
- ifelse(is.null(edge_color) &
- is_signed(g),
- "Edge Sign", "Edge Color")))
- } else p <- p + ggraph::scale_edge_colour_manual(values = getOption("mnet_cat", default = colorsafe_palette),
- guide = ggplot2::guide_legend(
- ifelse(is.null(edge_color) &
- is_signed(g),
- "Edge Sign", "Edge Color")))
- p
-}
-
-.graph_nodes <- function(p, g, node_color, node_shape, node_size) {
- out <- .infer_node_mapping(g, node_color, node_size, node_shape)
- if (is.null(node_color) & "Infected" %in% names(node_attribute(g))) {
- p <- map_infected_nodes(p, g, out)
- } else if (is.null(node_color) & any("diff_model" %in% names(attributes(g)))) {
- p <- map_diff_model_nodes(p, g, out)
- } else {
- p <- map_nodes(p, out)
- # Check legends
- if (length(unique(out[["nsize"]])) > 1)
- p <- p + ggplot2::guides(size = ggplot2::guide_legend(title = "Node Size"))
- if (length(unique(out[["nshape"]])) > 1)
- p <- p + ggplot2::guides(shape = ggplot2::guide_legend(
- title = ifelse(is_twomode(g) & is.null(node_shape), "Node Mode", "Node Shape")))
- if (length(unique(out[["ncolor"]])) > 1){
- if(length(unique(out[["ncolor"]])) == 2){
- p <- p + ggplot2::scale_colour_manual(values = getOption("mnet_highlight", default = c("grey","black")),
- guide = ggplot2::guide_legend("Node Color"))
- } else {
- p <- p + ggplot2::scale_colour_manual(values = getOption("mnet_cat", default = colorsafe_palette),
- guide = ggplot2::guide_legend("Node Color"))
- }
- }
- }
- # Consider rescaling nodes
- p <- p + ggplot2::scale_size(range = c(1/net_nodes(g)*50, 1/net_nodes(g)*100))
- p
-}
-
-.graph_labels <- function(p, g, layout) {
- if (layout == "circle" | layout == "concentric") {
- angles <- as.data.frame(cart2pol(as.matrix(p[["data"]][,1:2])))
- angles$degree <- angles$phi * 180/pi
- angles <- dplyr::case_when(p[["data"]][,2] == 0 & p[["data"]][,1] == 0 ~ 0.1,
- p[["data"]][,2] >= 0 & p[["data"]][,1] > 0 ~ angles$degree,
- p[["data"]][,2] < 0 & p[["data"]][,1] > 0 ~ angles$degree,
- p[["data"]][,1] == 1 ~ angles$degree,
- TRUE ~ angles$degree - 180)
- if (net_nodes(g) < 10) {
- hj <- ifelse(p[["data"]][,1] >= 0, -0.8, 1.8)
- } else if (net_nodes(g) < 20) {
- hj <- ifelse(p[["data"]][,1] >= 0, -0.4, 1.4)
- } else {
- hj <- ifelse(p[["data"]][,1] >= 0, -0.2, 1.2)
- }
- p <- p + ggraph::geom_node_text(ggplot2::aes(label = name), repel = TRUE,
- size = 3, hjust = hj, angle = angles) +
- ggplot2::coord_cartesian(xlim=c(-1.2,1.2), ylim=c(-1.2,1.2))
- } else if (layout %in% c("bipartite", "railway") | layout == "hierarchy" &
- length(unique(p[["data"]][["y"]])) <= 2) {
- p <- p + ggraph::geom_node_text(ggplot2::aes(label = name), angle = 90,
- size = 3, hjust = "outward", repel = TRUE,
- nudge_y = ifelse(p[["data"]][,2] == 1,
- 0.05, -0.05)) +
- ggplot2::coord_cartesian(ylim=c(-0.2, 1.2))
- } else if (layout == "hierarchy" & length(unique(p[["data"]][["y"]])) > 2) {
- p <- p + ggraph::geom_node_text(ggplot2::aes(label = name),
- size = 3, hjust = "inward", repel = TRUE)
- } else if (layout %in% c("alluvial", "lineage")) {
- p <- p + ggraph::geom_node_label(ggplot2::aes(label = name), size = 3,
- repel = TRUE, nudge_x = ifelse(p[["data"]][,1] == 1,
- 0.02, -0.02))
- } else {
- p <- p + ggraph::geom_node_label(ggplot2::aes(label = name),
- repel = TRUE, seed = 1234, size = 3)
- }
-}
-
-# `graphr()` helper functions
-reduce_categories <- function(g, node_group) {
- limit <- toCondense <- NULL
- if (sum(table(node_attribute(g, node_group)) <= 2) > 2 &
- length(unique(node_attribute(g, node_group))) > 2) {
- toCondense <- names(which(table(node_attribute(g, node_group)) <= 2))
- out <- ifelse(node_attribute(g, node_group) %in% toCondense,
- "Other", node_attribute(g, node_group))
- snet_info("The number of groups was reduced since there were groups with less than 2 nodes.")
- } else if (sum(table(node_attribute(g, node_group)) <= 2) == 2 &
- length(unique(node_attribute(g, node_group))) > 2) {
- limit <- stats::reorder(node_attribute(g, node_group),
- node_attribute(g, node_group),
- FUN = length, decreasing = TRUE)
- if (sum(utils::tail(attr(limit, "scores"), 2))) {
- toCondense <- utils::tail(levels(limit), 3)
- } else {
- toCondense <- utils::tail(levels(limit), 2)
- }
- out <- ifelse(node_attribute(g, node_group) %in% toCondense, "Other",
- node_attribute(g, node_group))
- snet_info("The number of groups was reduced since there were groups with less than 2 nodes.")
- } else if (sum(table(node_attribute(g, node_group)) <= 2) == 1 &
- length(unique(node_attribute(g, node_group))) > 2) {
- limit <- stats::reorder(node_attribute(g, node_group),
- node_attribute(g, node_group),
- FUN = length, decreasing = TRUE)
- toCondense <- utils::tail(levels(limit), 2)
- out <- ifelse(node_attribute(g, node_group) %in% toCondense, "Other",
- node_attribute(g, node_group))
- snet_info("The number of groups was reduced since there were groups with less than 2 nodes.")
- } else if (sum(table(node_attribute(g, node_group)) <= 2) == 1 &
- length(unique(node_attribute(g, node_group))) == 2) {
- out <- as.factor(node_attribute(g, node_group))
- snet_info("Node groups with 2 nodes or less can be cause issues for plotting ...")
- } else out <- as.factor(node_attribute(g, node_group))
- out
-}
-
-.infer_directed_edge_mapping <- function(g, edge_color, edge_size, node_size) {
- check_edge_variables(g, edge_color, edge_size)
- list("ecolor" = .infer_ecolor(g, edge_color),
- "esize" = .infer_esize(g, edge_size),
- "line_type" = .infer_line_type(g),
- "end_cap" = .infer_end_cap(g, node_size))
-}
-
-.infer_edge_mapping <- function(g, edge_color, edge_size) {
- check_edge_variables(g, edge_color, edge_size)
- list("ecolor" = .infer_ecolor(g, edge_color),
- "esize" = .infer_esize(g, edge_size),
- "line_type" = .infer_line_type(g))
-}
-
-.infer_ecolor <- function(g, edge_color){
- if (!is.null(edge_color)) {
- if (edge_color %in% names(tie_attribute(g))) {
- if ("tie_mark" %in% class(tie_attribute(g, edge_color))) {
- out <- factor(as.character(tie_attribute(g, edge_color)),
- levels = c("FALSE", "TRUE"))
- } else out <- as.factor(as.character(tie_attribute(g, edge_color)))
- if (length(unique(out)) == 1) {
- out <- rep("black", net_ties(g))
- snet_info("Please indicate a variable with more than one value or level when mapping edge colors.")
- }
- } else {
- out <- edge_color
- }
- } else if (is.null(edge_color) & is_signed(g)) {
- out <- factor(ifelse(igraph::E(g)$sign >= 0, "Positive", "Negative"),
- levels = c("Positive", "Negative"))
- if (length(unique(out)) == 1) {
- out <- "black"
- }
- } else {
- out <- "black"
- }
- out
-}
-
-.infer_esize <- function(g, edge_size){
- if (!is.null(edge_size)) {
- if (any(edge_size %in% names(tie_attribute(g)))) {
- out <- tie_attribute(g, edge_size)
- } else {
- out <- edge_size
- }
- } else if (is.null(edge_size) & is_weighted(g)) {
- out <- tie_attribute(g, "weight")
- } else {
- out <- 0.5
- }
- out
-}
-
-.infer_end_cap <- function(g, node_size) {
- nsize <- .infer_nsize(g, node_size)/2
- # Accounts for rescaling
- if (length(unique(nsize)) == 1) {
- out <- rep(unique(nsize), net_ties(g))
- } else {
- out <- g %>%
- tidygraph::activate("edges") %>%
- data.frame() %>%
- dplyr::left_join(data.frame(node_id = 1:length(node_names(g)),
- nsize = nsize),
- by = c("to" = "node_id"))
- out <- out$nsize
- out <- ((out - min(out)) / (max(out) - min(out))) *
- ((1 / net_nodes(g) * 100) - (1 / net_nodes(g)*50)) + (1 / net_nodes(g) * 50)
- }
- out
-}
-
-.infer_line_type <- function(g) {
- if (is_signed(g)) {
- out <- ifelse(as.numeric(tie_signs(g)) >= 0,
- "solid", "dashed")
- # ifelse(length(unique(out)) == 1, unique(out), out)
- } else out <- "solid"
- out
-}
-
-check_edge_variables <- function(g, edge_color, edge_size) {
- if (!is.null(edge_color)) {
- if (any(!tolower(edge_color) %in% tolower(igraph::edge_attr_names(g))) &
- any(!edge_color %in% grDevices::colors())) {
- snet_info("Please make sure you spelled `edge_color` variable correctly.")
- }
- }
- if (!is.null(edge_size)) {
- if (!is.numeric(edge_size) & any(!tolower(edge_size) %in% tolower(igraph::edge_attr_names(g)))) {
- snet_info("Please make sure you spelled `edge_size` variable correctly.")
- }
- }
-}
-
-map_directed_edges <- function(p, g, out) {
- if (length(out[["ecolor"]]) == 1 & length(out[["esize"]]) == 1) {
- p <- p + ggraph::geom_edge_arc(ggplot2::aes(end_cap = ggraph::circle(c(out[["end_cap"]]), 'mm')),
- edge_colour = out[["ecolor"]], edge_width = out[["esize"]],
- edge_linetype = out[["line_type"]],
- edge_alpha = 0.4, strength = ifelse(igraph::which_mutual(g), 0.2, 0),
- arrow = ggplot2::arrow(angle = 15, type = "closed",
- length = ggplot2::unit(2, 'mm')))
- } else if (length(out[["ecolor"]]) > 1 & length(out[["esize"]]) == 1) {
- p <- p + ggraph::geom_edge_arc(ggplot2::aes(edge_colour = out[["ecolor"]],
- end_cap = ggraph::circle(c(out[["end_cap"]]), 'mm')),
- edge_width = out[["esize"]], edge_linetype = out[["line_type"]],
- edge_alpha = 0.4, strength = ifelse(igraph::which_mutual(g), 0.2, 0),
- arrow = ggplot2::arrow(angle = 15, type = "closed",
- length = ggplot2::unit(2, 'mm')))
- } else if (length(out[["ecolor"]]) == 1 & length(out[["esize"]]) > 1) {
- p <- p + ggraph::geom_edge_arc(ggplot2::aes(edge_width = out[["esize"]],
- end_cap = ggraph::circle(c(out[["end_cap"]]), 'mm')),
- edge_colour = out[["ecolor"]], edge_linetype = out[["line_type"]],
- edge_alpha = 0.4, strength = ifelse(igraph::which_mutual(g), 0.2, 0),
- arrow = ggplot2::arrow(angle = 15, type = "closed",
- length = ggplot2::unit(2, 'mm')))
- } else {
- p <- p + ggraph::geom_edge_arc(ggplot2::aes(edge_colour = getOption("mnet_cat")[out[["ecolor"]]],
- edge_width = out[["esize"]],
- end_cap = ggraph::circle(c(out[["end_cap"]]), 'mm')),
- edge_linetype = out[["line_type"]],
- edge_alpha = 0.4, strength = ifelse(igraph::which_mutual(g), 0.2, 0),
- arrow = ggplot2::arrow(angle = 15, type = "closed",
- length = ggplot2::unit(2, 'mm')))
- }
- p
-}
-
-map_edges <- function(p, g, out) {
- if (length(out[["ecolor"]]) == 1 & length(out[["esize"]]) == 1) {
- p <- p + ggraph::geom_edge_link0(edge_width = out[["esize"]],
- edge_colour = out[["ecolor"]],
- edge_alpha = 0.4,
- edge_linetype = out[["line_type"]])
- } else if (length(out[["ecolor"]]) > 1 & length(out[["esize"]]) == 1) {
- p <- p + ggraph::geom_edge_link0(ggplot2::aes(edge_colour = out[["ecolor"]]),
- edge_width = out[["esize"]],
- edge_alpha = 0.4,
- edge_linetype = out[["line_type"]])
- } else if (length(out[["ecolor"]]) == 1 & length(out[["esize"]]) > 1) {
- p <- p + ggraph::geom_edge_link0(ggplot2::aes(edge_width = out[["esize"]]),
- edge_colour = out[["ecolor"]],
- edge_alpha = 0.4,
- edge_linetype = out[["line_type"]])
- } else {
- p <- p + ggraph::geom_edge_link0(ggplot2::aes(edge_width = out[["esize"]],
- edge_colour = out[["ecolor"]]),
- edge_alpha = 0.4, edge_linetype = out[["line_type"]])
- }
-}
-
-.infer_node_mapping <- function(g, node_color, node_size, node_shape) {
- check_node_variables(g, node_color, node_size)
- list("nshape" = .infer_nshape(g, node_shape),
- "nsize" = .infer_nsize(g, node_size),
- "ncolor" = .infer_ncolor(g, node_color))
-}
-
-.infer_nsize <- function(g, node_size) {
- if (!is.null(node_size)) {
- if (is.character(node_size)) {
- out <- node_attribute(g, node_size)
- } else out <- node_size
- if (length(node_size > 1) & all(out <= 1 & out >= 0)) out <- out * 10
- } else {
- out <- min(20, (250 / net_nodes(g)) / 2)
- }
- as.numeric(out)
-}
-
-.infer_nshape <- function(g, node_shape) {
- if (!is.null(node_shape)) {
- if (node_shape %in% names(node_attribute(g))) {
- out <- as.factor(as.character(node_attribute(g, node_shape)))
- } else out <- node_shape
- } else if (is_twomode(g) & is.null(node_shape)) {
- out <- ifelse(igraph::V(g)$type, "One", "Two")
- } else {
- out <- "circle"
- }
- out
-}
-
-.infer_ncolor <- function(g, node_color) {
- if (!is.null(node_color)) {
- if (node_color %in% names(node_attribute(g))) {
- if ("node_mark" %in% class(node_attribute(g, node_color))) {
- out <- factor(as.character(node_attribute(g, node_color)),
- levels = c("FALSE", "TRUE"))
- } else out <- as.factor(as.character(node_attribute(g, node_color)))
- if (length(unique(out)) == 1) {
- out <- rep("black", net_nodes(g))
- snet_info("Please indicate a variable with more than one value or level when mapping node colors.")
- }
- } else out <- node_color
- } else {
- out <- "black"
- }
- out
-}
-
-check_node_variables <- function(g, node_color, node_size) {
- if (!is.null(node_color)) {
- if (any(!tolower(node_color) %in% tolower(igraph::vertex_attr_names(g))) &
- any(!node_color %in% grDevices::colors())) {
- snet_info("Please make sure you spelled `node_color` variable correctly.")
- }
- }
- if (!is.null(node_size)) {
- if (!is.numeric(node_size) & any(!tolower(node_size) %in% tolower(igraph::vertex_attr_names(g)))) {
- snet_info("Please make sure you spelled `node_size` variable correctly.")
- }
- }
-}
-
-map_infected_nodes<- function(p, g, out) {
- node_color <- as.factor(ifelse(node_attribute(g, "Exposed"), "Exposed",
- ifelse(node_attribute(g, "Infected"),"Infected",
- ifelse(node_attribute(g, "Recovered"), "Recovered",
- "Susceptible"))))
- p + ggraph::geom_node_point(ggplot2::aes(color = node_color),
- size = out[["nsize"]], shape = out[["nshape"]]) +
- ggplot2::scale_color_manual(name = NULL, guide = ggplot2::guide_legend(""),
- values = c("Infected" = "#d73027",
- "Susceptible" = "#4575b4",
- "Exposed" = "#E6AB02",
- "Recovered" = "#66A61E"))
-}
-
-map_diff_model_nodes <- function(p, g, out) {
- node_adopts <- node_adoption_time(attr(g,"diff_model"))
- nshape <- ifelse(node_adopts == min(node_adopts), "Seed(s)",
- ifelse(node_adopts == Inf, "Non-Adopter", "Adopter"))
- node_color <- ifelse(is.infinite(node_adopts),
- max(node_adopts[!is.infinite(node_adopts)]) + 1,
- node_adopts)
- p + ggraph::geom_node_point(ggplot2::aes(shape = nshape, color = node_color),
- size = out[["nsize"]]) +
- ggplot2::scale_color_gradient(low = "#d73027", high = "#4575b4",
- breaks=c(min(node_color)+1,
- ifelse(any(nshape=="Non-Adopter"),
- max(node_color)-1,
- max(node_color))),
- labels=c("Early\nadoption", "Late\nadoption"),
- name = "Time of\nAdoption\n") +
- ggplot2::scale_shape_manual(name = "",
- breaks = c("Seed(s)", "Adopter", "Non-Adopter"),
- values = c("Seed(s)" = "triangle",
- "Adopter" = "circle",
- "Non-Adopter" = "square")) +
- ggplot2::guides(color = ggplot2::guide_colorbar(order = 1, reverse = TRUE),
- shape = ggplot2::guide_legend(order = 2))
-}
-
-map_nodes <- function(p, out) {
- if (length(out[["ncolor"]]) == 1 & length(out[["nsize"]]) == 1 & length(out[["nshape"]]) == 1) {
- p <- p + ggraph::geom_node_point(colour = out[["ncolor"]], size = out[["nsize"]],
- shape = out[["nshape"]])
- } else if (length(out[["ncolor"]]) > 1 & length(out[["nsize"]]) == 1 & length(out[["nshape"]]) == 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(colour = out[["ncolor"]]),
- size = out[["nsize"]], shape = out[["nshape"]])
- } else if (length(out[["ncolor"]]) == 1 & length(out[["nsize"]]) > 1 & length(out[["nshape"]]) == 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(size = out[["nsize"]]),
- colour = out[["ncolor"]], shape = out[["nshape"]])
- } else if (length(out[["ncolor"]]) == 1 & length(out[["nsize"]]) == 1 & length(out[["nshape"]]) > 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(shape = out[["nshape"]]),
- colour = out[["ncolor"]], size = out[["nsize"]])
- } else if (length(out[["ncolor"]]) > 1 & length(out[["nsize"]]) > 1 & length(out[["nshape"]]) == 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(colour = out[["ncolor"]], size = out[["nsize"]]),
- shape = out[["nshape"]])
- } else if (length(out[["ncolor"]]) > 1 & length(out[["nsize"]]) == 1 & length(out[["nshape"]]) > 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(colour = out[["ncolor"]], shape = out[["nshape"]]),
- size = out[["nsize"]])
- } else if (length(out[["ncolor"]]) == 1 & length(out[["nsize"]]) > 1 & length(out[["nshape"]]) > 1) {
- p <- p + ggraph::geom_node_point(ggplot2::aes(size = out[["nsize"]], shape = out[["nshape"]]),
- colour = out[["ncolor"]])
- } else {
- p <- p + ggraph::geom_node_point(ggplot2::aes(colour = out[["ncolor"]],
- shape = out[["nshape"]],
- size = out[["nsize"]]))
- }
- p
-}
-
-
-cart2pol <- function(xyz){
- stopifnot(is.numeric(xyz))
- if (is.vector(xyz) && (length(xyz) == 2 || length(xyz) ==
- 3)) {
- x <- xyz[1]
- y <- xyz[2]
- m <- 1
- n <- length(xyz)
- }
- else if (is.matrix(xyz) && (ncol(xyz) == 2 || ncol(xyz) ==
- 3)) {
- x <- xyz[, 1]
- y <- xyz[, 2]
- m <- nrow(xyz)
- n <- ncol(xyz)
- }
- else snet_abort("Input must be a vector of length 3 or a matrix with 3 columns.")
- phi <- atan2(y, x)
- r <- hypot(x, y)
- if (n == 2) {
- if (m == 1)
- prz <- c(phi, r)
- else prz <- cbind(phi, r)
- }
- else {
- if (m == 1) {
- z <- xyz[3]
- prz <- c(phi, r, z)
- }
- else {
- z <- xyz[, 3]
- prz <- cbind(phi, r, z)
- }
- }
- return(prz)
-}
-
-hypot <- function (x, y) {
- if ((length(x) == 0 && is.numeric(y) && length(y) <= 1) ||
- (length(y) == 0 && is.numeric(x) && length(x) <= 1))
- return(vector())
- if (!is.numeric(x) && !is.complex(x) || !is.numeric(y) &&
- !is.complex(y))
- snet_abort("Arguments 'x' and 'y' must be numeric or complex.")
- if (length(x) == 1 && length(y) > 1) {
- x <- rep(x, length(y))
- dim(x) <- dim(y)
- }
- else if (length(x) > 1 && length(y) == 1) {
- y <- rep(y, length(x))
- dim(y) <- dim(x)
- }
- if ((is.vector(x) && is.vector(y) && length(x) != length(y)) ||
- (is.matrix(x) && is.matrix(y) && dim(x) != dim(y)) ||
- (is.vector(x) && is.matrix(y)) || is.matrix(x) && is.vector(y))
- snet_abort("Arguments 'x' and 'y' must be of the same size.")
- x <- abs(x)
- y <- abs(y)
- m <- pmin(x, y)
- M <- pmax(x, y)
- ifelse(M == 0, 0, M * sqrt(1 + (m/M)^2))
-}
-
-# Longitudinal or comparative networks ####
-
-#' Easily graph a set of networks with sensible defaults
-#'
-#' @description
-#' This function provides users with an easy way to graph
-#' lists of network data for comparison.
-#'
-#' It builds upon this package's `graphr()` function,
-#' and inherits all the same features and arguments.
-#' See `graphr()` for more.
-#' However, it uses the `{patchwork}` package to plot the graphs
-#' side by side and, if necessary, in successive rows.
-#' This is useful for lists of networks that represent, for example,
-#' ego or component subgraphs of a network,
-#' or a list of a network's different types of tie or across time.
-#' By default just the first and last network will be plotted,
-#' but this can be overridden by the "waves" parameter.
-#'
-#' Where the graphs are of the same network (same nodes),
-#' the graphs may share a layout to facilitate comparison.
-#' By default, successive graphs will use the layout calculated for
-#' the "first" network, but other options include the "last" layout,
-#' or a mix, "both", of them.
-#' @family mapping
-#' @param netlist A list of manynet-compatible networks.
-#' @param waves Numeric, the number of plots to be displayed side-by-side.
-#' If missing, the number of plots will be reduced to the first and last
-#' when there are more than four plots.
-#' This argument can also be passed a vector selecting the waves to plot.
-#' @param based_on Whether the layout of the joint plots should
-#' be based on the "first" or the "last" network, or "both".
-#' @param ... Additional arguments passed to `graphr()`.
-#' @return Multiple `ggplot2::ggplot()` objects displayed side-by-side.
-#' @name map_graphs
-#' @examples
-#' #graphs(to_egos(ison_adolescents))
-#' #graphs(to_egos(ison_adolescents), waves = 8)
-#' #graphs(to_egos(ison_adolescents), waves = c(2, 4, 6))
-#' #graphs(play_diffusion(ison_adolescents))
-#' @export
-graphs <- function(netlist, waves,
- based_on = c("first", "last", "both"), ...) {
- thisRequires("patchwork")
- based_on <- match.arg(based_on)
- if (any(class(netlist) == "diff_model")){
- if (is_list(attr(netlist, "network"))) netlist <- attr(netlist, "network") else
- netlist <- to_waves(netlist)
- }
- if (missing(waves)) {
- if (length(netlist) > 4) {
- netlist <- netlist[c(1, length(netlist))]
- snet_info("Plotting first and last waves side-by-side. \nTo set the waves plotted use the 'waves = ' argument.")
- }
- } else if (!missing(waves)) {
- if (length(waves) == 1) netlist <- netlist[c(1:waves)] else
- netlist <- netlist[waves]
- }
- if (is.null(names(netlist))) names(netlist) <- rep("", length(netlist))
- if (length(unique(lapply(netlist, length))) == 1) {
- if (based_on == "first") {
- lay <- graphr(netlist[[1]], ...)
- x <- lay$data$x
- y <- lay$data$y
- } else if (based_on == "last") {
- lay <- graphr(netlist[[length(netlist)]], ...)
- x <- lay$data$x
- y <- lay$data$y
- } else if (based_on == "both") {
- lay <- graphr(netlist[[1]], ...)
- x1 <- lay$data$x
- y1 <- lay$data$y
- lay1 <- graphr(netlist[[length(netlist)]], ...)
- x <- (lay1$data$x + x1)/2
- y <- (lay1$data$y + y1)/2
- }
- gs <- lapply(1:length(netlist), function(i)
- graphr(netlist[[i]], x = x, y = y, ...) + ggtitle(names(netlist)[i]))
- } else {
- thisRequires("methods")
- if (!methods::hasArg("layout") & is_ego_network(netlist)) {
- gs <- lapply(1:length(netlist), function(i)
- graphr(netlist[[i]], layout = "star", center = names(netlist)[[i]], ...) +
- ggtitle(names(netlist)[i]))
- } else {
- snet_info("Layouts were not standardised since not all nodes appear across waves.")
- gs <- lapply(1:length(netlist), function(i)
- graphr(netlist[[i]], ...) + ggtitle(names(netlist)[i]))
- }
- }
- # if (all(c("Infected", "Exposed", "Recovered") %in% names(gs[[1]]$data))) {
- # gs <- .collapse_guides(gs)
- # }
- do.call(patchwork::wrap_plots, c(gs, list(guides = "collect")))
-}
-
-# `graphs()` helper functions
-is_ego_network <- function(nlist) {
- if (all(unique(names(nlist)) != "")) {
- length(names(nlist)) == length(unique(unlist(unname(lapply(nlist, node_names))))) &
- all(order_alphabetically(names(nlist)) ==
- order_alphabetically(unique(unlist(unname(lapply(nlist, node_names))))))
- } else FALSE
-}
-
-order_alphabetically <- function(v) {
- v[order(names(stats::setNames(v, v)))]
-}
-
-# Dynamic networks ####
-
-#' Easily animate dynamic networks with sensible defaults
-#'
-#' @description
-#' This function provides users with an easy way to graph
-#' dynamic network data for exploration and presentation.
-#'
-#' It builds upon this package's `graphr()` function,
-#' and inherits all the same features and arguments.
-#' See `graphr()` for more.
-#' However, it uses the `{gganimate}` package to animate the changes
-#' between successive iterations of a network.
-#' This is useful for networks in which the ties and/or the node or tie
-#' attributes are changing.
-#'
-#' A progress bar is shown if it takes some time to encoding all the
-#' .png files into a .gif.
-#' @name map_grapht
-#' @family mapping
-#' @param tlist The same migraph-compatible network listed according to
-#' a time attribute, waves, or slices.
-#' @param keep_isolates Logical, whether to keep isolate nodes in the graph.
-#' TRUE by default.
-#' If FALSE, removes nodes from each frame they are isolated in.
-#' @inheritParams map_graphr
-#' @importFrom igraph gsize as_data_frame get.edgelist
-#' @importFrom ggplot2 ggplot geom_segment geom_point geom_text
-#' scale_alpha_manual theme_void
-#' @importFrom ggraph create_layout
-#' @importFrom dplyr mutate select distinct left_join %>%
-#' @source https://blog.schochastics.net/posts/2021-09-15_animating-network-evolutions-with-gganimate/
-#' @return Shows a .gif image. Assigning the result of the function
-#' saves the gif to a temporary folder and the object holds the path to this file.
-#' @examples
-#' #ison_adolescents %>%
-#' # mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) %>%
-#' # to_waves(attribute = "year", cumulative = TRUE) %>%
-#' # grapht()
-#' #ison_adolescents %>%
-#' # mutate(gender = rep(c("male", "female"), times = 4),
-#' # hair = rep(c("black", "brown"), times = 4),
-#' # age = sample(11:16, 8, replace = TRUE)) %>%
-#' # mutate_ties(year = sample(1995:1998, 10, replace = TRUE),
-#' # links = sample(c("friends", "not_friends"), 10, replace = TRUE),
-#' # weekly_meetings = sample(c(3, 5, 7), 10, replace = TRUE)) %>%
-#' # to_waves(attribute = "year") %>%
-#' # grapht(layout = "concentric", membership = "gender",
-#' # node_shape = "gender", node_color = "hair",
-#' # node_size = "age", edge_color = "links",
-#' # edge_size = "weekly_meetings")
-#' #grapht(play_diffusion(ison_adolescents, seeds = 5))
-#' @export
-grapht <- function(tlist, keep_isolates = TRUE,
- layout, labels = TRUE,
- node_color, node_shape, node_size,
- edge_color, edge_size, ...,
- node_colour, edge_colour) {
- thisRequires("gganimate")
- thisRequires("gifski")
- # thisRequires("png")
- x <- y <- name <- status <- frame <- NULL
- # Check arguments
- if (missing(layout)) {
- if (length(tlist[[1]]) == 3) {
- layout <- "triad"
- } else if (length(tlist[[1]]) == 4) {
- layout <- "quad"
- } else if (is_twomode(tlist[[1]])) {
- layout <- "hierarchy"
- } else layout <- "stress"
- }
- if (missing(node_color) && missing(node_colour)) {
- node_color <- NULL
- } else if (missing(node_color)) {
- node_color <- as.character(substitute(node_colour))
- } else {
- node_color <- as.character(substitute(node_color))
- }
- if (missing(node_shape)) node_shape <- NULL else
- node_shape <- as.character(substitute(node_shape))
- if (missing(node_size)) node_size <- NULL else if (!is.numeric(node_size)) {
- node_size <- as.character(substitute(node_size))
- }
- if (missing(edge_color) && missing(edge_colour)) {
- edge_color <- NULL
- } else if (missing(edge_color)) {
- edge_color <- as.character(substitute(edge_colour))
- } else {
- edge_color <- as.character(substitute(edge_color))
- }
- if (missing(edge_size)) edge_size <- NULL else if (!is.numeric(edge_size)) {
- edge_size <- as.character(substitute(edge_size))
- }
- # Check if diffusion model
- if (inherits(tlist, "diff_model")) tlist <- to_waves(tlist)
- # Check if object is a list of lists
- if (!is.list(tlist[[1]])) {
- snet_abort("Please declare a migraph-compatible network listed according
- to a time attribute, waves, or slices.")
- }
- # Remove lists without edges
- tlist <- Filter(function(x) igraph::gsize(x) > 0, tlist)
- # Check names for groups
- if (!"name" %in% names(node_attribute(tlist[[1]]))) {
- labels <- FALSE
- for (i in seq_len(length(tlist))) {
- tlist[[i]] <- add_node_attribute(tlist[[i]], "name",
- as.character(seq_len(igraph::vcount(tlist[[i]]))))
- }
- }
- # Create an edge list
- edges_lst <- lapply(1:length(tlist), function(i)
- cbind(igraph::as_data_frame(tlist[[i]], "edges"),
- frame = ifelse(is.null(names(tlist)), i, names(tlist)[i])))
- # Check if all names are present in all lists
- if (length(unique(unname(lapply(tlist, length)))) != 1) {
- if (any(c(node_shape, node_color, node_size) %in% names(node_attribute(tlist[[1]])))) {
- node_info <- dplyr::distinct(do.call(rbind, lapply(1:length(tlist), function(i)
- tlist[[i]] %>% activate("nodes") %>% data.frame()))) # keep node info for latter
- } else node_info <- NULL
- tlist <- to_waves(as_tidygraph(do.call("rbind", edges_lst)), attribute = "frame")
- } else node_info <- NULL
- # Add separate layouts for each time point
- lay <- lapply(1:length(tlist), function(i)
- ggraph::create_layout(tlist[[i]], layout, ...))
- # Create a node list for each time point
- nodes_lst <- lapply(1:length(tlist), function(i) {
- cbind(igraph::as_data_frame(tlist[[i]], "vertices"),
- x = lay[[i]][, 1], y = lay[[i]][, 2],
- frame = ifelse(is.null(names(tlist)), i, names(tlist)[i]))
- })
- # Create an edge list for each time point
- edges_lst <- time_edges_lst(tlist, edges_lst, nodes_lst)
- # Get edge IDs for all edges
- all_edges <- do.call("rbind", lapply(tlist, igraph::get.edgelist))
- all_edges <- all_edges[!duplicated(all_edges), ]
- all_edges <- cbind(all_edges, paste0(all_edges[, 1], "-", all_edges[, 2]))
- # Add edges level information for edge transitions
- edges_lst <- transition_edge_lst(tlist, edges_lst, nodes_lst, all_edges)
- # Bind nodes and edges list
- edges_out <- do.call("rbind", edges_lst)
- nodes_out <- do.call("rbind", nodes_lst)
- if (!is.null(node_info)) {
- nodes_out <- dplyr::left_join(nodes_out, node_info[!duplicated(node_info$name),], by = "name")
- }
- # Delete nodes for each frame if isolate
- if (isFALSE(keep_isolates)) {
- nodes_out <- remove_isolates(edges_out, nodes_out)
- } else {
- if (nrow(nodes_out)/length(unique(nodes_out$frame)) > 30 &
- any(unlist(lapply(tlist, node_is_isolate)) == TRUE)) {
- snet_info("Please considering deleting isolates to improve visualisation.")
- }
- nodes_out$status <- TRUE
- }
- # Plot with ggplot2/ggraph and animate with gganimate
- p <- map_dynamic(edges_out, nodes_out, edge_color, node_shape,
- node_color, node_size, edge_size, labels) +
- gganimate::transition_states(states = frame, transition_length = 5,
- state_length = 10, wrap = FALSE) +
- gganimate::enter_fade() +
- gganimate::exit_fade() +
- ggplot2::labs(title = "{closest_state}")
- gganimate::animate(p, duration = 2*length(tlist), start_pause = 5,
- end_pause = 10, renderer = gganimate::gifski_renderer())
-}
-
-map_dynamic <- function(edges_out, nodes_out, edge_color, node_shape,
- node_color, node_size, edge_size, labels) {
- x <- xend <- y <- yend <- id <- status <- Infected <- name <- NULL
- alphad <- ifelse(nodes_out$status == TRUE, 1, 0)
- alphae <- ifelse(edges_out$status == TRUE, 1, 0)
- if (all(unique(alphae) == 1)) alphae <- 0.8
- # Plot edges
- if (!is.null(edge_color)) {
- # Remove NAs in edge color, if declared
- if (edge_color %in% names(edges_out)) {
- edge_color <- .check_color(edges_out[[edge_color]])
- }
- } else edge_color <- "black"
- if (!is.null(edge_size)) {
- if (edge_size %in% names(edges_out)) {
- edge_size <- as.numeric(edges_out[[edge_size]])
- edge_size <- ifelse(is.na(edge_size), 0.5, edge_size)
- }
- } else edge_size <- 0.5
- p <- ggplot2::ggplot() +
- ggplot2::geom_segment(aes(x = x, xend = xend, y = y, yend = yend, group = id),
- alpha = alphae, data = edges_out, color = edge_color,
- linewidth = edge_size, show.legend = FALSE)
- # Set node shape, color, and size
- if (!is.null(node_shape)) {
- if (node_shape %in% names(nodes_out)) {
- node_shape <- as.factor(nodes_out[[node_shape]])
- if (!any(grepl("circle|square|triangle", node_shape))) {
- node_shape <- c("circle", "square", "triangle")[node_shape]
- }
- }
- } else node_shape <- "circle"
- if (!is.null(node_color)) {
- if (node_color %in% names(nodes_out)) {
- node_color <- .check_color(nodes_out[[node_color]])
- }
- } else if (is.null(node_color) & "Infected" %in% names(nodes_out)) {
- node_color <- as.factor(ifelse(nodes_out[["Exposed"]], "Exposed",
- ifelse(nodes_out[["Infected"]],"Infected",
- ifelse(nodes_out[["Recovered"]], "Recovered",
- "Susceptible"))))
- } else node_color <- "darkgray"
- if (!is.null(node_size)) {
- if (node_size %in% names(nodes_out)) {
- node_size <- nodes_out[[node_size]]
- }
- } else if (nrow(nodes_out) > 100) {
- node_size <- 3
- } else node_size <- nrow(nodes_out)/length(unique(nodes_out$frame))
- # Add labels
- if (isTRUE(labels)) {
- p <- p + ggplot2::geom_text(aes(x, y, label = name), alpha = alphad,
- data = nodes_out, color = "black",
- hjust = -0.2, vjust = -0.2, show.legend = FALSE)
- }
- # Plot nodes
- if ("Infected" %in% names(nodes_out)) {
- p <- p + ggplot2::geom_point(aes(x, y, group = name, color = node_color),
- size = node_size, shape = node_shape, data = nodes_out) +
- ggplot2::scale_color_manual(name = NULL, values = c("Infected" = "#d73027",
- "Susceptible" = "#4575b4",
- "Exposed" = "#E6AB02",
- "Recovered" = "#66A61E")) +
- ggplot2::theme_void() +
- ggplot2::theme(legend.position = "bottom")
- } else {
- p <- p + ggplot2::geom_point(aes(x, y, group = name), alpha = alphad,
- size = node_size, data = nodes_out,
- color = node_color, shape = node_shape,
- show.legend = FALSE) +
- ggplot2::theme_void()
- }
- p
-}
-
-# `graphd()` helper functions
-.check_color <- function(v) {
- color <- grDevices::colors()
- color <- color[!color %in% "black"]
- v <- ifelse(is.na(v), "black", v)
- if (!any(grepl(paste(color, collapse = "|"), v)) | any(grepl("^#", v))) {
- for(i in unique(v)) {
- if (i != "black") {
- v[v == i] <- sample(color, 1)
- }
- }
- }
- v
-}
-
-time_edges_lst <- function(tlist, edges_lst, nodes_lst, edge_color) {
- lapply(1:length(tlist), function(i) {
- edges_lst[[i]]$x <- nodes_lst[[i]]$x[match(edges_lst[[i]]$from,
- nodes_lst[[i]]$name)]
- edges_lst[[i]]$y <- nodes_lst[[i]]$y[match(edges_lst[[i]]$from,
- nodes_lst[[i]]$name)]
- edges_lst[[i]]$xend <- nodes_lst[[i]]$x[match(edges_lst[[i]]$to,
- nodes_lst[[i]]$name)]
- edges_lst[[i]]$yend <- nodes_lst[[i]]$y[match(edges_lst[[i]]$to,
- nodes_lst[[i]]$name)]
- edges_lst[[i]]$id <- paste0(edges_lst[[i]]$from, "-", edges_lst[[i]]$to)
- edges_lst[[i]]$status <- TRUE
- edges_lst[[i]]
- })
-}
-
-transition_edge_lst <- function(tlist, edges_lst, nodes_lst, all_edges) {
- x <- lapply(1:length(tlist), function(i) {
- idx <- which(!all_edges[, 3] %in% edges_lst[[i]]$id)
- if (length(idx) != 0) {
- tmp <- data.frame(from = all_edges[idx, 1], to = all_edges[idx, 2],
- id = all_edges[idx, 3])
- tmp$x <- nodes_lst[[i]]$x[match(tmp$from, nodes_lst[[i]]$name)]
- tmp$y <- nodes_lst[[i]]$y[match(tmp$from, nodes_lst[[i]]$name)]
- tmp$xend <- nodes_lst[[i]]$x[match(tmp$to, nodes_lst[[i]]$name)]
- tmp$yend <- nodes_lst[[i]]$y[match(tmp$to, nodes_lst[[i]]$name)]
- tmp$frame <- ifelse(is.null(names(tlist)), i, names(tlist)[i])
- tmp$status <- FALSE
- edges_lst[[i]] <- dplyr::bind_rows(edges_lst[[i]], tmp)
- }
- edges_lst[[i]]
- })
-}
-
-remove_isolates <- function(edges_out, nodes_out) {
- status <- frame <- from <- to <- framen <- NULL
- # Create node metadata for node presence in certain frame
- meta <- edges_out %>%
- dplyr::filter(status == TRUE) %>%
- dplyr::mutate(framen = match(frame, unique(frame)),
- meta = ifelse(framen > 1, paste0(from, (framen - 1)), from)) %>%
- dplyr::select(meta, status) %>%
- dplyr::distinct()
- metab <- edges_out %>%
- dplyr::filter(status == TRUE) %>%
- dplyr::mutate(framen = match(frame, unique(frame)),
- meta = ifelse(framen > 1, paste0(to, (framen - 1)), to)) %>%
- dplyr::select(meta, status) %>%
- rbind(meta) %>%
- dplyr::distinct()
- # Mark nodes that are isolates
- nodes_out$meta <- rownames(nodes_out)
- # Join data
- nodes_out <- dplyr::left_join(nodes_out, metab, by = "meta") %>%
- dplyr::mutate(status = ifelse(is.na(status), FALSE, TRUE)) %>%
- dplyr::distinct()
-}
diff --git a/R/map_layouts.R b/R/map_layouts.R
deleted file mode 100644
index 7e55dfe8..00000000
--- a/R/map_layouts.R
+++ /dev/null
@@ -1,565 +0,0 @@
-# Configurational ####
-
-#' Layout algorithms based on configurational positions
-#'
-#' @description
-#' Configurational layouts locate nodes at symmetric coordinates
-#' to help illustrate particular configurations.
-#' Currently "triad" and "quad" layouts are available.
-#' The "configuration" layout will choose the appropriate configurational
-#' layout automatically.
-#'
-#' @name map_layout_configuration
-#' @family mapping
-#' @inheritParams map_layout_partition
-NULL
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_configuration <- function(.data,
- circular = FALSE, times = 1000){
- if (net_nodes(.data) == 2) {
- layout_tbl_graph_dyad(.data, circular = circular, times = times)
- } else if (net_nodes(.data) == 3) {
- layout_tbl_graph_triad(.data, circular = circular, times = times)
- } else if (net_nodes(.data) == 4) {
- layout_tbl_graph_tetrad(.data, circular = circular, times = times)
- } else if (net_nodes(.data) == 5) {
- layout_tbl_graph_pentad(.data, circular = circular, times = times)
- } else if (net_nodes(.data) == 6) {
- layout_tbl_graph_hexad(.data, circular = circular, times = times)
- }
-}
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_dyad <- function(.data,
- circular = FALSE, times = 1000){
- res <- matrix(c(0,0,
- 1,0), 2, 2, byrow = TRUE)
- .to_lo(res)
-}
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_triad <- function(.data,
- circular = FALSE, times = 1000){
- res <- matrix(c(0,0,
- 2,3.5,
- 4,0), 3, 2, byrow = TRUE)
- .to_lo(res)
-}
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_tetrad <- function(.data,
- circular = FALSE, times = 1000){
- res <- matrix(c(0,0,
- 0,1,
- 1,0,
- 1,1), 4, 2, byrow = TRUE)
- .to_lo(res)
-}
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_pentad <- function(.data,
- circular = FALSE, times = 1000){
- res <- matrix(c(0,1,
- -0.9511,0.3090,
- -0.5878,-0.8090,
- 0.5878,-0.8090,
- 0.9511,0.3090), 5, 2, byrow = TRUE)
- .to_lo(res)
-}
-
-#' @rdname map_layout_configuration
-#' @export
-layout_tbl_graph_hexad <- function(.data,
- circular = FALSE, times = 1000){
- res <- matrix(c(1,0,
- 1/2,sqrt(3)/2,
- -1/2,sqrt(3)/2,
- -1,0,
- -1/2,-sqrt(3)/2,
- 1/2,-sqrt(3)/2), 6, 2, byrow = TRUE)
- .to_lo(res)
-}
-
-# Partitions ####
-
-#' Layout algorithms based on bi- or other partitions
-#'
-#' @description
-#' These algorithms layout networks based on two or more partitions,
-#' and are recommended for use with `graphr()` or `{ggraph}`.
-#' Note that these layout algorithms use `{Rgraphviz}`,
-#' a package that is only available on Bioconductor.
-#' It will first need to be downloaded using `BiocManager::install("Rgraphviz")`.
-#' If it has not already been installed, there is a prompt the first time
-#' these functions are used though.
-#'
-#' The "hierarchy" layout layers the first node set along the bottom,
-#' and the second node set along the top,
-#' sequenced and spaced as necessary to minimise edge overlap.
-#' The "alluvial" layout is similar to "hierarchy",
-#' but places successive layers horizontally rather than vertically.
-#' The "railway" layout is similar to "hierarchy",
-#' but nodes are aligned across the layers.
-#' The "ladder" layout is similar to "railway",
-#' but places successive layers horizontally rather than vertically.
-#' The "concentric" layout places a "hierarchy" layout
-#' around a circle, with successive layers appearing as concentric circles.
-#' The "multilevel" layout places successive layers as multiple levels.
-#' The "lineage" layout ranks nodes in Y axis according to values.
-#' @name map_layout_partition
-#' @inheritParams mark_is
-#' @param circular Should the layout be transformed into a radial representation.
-#' Only possible for some layouts. Defaults to FALSE.
-#' @param times Maximum number of iterations, where appropriate
-#' @param radius A vector of radii at which the concentric circles
-#' should be located for "concentric" layout.
-#' By default this is equal placement around an empty centre,
-#' unless one (the core) is a single node,
-#' in which case this node occupies the centre of the graph.
-#' @param order.by An attribute label indicating the (decreasing) order
-#' for the nodes around the circles for "concentric" layout.
-#' By default ordering is given by a bipartite placement that reduces
-#' the number of edge crossings.
-#' @param membership A node attribute or a vector to draw concentric circles
-#' for "concentric" layout.
-#' @param center Further split "hierarchical" layouts by
-#' declaring the "center" argument as the "events", "actors",
-#' or by declaring a node name in hierarchy layout.
-#' Defaults to NULL.
-#' @param level A node attribute or a vector to hierarchically order levels for
-#' "multilevel" layout.
-#' @param rank A numerical node attribute to place nodes in Y axis
-#' according to values for "lineage" layout.
-#' @family mapping
-#' @source
-#' Diego Diez, Andrew P. Hutchins and Diego Miranda-Saavedra. 2014.
-#' "Systematic identification of transcriptional regulatory modules from
-#' protein-protein interaction networks".
-#' _Nucleic Acids Research_, 42 (1) e6.
-NULL
-
-#' @rdname map_layout_partition
-#' @examples
-#' #graphr(ison_southern_women, layout = "hierarchy", center = "events",
-#' # node_color = "type", node_size = 3)
-#' @export
-layout_tbl_graph_hierarchy <- function(.data, center = NULL,
- circular = FALSE, times = 1000) {
- if (is.null(center)) {
- thisRequiresBio("Rgraphviz")
- prep <- as_matrix(.data)
- if (anyDuplicated(rownames(prep))) {
- rownames(prep) <- seq_len(nrow(prep))
- colnames(prep) <- seq_len(ncol(prep)) + max(nrow(prep))
- }
- if (any(prep<0)) prep[prep<0] <- 0
- out <- as_graphAM(prep)
- out <- suppressMessages(Rgraphviz::layoutGraph(out, layoutType = 'dot',
- attrs = list(graph = list(rankdir = "BT"))))
- nodeX <- .rescale(out@renderInfo@nodes$nodeX)
- nodeY <- .rescale(out@renderInfo@nodes$nodeY)
- if (is_twomode(.data) & "name" %in% igraph::vertex_attr_names(.data)) {
- names <- igraph::vertex_attr(.data, "name")
- nodeX <- nodeX[order(match(names(nodeX), names))]
- nodeY <- nodeY[order(match(names(nodeY), names))]
- }
- # nodeY <- abs(nodeY - max(nodeY))
- out <- .to_lo(cbind(nodeX, nodeY))
- } else {
- if (!is_twomode(.data)) snet_abort("Please declare a two-mode network.")
- net <- as_matrix(.data)
- nn <- dim(net)[1]
- mm <- dim(net)[2]
- if (center == "actors") {
- Act <- cbind(rep(1, nrow(net)), nrm(rng(nn)))
- Evt1 <- cbind(rep(0, ceiling(ncol(net)/2)), nrm(rng(ceiling(mm/2))))
- Evt2 <- cbind(rep(2, floor(ncol(net)/2)), nrm(rng(floor(mm/2))))
- crd <- rbind(Act, Evt1, Evt2)
- crd[which(is.nan(crd))] <- 0.5
- rownames(crd) <- c(dimnames(net)[[1]], dimnames(net)[[2]])
- } else if (center == "events") {
- Act1 <- cbind(rep(0, ceiling(nrow(net)/2)), nrm(rng(ceiling(nn/2))))
- Act2 <- cbind(rep(2, floor(nrow(net)/2)), nrm(rng(floor(nn/2))))
- Evt <- cbind(rep(1, ncol(net)), nrm(rng(mm)))
- crd <- rbind(Act1, Act2, Evt)
- crd[which(is.nan(crd))] <- 0.5
- rownames(crd) <- c(dimnames(net)[[1]], dimnames(net)[[2]])
- } else {
- if (center %in% node_names(.data)) {
- side1 <- suppressWarnings(cbind(rep(0, nrow(net)), nrm(rng(nn))))
- side2 <- suppressWarnings(cbind(rep(2, ncol(net)), nrm(rng(mm))))
- if (any(rownames(net) == center)) {
- side1[,1] <- ifelse(rownames(net) == center, 1, side1[,1])
- side1[,2] <- ifelse(rownames(net) == center, 0.5, side1[,2])
- } else {
- side2[,1] <- ifelse(rownames(net) == center, 1, side2[,1])
- side2[,2] <- ifelse(rownames(net) == center, 0.5, side2[,2])
- }
- crd <- rbind(side1, side2)
- crd[which(is.nan(crd))] <- 0.5
- rownames(crd) <- c(dimnames(net)[[1]], dimnames(net)[[2]])
- } else snet_abort("Please declare actors, events, or a node name as center.")
- }
- out <- .to_lo(crd)
- }
- out
-}
-
-#' @rdname map_layout_partition
-#' @examples
-#' #graphr(ison_southern_women, layout = "alluvial")
-#' @export
-layout_tbl_graph_alluvial <- function(.data,
- circular = FALSE, times = 1000){
- thisRequiresBio("Rgraphviz")
- prep <- as_matrix(.data, twomode = FALSE)
- if(anyDuplicated(rownames(prep))){
- rownames(prep) <- seq_len(nrow(prep))
- colnames(prep) <- seq_len(ncol(prep))
- }
- if(any(prep<0)) prep[prep<0] <- 0
- out <- as_graphAM(prep)
- out <- suppressMessages(Rgraphviz::layoutGraph(out, layoutType = 'dot',
- attrs = list(graph = list(rankdir = "LR"))))
- nodeX <- .rescale(out@renderInfo@nodes$nodeX)
- nodeY <- .rescale(out@renderInfo@nodes$nodeY)
- # nodeY <- abs(nodeY - max(nodeY))
- .to_lo(cbind(nodeX, nodeY))
-}
-
-#' @rdname map_layout_partition
-#' @export
-layout_tbl_graph_railway <- function(.data,
- circular = FALSE, times = 1000) {
- res <- layout_tbl_graph_hierarchy(as_igraph(.data))
- res$x <- c(match(res[res[,2]==0,1], sort(res[res[,2]==0,1])),
- match(res[res[,2]==1,1], sort(res[res[,2]==1,1])))
- res
-}
-
-#' @rdname map_layout_partition
-#' @export
-layout_tbl_graph_ladder <- function(.data,
- circular = FALSE, times = 1000){
- res <- layout_tbl_graph_alluvial(as_igraph(.data))
- res$y <- c(match(res[res[,2]==1,1], sort(res[res[,2]==1,1])),
- match(res[res[,2]==0,1], sort(res[res[,2]==0,1])))
- res
-}
-
-#' @rdname map_layout_partition
-#' @examples
-#' #graphr(ison_southern_women, layout = "concentric", membership = "type",
-#' # node_color = "type", node_size = 3)
-#' @export
-layout_tbl_graph_concentric <- function(.data, membership,
- radius = NULL,
- order.by = NULL,
- circular = FALSE, times = 1000) {
- if (any(igraph::vertex_attr(.data, "name") == "")) {
- ll <- unlist(lapply(seq_len(length(.data)), function(x) {
- ifelse(igraph::vertex_attr(.data, "name")[x] == "",
- paste0("ramdom", x), igraph::vertex_attr(.data, "name")[x])
- }))
- .data <- set_vertex_attr(.data, "name", value = ll)
- }
- if (missing(membership)) {
- if (is_twomode(.data)) membership <- node_is_mode(.data) else
- snet_abort("Please pass the function a `membership` node attribute or a vector.")
- } else {
- if (length(membership) > 1 & length(membership) != length(.data)) {
- snet_abort("Please pass the function a `membership` node attribute or a vector.")
- } else if (length(membership) != length(.data)) {
- membership <- node_attribute(.data, membership)
- }
- }
- names(membership) <- node_names(.data)
- membership <- to_list(membership)
- all_c <- unlist(membership, use.names = FALSE)
- if (any(table(all_c) > 1)) snet_abort("Duplicated nodes in layers!")
- if (is_labelled(.data)) all_n <- node_names(.data) else all_n <- 1:net_nodes(.data)
- sel_other <- all_n[!all_n %in% all_c]
- if (length(sel_other) > 0) membership[[length(membership) + 1]] <- sel_other
- if (is.null(radius)) {
- radius <- seq(0, 1, 1/(length(membership)))
- if (length(membership[[1]]) == 1)
- radius <- radius[-length(radius)] else radius <- radius[-1]
- }
- if (!is.null(order.by)) {
- order.values <- lapply(order.by,
- function(b) node_attribute(.data, b))
- } else {
- if (is_twomode(.data) & length(membership) == 2) {
- xnet <- as_matrix(to_multilevel(.data))[membership[[2-1]],
- membership[[2]]]
- lo <- layout_tbl_graph_hierarchy(as_igraph(xnet, twomode = TRUE))
- lo$names <- node_names(.data)
- if (ncol(lo) == 2) lo[,1] <- seq_len(dim(lo)[1])
- order.values <- lapply(1:0, function(x)
- if(ncol(lo) >= 3) sort(lo[lo[,2] == x,])[,3]
- else sort(lo[lo[,2] == x,1]))
- } else order.values <- membership[order(sapply(membership, length))]
- # order.values <- getNNvec(.data, members)
- }
- res <- matrix(NA, nrow = length(all_n), ncol = 2)
- for (k in seq_along(membership)) {
- r <- radius[k]
- l <- order.values[[k]]
- if(is_labelled(.data))
- l <- match(l, node_names(.data))
- res[l, ] <- getCoordinates(l, r)
- }
- .to_lo(res)
-}
-
-#' @rdname map_layout_partition
-#' @examples
-#' #graphr(ison_lotr, layout = "multilevel",
-#' # node_color = "Race", level = "Race", node_size = 3)
-#' @export
-layout_tbl_graph_multilevel <- function(.data, level, circular = FALSE) {
- if (missing(level)) {
- if (any(grepl("lvl", names(node_attribute(.data))))) {
- message("Level attribute 'lvl' found in data.")
- } else {
- snet_abort("Please pass the function a `level` node attribute or a vector.")
- }
- } else {
- if (length(level) > 1 & length(level) != length(.data)) {
- snet_abort("Please pass the function a `level` node attribute or a vector.")
- } else if (length(level) != length(.data)) {
- level <- as.factor(node_attribute(.data, level))
- }
- }
- out <- igraph::set_vertex_attr(.data, "lvl", value = level)
- thisRequires("graphlayouts")
- out <- graphlayouts::layout_as_multilevel(out, alpha = 25)
- .to_lo(out)
-}
-
-#' @rdname map_layout_partition
-#' @examples
-#' # ison_adolescents %>%
-#' # mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2),
-#' # cut = node_is_cutpoint(ison_adolescents)) %>%
-#' # graphr(layout = "lineage", rank = "year", node_color = "cut",
-#' # node_size = migraph::node_degree(ison_adolescents)*10)
-#' @export
-layout_tbl_graph_lineage <- function(.data, rank, circular = FALSE) {
- if (length(rank) > 1 & length(rank) != length(.data)) {
- snet_abort("Please pass the function a `rank` node attribute or a vector.")
- } else if (length(rank) != length(.data)) {
- rank <- as.numeric(node_attribute(.data, rank))
- }
- thisRequiresBio("Rgraphviz")
- out <- layout_tbl_graph_alluvial(
- as_igraph(mutate(.data, type = ifelse(
- rank > mean(rank), TRUE, FALSE)), twomode = TRUE))
- out$x <- .rescale(rank)
- .check_dup(out)
-}
-
-.rescale <- function(vector){
- (vector - min(vector)) / (max(vector) - min(vector))
-}
-
-.to_lo <- function(mat) {
- res <- as.data.frame(mat)
- names(res) <- c("x","y")
- res
-}
-
-to_list <- function(members) {
- out <- lapply(sort(unique(members)), function(x){
- y <- which(members==x)
- if(!is.null(names(y))) names(y) else y
- })
- names(out) <- unique(members)
- out
-}
-
-.check_dup <- function(mat) {
- mat$y <- ifelse(duplicated(mat[c('x','y')]), mat$y*0.95, mat$y)
- mat
-}
-
-#' @importFrom igraph degree
-getNNvec <- function(.data, members){
- lapply(members, function(circle){
- diss <- 1 - stats::cor(to_multilevel(as_matrix(.data))[, circle])
- diag(diss) <- NA
- if(is_labelled(.data))
- starts <- names(sort(igraph::degree(.data)[circle], decreasing = TRUE)[1])
- else starts <- paste0("V",1:net_nodes(.data))[sort(igraph::degree(.data)[circle],
- decreasing = TRUE)[1]]
- if(length(circle)>1)
- starts <- c(starts, names(which.min(diss[starts,])))
- out <- starts
- if(length(circle)>2){
- for(i in 1:(length(circle)-2)){
- diss <- diss[,!colnames(diss) %in% starts]
- if(is.matrix(diss)){
- side <- names(which.min(apply(diss[starts,], 1, min, na.rm = TRUE)))
- new <- names(which.min(diss[side,]))
- } else {
- side <- names(which.min(diss[starts]))
- new <- setdiff(circle,out)
- }
- if(side == out[1]){
- out <- c(new, out)
- starts <- c(new, starts[2])
- } else {
- out <- c(out, new)
- starts <- c(starts[1], new)
- }
- }
- }
- out
- })
-}
-
-getCoordinates <- function(x, r) {
- l <- length(x)
- d <- 360/l
- c1 <- seq(0, 360, d)
- c1 <- c1[1:(length(c1) - 1)]
- tmp <- t(vapply(c1,
- function(cc) c(cos(cc * pi/180) *
- r, sin(cc *
- pi/180) * r),
- FUN.VALUE = numeric(2)))
- rownames(tmp) <- x
- tmp
-}
-
-rng <- function(r) {
- if (r == 1L) return(0)
- if (r > 1L) {
- x <- vector()
- x <- append(x, (-1))
- for (i in 1:(r - 1)) x <- append(x, ((-1) + (2L/(r - 1L)) * i))
- return(x * (r/50L))
- } else snet_abort("no negative values")
-}
-
-nrm <- function(x, digits = 3) {
- if (isTRUE(length(x) == 1L) == TRUE) return(x)
- if (is.array(x) == TRUE) {
- xnorm <- (x[, 1] - min(x[, 1]))/(max(x[, 1]) - min(x[, 1]))
- rat <- (max(x[, 1]) - min(x[, 1]))/(max(x[, 2]) - min(x[, 2]))
- ynorm <- ((x[, 2] - min(x[, 2]))/(max(x[, 2]) - min(x[, 2]))) * (rat)
- ifelse(isTRUE(rat > 0) == FALSE,
- ynorm <- ((x[, 2] - min(x[, 2]))/(max(x[, 2]) -
- min(x[, 2]))) * (1L/rat), NA)
- return(round(data.frame(X = xnorm, Y = ynorm), digits))
- }
- else if (is.vector(x) == TRUE) {
- return(round((x - min(x))/(max(x) - min(x)), digits))
- }
-}
-
-# Grid ####
-
-#' Layouts for snapping layouts to a grid
-#'
-#' @description The function uses approximate pattern matching
-#' to redistribute coarse layouts on square grid points, while
-#' preserving the topological relationships among the nodes (see Inoue et al. 2012).
-#' @references
-#' Inoue, Kentaro, Shinichi Shimozono, Hideaki Yoshida, and Hiroyuki Kurata. 2012.
-#' “Application of Approximate Pattern Matching in Two Dimensional Spaces to Grid Layout for Biochemical Network Maps” edited by J. Bourdon.
-#' _PLoS ONE_ 7(6):e37739.
-#' \doi{https://doi.org/10.1371/journal.pone.0037739}.
-#' @keywords internal
-depth_first_recursive_search <- function(layout) {
- if("ggraph" %in% class(layout)) layout <- layout$data[,c("x","y")]
- layout <- as.data.frame(layout)
- dims <- ceiling(2 * sqrt(nrow(layout)))
- # evens <- 0:dims[0:dims %% 2 == 0]
- vacant_points <- expand.grid(seq.int(0, dims, 1), seq.int(0, dims, 1)) # create options
- vacant_points <- vacant_points - floor(dims / 2) # centre options
- names(vacant_points) <- c("x", "y")
- gridout <- layout[order(abs(layout[,1]) + abs(layout[,2])), ] # sort centroid distance
- nodes <- seq_len(nrow(gridout))
- for (i in nodes) {
- dists <- as.matrix(stats::dist(rbind(gridout[i, 1:2], vacant_points),
- method = "manhattan"))[, 1]
- mindist <- which(dists == min(dists[2:length(dists)]))[1] - 1
- vacpoint <- vacant_points[mindist, ]
- changes <- vacpoint - gridout[i, 1:2]
- gridout[nodes >= i, 1] <- gridout[nodes >= i, 1] +
- changes[[1]]
- gridout[nodes >= i, 2] <- gridout[nodes >= i, 2] +
- changes[[2]]
- vacant_points <- vacant_points[-mindist, ]
- }
- gridout[order(row.names(gridout)),] # reorder from centroid
- # gridout
- # plot(gridout[order(row.names(gridout)),])
-}
-
-# localmin <- function(layout, graph) {
-# repeat {
-# f0 <- sum(cost_function(layout, graph))
-# L <- get_vacant_points(layout)
-# for (a in seq_len(nrow(layout))) {
-# out <- t(apply(L, 1, function(y) {
-# layout_new <- layout
-# layout_new[a, 1:2] <- y
-# c(a, y, sum(cost_function(layout_new, graph)))
-# }))
-# }
-# if (out[which.min(out[, 4]), 4] < f0) {
-# layout[out[which.min(out[, 4]), 1], 1:2] <- out[which.min(out[, 4]), 2:3]
-# } else{
-# break
-# }
-# }
-# layout
-# }
-#
-# get_vacant_points <- function(layout) {
-# all_points <- expand.grid(min(layout$x):max(layout$x),
-# min(layout$y):max(layout$y))
-# names(all_points) <- c("x", "y")
-# vacant_points <- rbind(all_points,
-# layout[, c("x", "y")])
-# vacant_points <- subset(vacant_points,
-# !(duplicated(vacant_points) |
-# duplicated(vacant_points, fromLast = TRUE)))
-# vacant_points
-# }
-#
-# cost_function <- function(layout, graph, max_repulse_distance = max(layout[, 1]) * .75) {
-# d <- as.matrix(dist(layout[, 1:2], method = "manhattan"))
-# a <- as_matrix(graph)
-# i <- diag(nrow(a))
-# m <- a + i
-# w <- ifelse(m > 0, 3,
-# ifelse(m == 0 & m %*% t(m) > 0, 0, -2)) # only three levels here
-# # see Li and Kurata (2005: 2037) for more granulated option
-# ifelse(w >= 0, w * d, w * min(d, max_repulse_distance))
-# }
-#
-# plot_gl <- function(x, tmax, tmin, rmin, fmin, ne, rc, p) {
-# l <- index <- a <- NULL # initialize variables to avoid CMD check notes
-# x <- as_tidygraph(x)
-# lo <- ggraph::create_layout(x, layout = "igraph", algorithm = "randomly")
-# lo[, 1] <- round(lo[, 1] * 1000)
-# lo[, 2] <- round(lo[, 2] * 1000)
-# dists <- as.matrix(dist(lo[, 1:2], method = "manhattan"))
-# colMax <- function(data) apply(data, MARGIN = 1, FUN = max, na.rm = TRUE)
-# diag(dists) <- NA
-# rsep <- l * sum(ifelse(colMax(a / dists - 1) > 0, colMax(a / dists - 1), 0))
-# ggraph::ggraph(x, graph = lo) +
-# ggraph::geom_edge_link(ggplot2::aes(alpha = ggplot2::stat(index)),
-# show.legend = FALSE) +
-# ggraph::geom_node_point()
-# }
-
diff --git a/R/map_palettes.R b/R/map_palettes.R
deleted file mode 100644
index d1cf1128..00000000
--- a/R/map_palettes.R
+++ /dev/null
@@ -1,42 +0,0 @@
-#' Many palettes generator
-#'
-#' @param palette Name of desired palette. Current choices are:
-#' \code{IHEID}, \code{Centres}, \code{SDGs}, \code{ETHZ}, \code{RUG},
-#' and \code{UZH}.
-#' @param n Number of colors desired. If omitted, uses all colours.
-#' @param type Either "continuous" or "discrete". Use continuous if you want
-#' to automatically interpolate between colours.
-#' @return A graphic display of colours in palette.
-#' @name map_palettes
-#' @source Adapted from \url{https://github.com/karthik/wesanderson/blob/master/R/colors.R}
-#' @examples
-#' many_palettes()
-#' #many_palettes("IHEID")
-#' @export
-many_palettes <- function(palette, n, type = c("discrete", "continuous")) {
- type <- match.arg(type)
- if (missing(palette)) {
- pal <- corp_palette(c("IHEID", "SDGs", "Centres", "ETHZ", "RUG", "UZH"))
- } else pal <- corp_palette(palette)
- if (is.null(pal))
- snet_abort("Palette not found.")
- if (missing(n)) {
- n <- length(pal)
- }
- if (type == "discrete" && n > length(pal)) {
- snet_abort("Number of requested colors greater than what palette can offer")
- }
- x <- switch(type, continuous = grDevices::colorRampPalette(pal)(n),
- discrete = pal[1:n])
- old <- graphics::par(mar = c(0.5, 0.5, 0.5, 0.5))
- on.exit(graphics::par(old))
- graphics::image(1:n, 1, as.matrix(1:n), col = x,
- ylab = "", xaxt = "n", yaxt = "n", bty = "n")
- graphics::rect(0, 0.9, n + 1, 1.1, col = grDevices::rgb(1, 1, 1, 0.8),
- border = NA)
- graphics::text(c(seq_len(n)), 1, labels = attr(x, "name"),
- cex = 0.9, family = "serif", srt = 90)
-}
-
-colorsafe_palette <- c("#d73027", "#4575b4", "#1B9E77","#D95F02","#7570B3",
- "#E7298A", "#66A61E","#E6AB02","#A6761D","#666666")
diff --git a/R/map_plot.R b/R/map_plot.R
deleted file mode 100644
index 51ba1c05..00000000
--- a/R/map_plot.R
+++ /dev/null
@@ -1,272 +0,0 @@
-#' #' Plot agreements network
-#' #'
-#' #' @description Facilitates plotting of 'many' data.
-#' #' @param dataset A dataset from one of the many packages
-#' #' or a "consolidated" database.
-#' #' @param actor An actor variable.
-#' #' "stateID", by default.
-#' #' @param treaty_type The type of treaties to be returned.
-#' #' NULL, by default.
-#' #' Other options are "bilateral" or "multilateral".
-#' #' @param key An ID column to collapse by.
-#' #' By default "manyID".
-#' #' @param layout How do you want the plot to look like?
-#' #' An `{ggraph}` layout algorithm.
-#' #' If not declared, reasonable defaults are used.
-#' #' @name plot_
-#' NULL
-#'
-#' #' @rdname plot_
-#' #' @importFrom dplyr %>% select mutate distinct rename
-#' #' @return A network of agreements' relations.
-#' #' @examples
-#' #' \donttest{
-#' #' #agreements <- dplyr::filter(manyenviron::agreements$ECOLEX,
-#' #' #Begin > "2000-01-01" & Begin < "2000-12-12")
-#' #' #plot_agreements(agreements)
-#' #'}
-#' #' @export
-#' plot_agreements <- function(dataset, treaty_type = NULL, key = "manyID",
-#' layout = "circle") {
-#' manyID <- treatyID <- name <- NULL
-#' if (key == "manyID") {
-#' out <- dplyr::select(dataset, manyID) %>%
-#' dplyr::rename(key = manyID) %>%
-#' dplyr::distinct()
-#' } else if (key == "treatyID") {
-#' out <- dplyr::select(dataset, treatyID) %>%
-#' dplyr::rename(key == treatyID) %>%
-#' dplyr::distinct()
-#' } else snet_abort("Please declare either 'manyID' or 'treatyID'.")
-#' if (!is.null(treaty_type)) {
-#' if (treaty_type == "bilateral") {
-#' out <- out[grep("-", out$key),]
-#' }
-#' if (treaty_type == "multilateral") {
-#' out <- out[grep("-", out$key, invert = TRUE),]
-#' }
-#' }
-#' dplyr::mutate(out,
-#' link = ifelse(grepl(":", key), sapply(strsplit(key, ":"),
-#' "[", 2), "NA"),
-#' key = gsub("\\:.*", "", key)) %>%
-#' as_tidygraph() %>%
-#' dplyr::filter(name != "NA") %>%
-#' graphr(layout = layout)
-#' }
-#'
-#' #' @rdname plot_
-#' #' @importFrom dplyr %>% select distinct all_of rename
-#' #' @return A network of agreements' memberships.
-#' #' @examples
-#' #' \donttest{
-#' #' #memberships <- dplyr::filter(manyenviron::memberships$ECOLEX_MEM,
-#' #' #Begin > "2000-01-01" & Begin < "2000-01-31")
-#' #' #plot_memberships(memberships)
-#' #'}
-#' #' @export
-#' plot_memberships <- function(dataset, actor = "stateID", treaty_type = NULL,
-#' key = "manyID", layout = "bipartite") {
-#' manyID <- treatyID <- name <- NULL
-#' if (key == "manyID") {
-#' out <- dplyr::select(dataset, manyID, dplyr::all_of(actor)) %>%
-#' dplyr::rename(key = manyID) %>%
-#' dplyr::distinct()
-#' } else if (key == "treatyID") {
-#' out <- dplyr::select(dataset, treatyID, dplyr::all_of(actor)) %>%
-#' dplyr::rename(key == treatyID) %>%
-#' dplyr::distinct()
-#' } else snet_abort("Please declare either 'manyID' or 'treatyID'.")
-#' if (!is.null(treaty_type)) {
-#' if (treaty_type == "bilateral") {
-#' out <- out[grep("-", out$key),]
-#' }
-#' if (treaty_type == "multilateral") {
-#' out <- out[grep("-", out$key, invert = TRUE),]
-#' }
-#' }
-#' stats::na.omit(out) %>%
-#' as_tidygraph() %>%
-#' mutate(type = ifelse(grepl("[0-9][0-9][0-9][0-9][A-Za-z]",
-#' name), TRUE, FALSE)) %>%
-#' graphr(layout = layout)
-#' }
-#'
-#' #' @rdname plot_
-#' #' @importFrom dplyr %>% select mutate distinct filter rename
-#' #' @return A plot of agreements' lineages.
-#' #' @examples
-#' #' \donttest{
-#' #' #lineage <- dplyr::filter(manyenviron::agreements$HUGGO,
-#' #' #Begin > "2000-01-01", Begin < "2001-12-31")
-#' #' #plot_lineage(lineage)
-#' #' }
-#' #' @export
-#' plot_lineage <- function(dataset, treaty_type = NULL, key = "manyID",
-#' layout = "nicely") {
-#' manyID <- treatyID <- name <- NULL
-#' if (key == "manyID") {
-#' out <- dplyr::select(dataset, manyID) %>%
-#' dplyr::rename(key = manyID) %>%
-#' dplyr::distinct()
-#' } else if (key == "treatyID") {
-#' out <- dplyr::select(dataset, treatyID) %>%
-#' dplyr::rename(key == treatyID) %>%
-#' dplyr::distinct()
-#' } else snet_abort("Please declare either 'manyID' or 'treatyID'.")
-#' if (!is.null(treaty_type)) {
-#' if (treaty_type == "bilateral") {
-#' out <- out[grep("-", out$key),]
-#' }
-#' if (treaty_type == "multilateral") {
-#' out <- out[grep("-", out$key, invert = TRUE),]
-#' }
-#' }
-#' out %>%
-#' dplyr::filter(grepl(":", key)) %>%
-#' dplyr::mutate(key1 = gsub(".*\\:", "", key),
-#' key = gsub("\\:.*", "", key)) %>%
-#' dplyr::distinct() %>%
-#' as_tidygraph() %>%
-#' graphr(layout = "nicely")
-#' }
-#'
-#' #' @rdname plot_
-#' #' @param date String date from the network snapshot.
-#' #' Used by \code{{cshapes}} to plot the correct map.
-#' #' By default, 2019-12-31.
-#' #' Date can be between 1886-01-01 and 2019-12-31.
-#' #' @param theme Theme you would like to use to plot the graph.
-#' #' bey defalt, "light".
-#' #' Available themes are "light", "dark", and "earth".
-#' #' @details `plot_map()` creates a plot of the a unimodal geographical network
-#' #' at a single point in time.
-#' #' @importFrom dplyr mutate inner_join rename filter
-#' #' @return A map of a country level geographical network.
-#' #' @examples
-#' #' \donttest{
-#' #' #memberships <- dplyr::filter(manyenviron::memberships$ECOLEX_MEM,
-#' #' #Begin > "2000-01-01" & Begin < "2000-12-12")
-#' #' #plot_map(memberships, actor = "stateID") +
-#' #' #ggplot2::labs(title = "Bilateral International Environmental Treaties Signed in the year 2000",
-#' #' #subtitle = "Ecolex data")
-#' #'}
-#' #' @export
-#' plot_map <- function(dataset, actor = "stateID", treaty_type = NULL,
-#' date = "2019-12-31", theme = "light") {
-#' # check packages
-#' thisRequires("cshapes")
-#' thisRequires("manydata")
-#' # Checks for correct input
-#' weight <- .data <- NULL
-#' # Step 1: get membership list
-#' dataset <- manydata::call_treaties(dataset = dataset, actor = actor,
-#' treaty_type = treaty_type)
-#' # Step 2: set up empty matrix
-#' actor <- unique(unlist(strsplit(dataset$Memberships, ", ")))
-#' out <- matrix(0, nrow = length(actor), ncol = length(actor))
-#' rownames(out) <- actor
-#' colnames(out) <- actor
-#' # Step 3: fill matrix with values
-#' for (k in colnames(out)) {
-#' m <- data.frame(table(unlist(strsplit(grep(k, dataset$Memberships,
-#' value = TRUE), ", "))))
-#' m <- m[!(m$Var1 %in% k),]
-#' out[k, ] <- ifelse(names(out[k,]) %in% m$Var1 == TRUE, m$Freq/100, out[k,])
-#' }
-#' out <- igraph::get.data.frame(igraph::graph.adjacency(out, weighted = TRUE))
-#' # Step 4 = get theme
-#' if (theme == "dark") {
-#' maptheme <- maptheme(palette = c("#FFFAFA", "#596673"))
-#' countrycolor <- "#FFFAFA"
-#' }
-#' if (theme == "earth") {
-#' maptheme <- maptheme(palette = c("#79B52F", "#4259FD"))
-#' countrycolor <- "#79B52F"
-#' }
-#' if (theme == "light") {
-#' maptheme <- maptheme(palette = c("#596673", "#FFFAFA"))
-#' countrycolor <- "#596673"
-#' }
-#' # Step 5: import the historical shapefile data
-#' cshapes <- cshapes::cshp(as.Date(date), useGW = FALSE)
-#' coment <- vapply(countryregex[, 3], # add stateID abbreviations
-#' function(x) grepl(x, cshapes$country_name,
-#' ignore.case = TRUE, perl = TRUE) * 1,
-#' FUN.VALUE = double(length(cshapes$country_name)))
-#' colnames(coment) <- countryregex[, 1]
-#' rownames(coment) <- cshapes$country_name
-#' ab <- apply(coment, 1, function(x) paste(names(x[x == 1]),
-#' collapse = "_"))
-#' ab[ab == ""] <- NA
-#' cshapes <- dplyr::mutate(cshapes, stateID = unname(ab))
-#' # Step 6: create edges with from/to lat/long
-#' edges <- out %>%
-#' dplyr::inner_join(cshapes, by = c("from" = "stateID")) %>%
-#' dplyr::rename(x = .data$caplong, y = .data$caplat) %>%
-#' dplyr::inner_join(cshapes, by = c("to" = "stateID")) %>%
-#' dplyr::rename(xend = .data$caplong, yend = .data$caplat)
-#' # Step 7: Create plotted network from computed edges
-#' g <- as_tidygraph(edges)
-#' # Step 8: Get the country shapes from the edges dataframe
-#' country_shapes <- ggplot2::geom_sf(data = cshapes$geometry,
-#' fill = countrycolor)
-#' # Step 9: generate the point coordinates for capitals
-#' cshapes_pos <- cshapes %>%
-#' dplyr::filter(.data$stateID %in% node_names(g)) %>%
-#' dplyr::rename(x = .data$caplong, y = .data$caplat)
-#' # Reorder things according to nodes in plotted network g
-#' cshapes_pos <- cshapes_pos[match(node_names(g),
-#' cshapes_pos[["stateID"]]), ]
-#' # Step 10: generate the layout
-#' lay <- ggraph::create_layout(g, layout = cshapes_pos)
-#' edges$circular <- rep(FALSE, nrow(edges))
-#' edges$edge.id <- rep(1, nrow(edges))
-#' # Step 11: plot
-#' ggraph::ggraph(lay) +
-#' country_shapes +
-#' ggraph::geom_edge_arc(data = edges, ggplot2::aes(edge_width = weight),
-#' strength = 0.33, alpha = 0.25) +
-#' ggraph::scale_edge_width_continuous(range = c(0.5, 2), # scales edge widths
-#' guide = "none") +
-#' ggraph::geom_node_point(shape = 21, # draw nodes
-#' fill = "white", color = "black", stroke = 0.5) +
-#' ggraph::geom_node_text(ggplot2::aes(label = node_names(g)),
-#' repel = TRUE, size = 3, color = "white",
-#' fontface = "bold") +
-#' maptheme
-#' }
-#'
-#' # Helper function providing the network map function with a few map themes.
-#' maptheme <- function(palette = c("#FFFAFA", "#596673")) {
-#' oceancolor <- palette[2]
-#' titlecolor <- ifelse(is_dark(palette[2]), "white", "black")
-#' # Create map theme
-#' maptheme <- ggplot2::theme(panel.grid = ggplot2::element_blank()) +
-#' ggplot2::theme(axis.text = ggplot2::element_blank()) +
-#' ggplot2::theme(axis.ticks = ggplot2::element_blank()) +
-#' ggplot2::theme(axis.title = ggplot2::element_blank()) +
-#' ggplot2::theme(legend.position = "bottom") +
-#' ggplot2::theme(panel.grid = ggplot2::element_blank()) +
-#' ggplot2::theme(panel.background = ggplot2::element_blank()) +
-#' ggplot2::theme(plot.background = ggplot2::element_rect(fill = oceancolor)) +
-#' ggplot2::theme(plot.title = ggplot2::element_text(color = titlecolor,
-#' hjust = 0.1, vjust = 0.1),
-#' plot.subtitle = ggplot2::element_text(color = titlecolor,
-#' hjust = 0.065,
-#' vjust = 0.1),
-#' plot.caption = ggplot2::element_text(color = titlecolor, hjust = 0.96)) +
-#' ggplot2::theme(plot.margin = ggplot2::unit(c(0, 0, 0.5, 0), "cm"))
-#' # This function returns a map theme for ggplot
-#' maptheme
-#' }
-#'
-#' # Helper function to check whether a color is light or dark:
-#' is_dark <- function(hex) {
-#' # Google luma formula for details.
-#' luma <- 0.33 * grDevices::col2rgb(hex)[[1]] +
-#' 0.5 * grDevices::col2rgb(hex)[[2]] +
-#' 0.16 * grDevices::col2rgb(hex)[[3]]
-#' isdark <- ifelse(luma < 186, TRUE, FALSE)
-#' isdark
-#' }
diff --git a/R/map_theme.R b/R/map_theme.R
deleted file mode 100644
index c3d6f895..00000000
--- a/R/map_theme.R
+++ /dev/null
@@ -1,558 +0,0 @@
-#' Many themes
-#'
-#' @description
-#' These functions enable graphs to be easily and quickly themed,
-#' e.g. changing the default colour of the graph's vertices and edges.
-#' @name map_themes
-#' @param base_size Font size, by default 12.
-#' @param base_family Font family, by default "sans".
-#' @examples
-#' to_mentoring(ison_brandes) %>%
-#' mutate(color = c(rep(c(1,2,3), 3), 3)) %>%
-#' graphr(node_color = "color") +
-#' labs(title = "Who leads and who follows?") +
-#' scale_color_iheid() +
-#' theme_iheid()
-NULL
-
-#' @rdname map_themes
-#' @param theme String naming a theme.
-#' By default "default".
-#' @export
-set_manynet_theme <- function(theme = "default"){
- theme_opts <- c("default",
- "iheid", "ethz", "uzh", "rug",
- "crisp", "neon", "rainbow")
- if(theme %in% theme_opts){
- options(mnet_theme = theme)
- set_highlight_theme(theme)
- set_background_theme(theme)
- set_categorical_theme(theme)
- cli::cli_alert_success("Theme set to {.emph {theme}}.")
- } else {
- cli::cli_alert_warning("Please choose one of the available themes: {.emph {theme_opts}}.")
- }
-}
-
-set_highlight_theme <- function(theme){
- if(theme == "iheid"){
- options(mnet_highlight = c("#000010","#E20020"))
- } else if(theme == "rug"){
- options(mnet_highlight = c("#000000", "#dc002d"))
- } else if(theme == "uzh"){
- options(mnet_highlight = c("#a3adb7", "#dc6027"))
- } else if(theme == "ethz"){
- options(mnet_highlight = c("#6F6F6F", "#0028a5"))
- } else if(theme == "crisp"){
- options(mnet_highlight = c("#FFFFFA", "#101314"))
- } else if(theme == "neon"){
- options(mnet_highlight = c("#5aeafd", "#54fe4b"))
- } else if(theme == "rainbow"){
- options(mnet_highlight = c('#1965B0', '#DC050C'))
- } else {
- options(mnet_highlight = c("#4576B5", "#D83127"))
- }
-}
-
-set_background_theme <- function(theme){
- if(theme == "neon"){
- options(mnet_background = "#070f23")
- } else {
- options(mnet_background = "#FFFFFF")
- }
-}
-
-set_categorical_theme <- function(theme){
- if(theme == "iheid"){
- options(mnet_cat = c("#006564","#0094D8","#622550",
- "#268D2B","#3E2682","#820C2B",
- "#008F92","#006EAA","#A8086E"))
- } else if(theme == "ethz"){
- options(mnet_cat = c("#215CAF","#007894","#627313",
- "#8E6713","#B7352D","#A7117A","#6F6F6F"))
- } else if(theme == "uzh"){
- options(mnet_cat = c("#0028A5","#4AC9E3","#A4D233",
- "#FFC845","#FC4C02","#BF0D3E",
- "#BDC9E8","#DBF4F9","#ECF6D6",
- "#FFF4DA","#FFDBCC","#FBC6D4",
- "#7596FF","#B7E9F4","#DBEDAD",
- "#FFE9B5","#FEB799","#F78CAA",
- "#3062FF","#92DFEE","#C8E485",
- "#FFDE8F","#FE9367","#F3537F",
- "#001E7C","#1EA7C4","#7CA023",
- "#F3AB00","#BD3902","#8F0A2E",
- "#001452","#147082","#536B18",
- "#A27200","#7E2601","#60061F"))
-
- } else if(theme == "rainbow"){
- options(mnet_cat = c('#E8ECFB', '#D9CCE3', '#D1BBD7',
- '#CAACCB', '#BA8DB4', '#AE76A3',
- '#AA6F9E', '#994F88', '#882E72',
- '#1965B0', '#437DBF', '#5289C7',
- '#6195CF', '#7BAFDE',
- '#4EB265', '#90C987', '#CAE0AB',
- '#F7F056', '#F7CB45', '#F6C141',
- '#F4A736', '#F1932D', '#EE8026',
- '#E8601C', '#E65518', '#DC050C',
- '#A5170E', '#72190E', '#42150A'))
- } else {
- options(mnet_cat = c("#1B9E77","#4575b4","#d73027",
- "#66A61E","#E6AB02","#D95F02","#7570B3",
- "#A6761D","#E7298A","#666666"))
- }
-}
-
-
-
-#' @rdname map_themes
-#' @export
-theme_iheid <- function(base_size = 12, base_family = "serif") {
- colors <- corp_palette("IHEID")
- (ggplot2::theme_minimal(base_size = base_size, base_family = base_family)
- + ggplot2::theme(
- line = ggplot2::element_line(colors["IHEIDBlack"]),
- rect = ggplot2::element_rect(fill = "#FFFFFF", linetype = 1,
- linewidth = 0.6, colour = colors["IHEIDGrey"]),
- title = ggplot2::element_text(colour = colors["IHEIDRed"],
- size = base_size, family = base_family,
- face = "bold"),
- plot.subtitle = ggplot2::element_text(colors["IHEIDGrey"],
- size = base_size*0.85, family = base_family,
- face = "bold.italic"),
- plot.caption = ggplot2::element_text(colors["IHEIDBlack"],
- size = base_size*0.75,
- family = base_family,
- face = "italic"),
- axis.title = ggplot2::element_blank(),
- axis.text = ggplot2::element_text(),
- axis.ticks = ggplot2::element_blank(),
- axis.line = ggplot2::element_blank(),
- axis.title.x = ggplot2::element_blank(),
- axis.text.x = ggplot2::element_blank(),
- axis.ticks.x = ggplot2::element_blank(),
- axis.title.y = ggplot2::element_blank(),
- axis.text.y = ggplot2::element_blank(),
- axis.ticks.y = ggplot2::element_blank(),
- panel.grid.major = ggplot2::element_blank(),
- panel.grid.minor = ggplot2::element_blank(),
- legend.background = ggplot2::element_rect(),
- legend.position = "bottom",
- legend.direction = "horizontal",
- legend.box = "vertical",
- strip.background = ggplot2::element_rect()))
-}
-
-#' @rdname map_themes
-#' @export
-theme_ethz <- function(base_size = 12, base_family = "sans") {
- colors <- corp_palette("ETHZ")
- (ggplot2::theme_minimal(base_size = base_size, base_family = base_family)
- + ggplot2::theme(
- line = ggplot2::element_line(colors["ETHZ_Green"]),
- rect = ggplot2::element_rect(fill = "#FFFFFF", linetype = 1,
- linewidth = 0.6, colour = colors["ETHZ_Petrol"]),
- title = ggplot2::element_text(colour = "black",
- size = base_size, family = base_family,
- face = "bold.italic"),
- plot.subtitle = ggplot2::element_text("black",
- size = base_size*0.85,
- family = base_family,
- face = "bold.italic"),
- plot.caption = ggplot2::element_text("black",
- size = base_size*0.75,
- family = base_family,
- face = "italic"),
- axis.title = ggplot2::element_blank(),
- axis.text = ggplot2::element_text(),
- axis.ticks = ggplot2::element_blank(),
- axis.line = ggplot2::element_blank(),
- axis.title.x = ggplot2::element_blank(),
- axis.text.x = ggplot2::element_blank(),
- axis.ticks.x = ggplot2::element_blank(),
- axis.title.y = ggplot2::element_blank(),
- axis.text.y = ggplot2::element_blank(),
- axis.ticks.y = ggplot2::element_blank(),
- panel.grid.major = ggplot2::element_blank(),
- panel.grid.minor = ggplot2::element_blank(),
- legend.background = ggplot2::element_rect(),
- legend.position = "bottom",
- legend.direction = "horizontal",
- legend.box = "vertical",
- strip.background = ggplot2::element_rect()))
-}
-
-#' @rdname map_themes
-#' @export
-theme_uzh <- function(base_size = 12, base_family = "sans") {
- colors <- corp_palette("UZH")
- (ggplot2::theme_minimal(base_size = base_size, base_family = base_family)
- + ggplot2::theme(
- line = ggplot2::element_line(colors["UZH_Blue"]),
- rect = ggplot2::element_rect(fill = "#FFFFFF", linetype = 1,
- linewidth = 0.6, colour = colors["UZH_Grey"]),
- title = ggplot2::element_text(colour = colors["UZH_Orange"],
- size = base_size, family = base_family,
- face = "bold"),
- plot.subtitle = ggplot2::element_text(colors["UZH_Blue"],
- size = base_size*0.85,
- family = base_family,
- face = "bold.italic"),
- plot.caption = ggplot2::element_text(colors["UZH_Grey"],
- size = base_size*0.75,
- family = base_family,
- face = "italic"),
- axis.title = ggplot2::element_blank(),
- axis.text = ggplot2::element_text(),
- axis.ticks = ggplot2::element_blank(),
- axis.line = ggplot2::element_blank(),
- axis.title.x = ggplot2::element_blank(),
- axis.text.x = ggplot2::element_blank(),
- axis.ticks.x = ggplot2::element_blank(),
- axis.title.y = ggplot2::element_blank(),
- axis.text.y = ggplot2::element_blank(),
- axis.ticks.y = ggplot2::element_blank(),
- panel.grid.major = ggplot2::element_blank(),
- panel.grid.minor = ggplot2::element_blank(),
- legend.background = ggplot2::element_rect(),
- legend.position = "bottom",
- legend.direction = "horizontal",
- legend.box = "vertical",
- strip.background = ggplot2::element_rect()))
-}
-
-#' @rdname map_themes
-#' @export
-theme_rug <- function(base_size = 12, base_family = "mono") {
- colors <- corp_palette("RUG")
- (ggplot2::theme_minimal(base_size = base_size, base_family = base_family)
- + ggplot2::theme(
- line = ggplot2::element_line(colors["RUG_Red"]),
- rect = ggplot2::element_rect(fill = "#FFFFFF", linetype = 1,
- linewidth = 0.6, colour = colors["RUG_Black"]),
- title = ggplot2::element_text(colour = colors["RUG_Red"],
- size = base_size, family = base_family,
- face = "bold"),
- plot.subtitle = ggplot2::element_text(colors["RUG_Black"],
- size = base_size*0.85,
- family = base_family,
- face = "bold.italic"),
- plot.caption = ggplot2::element_text(colors["RUG_Red"],
- size = base_size*0.75,
- family = base_family,
- face = "italic"),
- axis.title = ggplot2::element_blank(),
- axis.text = ggplot2::element_text(),
- axis.ticks = ggplot2::element_blank(),
- axis.line = ggplot2::element_blank(),
- axis.title.x = ggplot2::element_blank(),
- axis.text.x = ggplot2::element_blank(),
- axis.ticks.x = ggplot2::element_blank(),
- axis.title.y = ggplot2::element_blank(),
- axis.text.y = ggplot2::element_blank(),
- axis.ticks.y = ggplot2::element_blank(),
- panel.grid.major = ggplot2::element_blank(),
- panel.grid.minor = ggplot2::element_blank(),
- legend.background = ggplot2::element_rect(),
- legend.position = "bottom",
- legend.direction = "horizontal",
- legend.box = "vertical",
- strip.background = ggplot2::element_rect()))
-}
-
-#' Many scales
-#'
-#' @description These functions enable to add color scales to be graphs.
-#' @name map_scales
-#' @param direction Direction for using palette colors.
-#' @param ... Extra arguments passed to `ggplot2::discrete_scale()`.
-#' @examples
-#' #ison_brandes %>%
-#' #mutate(core = migraph::node_is_core(ison_brandes)) %>%
-#' #graphr(node_color = "core") +
-#' #scale_color_iheid()
-#' #graphr(ison_physicians[[1]], edge_color = "type") +
-#' #scale_edge_color_ethz()
-NULL
-
-#' @rdname map_scales
-#' @export
-scale_fill_iheid <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "IHEID", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_iheid <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "IHEID", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_iheid <- scale_colour_iheid
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_iheid <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "IHEID", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_iheid <- scale_edge_colour_iheid
-
-#' Centres color scales
-#'
-#' @rdname map_scales
-#' @export
-scale_fill_centres <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "Centres", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_centres <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "Centres", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_centres <- scale_colour_centres
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_centres <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "Centres", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_centres <- scale_edge_colour_centres
-
-#' SDGs color scales
-#'
-#' @rdname map_scales
-#' @export
-scale_fill_sdgs <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "SDGs", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_sdgs <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "SDGs", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_sdgs <- scale_colour_sdgs
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_sdgs <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "SDGs", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_sdgs <- scale_edge_colour_sdgs
-
-#' ETHZ color scales
-#'
-#' @rdname map_scales
-#' @export
-scale_fill_ethz <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "ETHZ", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_ethz <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "ETHZ", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_ethz <- scale_colour_ethz
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_ethz <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "ETHZ", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_ethz <- scale_edge_colour_ethz
-
-#' UZH color scales
-#'
-#' @rdname map_scales
-#' @export
-scale_fill_uzh <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "UZH", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_uzh <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "UZH", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_uzh <- scale_colour_uzh
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_uzh <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "UZH", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_uzh <- scale_edge_colour_uzh
-
-#' RUG color scales
-#'
-#' @rdname map_scales
-#' @export
-scale_fill_rug <- function(direction = 1, ...) {
- ggplot2::discrete_scale("fill",
- palette = palette_gen(palette = "RUG", direction),
- na.value = "grey", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_colour_rug <- function(direction = 1, ...) {
- ggplot2::discrete_scale("colour",
- palette = palette_gen(palette = "RUG", direction),
- na.value = "grey", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_color_rug <- scale_colour_rug
-
-#' @rdname map_scales
-#' @export
-scale_edge_colour_rug <- function(direction = 1, ...) {
- ggplot2::discrete_scale("edge_colour",
- palette = palette_gen(palette = "RUG", direction),
- na.value = "black", name = "", ...)
-}
-
-#' @rdname map_scales
-#' @export
-scale_edge_color_rug <- scale_edge_colour_rug
-
-# Helper functions
-corp_color <- function(...) {
- corp_colors <- c(`IHEIDRed` = "#E20020", `IHEIDBlack` = "#000010",
- `IHEIDGrey` = "#6f7072", `AHCD` = "#622550",
- `CFFD` = "#0094D8", `CIES` = "#268D2B",
- `CTEI` = "#008F92", `CGEN` = "#820C2B",
- `CCDP` = "#3E2682", `GLGC` = "#006564",
- `GLHC` = "#A8086E", `GLMC` = "#006EAA",
- `NoPoverty` = "#e5243b", `ZeroHunger` = "#DDA63A",
- `GoodHealth` = "#4C9F38",
- `QualityEducation` = "#C5192D",
- `GenderEquality` = "#FF3A21", `CleanWater` = "#26BDE2",
- `CleanEnergy` = "#FCC30B",
- `EconomicGrowth` = "#A21942",
- `Innovation` = "#FD6925",
- `ReducedInequalities` = "#DD1367",
- `SustainableCities` = "#FD9D24",
- `ResponsibleConsumption` = "#BF8B2E",
- `ClimateAction` = "#3F7E44", `BelowWater` = "#0A97D9",
- `OnLand` = "#56C02B", `StrongInstitutions` = "#00689D",
- `GoalPartnerships` = "#19486A",
- `ETHZ_Blue` = "#215CAF", `ETHZ_Petrol` = "#007894",
- `ETHZ_Green` = "#627313", `ETHZ_Bronze` = "#8E6713",
- `ETHZ_Red`= "#B7352D", `ETHZ_Purple` = "#A7117A",
- `ETHZ_Grey` = "#6F6F6F", `UZH_Blue` = "#0028a5",
- `UZH_Grey` = "#a3adb7", `UZH_Orange` = "#dc6027",
- `RUG_Red` = "#dc002d", `RUG_Black` = "#000000")
- cols <- c(...)
- if (is.null(cols))
- return (corp_colors)
- corp_colors[cols]
-}
-
-corp_palette <- function(palette, ...) {
- corp_palettes <- list(`IHEID` = corp_color("IHEIDRed",
- "IHEIDBlack",
- "IHEIDGrey"),
- `Centres` = corp_color("AHCD", "CFFD",
- "CIES", "CTEI",
- "CGEN", "CCDP",
- "GLGC", "GLHC",
- "GLMC"),
- `SDGs` = corp_color("NoPoverty",
- "ZeroHunger",
- "GoodHealth",
- "QualityEducation",
- "GenderEquality",
- "CleanWater",
- "CleanEnergy",
- "EconomicGrowth",
- "Innovation",
- "ReducedInequalities",
- "SustainableCities",
- "ResponsibleConsumption",
- "ClimateAction",
- "BelowWater",
- "OnLand",
- "StrongInstitutions",
- "GoalPartnerships"),
- `ETHZ` = corp_color("ETHZ_Blue", "ETHZ_Petrol",
- "ETHZ_Green", "ETHZ_Bronze",
- "ETHZ_Red", "ETHZ_Purple", "ETHZ_Grey"),
- `UZH` = corp_color("UZH_Blue", "UZH_Grey", "UZH_Orange"),
- `RUG` = corp_color("RUG_Red", "RUG_Black"))
- unlist(unname(corp_palettes[c(palette)]))
-}
-
-palette_gen <- function(palette, direction = 1) {
- function(n) {
- if (n > length(corp_palette(palette)))
- warning("Not enough colors in this palette!")
- else {
- all_colors <- corp_palette(palette)
- all_colors <- unname(unlist(all_colors))
- all_colors <- if (direction >= 0) all_colors else rev(all_colors)
- color_list <- all_colors[1:n]
- }
- }
-}
-
diff --git a/R/mark_ties.R b/R/mark_ties.R
index f2f14b28..e7939d41 100644
--- a/R/mark_ties.R
+++ b/R/mark_ties.R
@@ -86,8 +86,8 @@ tie_is_bridge <- function(.data){
#' @importFrom igraph all_shortest_paths
#' @examples
#' ison_adolescents %>%
-#' mutate_ties(route = tie_is_path(from = "Jane", to = 7)) %>%
-#' graphr(edge_colour = "route")
+#' mutate_ties(route = tie_is_path(from = "Jane", to = 7))
+#' #graphr(edge_colour = "route")
#' @export
tie_is_path <- function(.data, from, to, all_paths = FALSE){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -125,8 +125,8 @@ NULL
#' @importFrom igraph triangles
#' @examples
#' ison_monks %>% to_uniplex("like") %>%
-#' mutate_ties(tri = tie_is_triangular()) %>%
-#' graphr(edge_color = "tri")
+#' mutate_ties(tri = tie_is_triangular())
+#' #graphr(edge_color = "tri")
#' @export
tie_is_triangular <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -146,8 +146,8 @@ tie_is_triangular <- function(.data){
#' @rdname mark_triangles
#' @examples
#' ison_adolescents %>% to_directed() %>%
-#' mutate_ties(trans = tie_is_transitive()) %>%
-#' graphr(edge_color = "trans")
+#' mutate_ties(trans = tie_is_transitive())
+#' #graphr(edge_color = "trans")
#' @export
tie_is_transitive <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -163,8 +163,8 @@ tie_is_transitive <- function(.data){
#' @rdname mark_triangles
#' @examples
#' ison_adolescents %>% to_directed() %>%
-#' mutate_ties(trip = tie_is_triplet()) %>%
-#' graphr(edge_color = "trip")
+#' mutate_ties(trip = tie_is_triplet())
+#' #graphr(edge_color = "trip")
#' @export
tie_is_triplet <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -185,8 +185,8 @@ tie_is_triplet <- function(.data){
#' @rdname mark_triangles
#' @examples
#' ison_adolescents %>% to_directed() %>%
-#' mutate_ties(cyc = tie_is_cyclical()) %>%
-#' graphr(edge_color = "cyc")
+#' mutate_ties(cyc = tie_is_cyclical())
+#' #graphr(edge_color = "cyc")
#' @export
tie_is_cyclical <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -202,8 +202,8 @@ tie_is_cyclical <- function(.data){
#' @rdname mark_triangles
#' @examples
#' ison_monks %>% to_uniplex("like") %>%
-#' mutate_ties(simmel = tie_is_simmelian()) %>%
-#' graphr(edge_color = "simmel")
+#' mutate_ties(simmel = tie_is_simmelian())
+#' #graphr(edge_color = "simmel")
#' @export
tie_is_simmelian <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
@@ -218,8 +218,8 @@ tie_is_simmelian <- function(.data){
#' @rdname mark_triangles
#' @examples
#' generate_random(8, directed = TRUE) %>%
-#' mutate_ties(forbid = tie_is_forbidden()) %>%
-#' graphr(edge_color = "forbid")
+#' mutate_ties(forbid = tie_is_forbidden())
+#' #graphr(edge_color = "forbid")
#' @export
tie_is_forbidden <- function(.data){
if(missing(.data)) {expect_edges(); .data <- .G()}
diff --git a/R/measure_centrality.R b/R/measure_centrality.R
index a58f0944..88c8dc85 100644
--- a/R/measure_centrality.R
+++ b/R/measure_centrality.R
@@ -444,9 +444,7 @@ node_stress <- function(.data, normalized = TRUE){
#' @importFrom igraph edge_betweenness
#' @examples
#' (tb <- tie_betweenness(ison_adolescents))
-#' plot(tb)
-#' ison_adolescents %>% mutate_ties(weight = tb) %>%
-#' graphr()
+#' ison_adolescents %>% mutate_ties(weight = tb)
#' @export
tie_betweenness <- function(.data, normalized = TRUE){
if(missing(.data)) {expect_nodes(); .data <- .G()}
@@ -783,9 +781,7 @@ node_vitality <- function(.data, normalized = TRUE){
#' @rdname measure_central_close
#' @examples
#' (ec <- tie_closeness(ison_adolescents))
-#' plot(ec)
-#' ison_adolescents %>% mutate_ties(weight = ec) %>%
-#' graphr()
+#' ison_adolescents %>% mutate_ties(weight = ec)
#' @export
tie_closeness <- function(.data, normalized = TRUE){
if(missing(.data)) {expect_nodes(); .data <- .G()}
diff --git a/R/measure_features.R b/R/measure_features.R
index 54a61794..3c1d60bc 100644
--- a/R/measure_features.R
+++ b/R/measure_features.R
@@ -33,22 +33,62 @@
NULL
#' @rdname measure_features
-#' @examples
-#' net_core(ison_adolescents)
-#' net_core(ison_southern_women)
+#' @param method Which method of the following to use to calculate the fit of
+#' the core assignment to a core-periphery model.
+#' "correlation" calculates the correlation between the empirical network and
+#' an ideal typical network, and "ident" calculates the Euclidean distances
+#' between the same.
+#' "ndiff", however, calculates how distinct the core and periphery groups are
+#' based on the difference in coreness scores between the least core-like
+#' member of the core and the most core-like member of the periphery.
+#' "diff" is similar to "ndiff", but multiplies the raw "ndiff" score by the
+#' square root of the size of the core, thus penalising large cores.
+#' @section Core-Periphery:
+#' `net_core()` calculates the Pearson correlation between the given network,
+#' where the nodes in the core are assigned by some given mark, and an ideal
+#' typical core-periphery network with the same number of nodes in the core
+#' and the periphery.
#' @references
#' ## On core-periphery
#' Borgatti, Stephen P., and Martin G. Everett. 2000.
#' “Models of Core/Periphery Structures.”
#' _Social Networks_ 21(4):375–95.
#' \doi{10.1016/S0378-8733(99)00019-2}
+#' @examples
+#' net_core(ison_adolescents)
+#' net_core(ison_southern_women)
#' @export
net_core <- function(.data,
- mark = NULL){
+ mark = NULL,
+ method = c("correlation","ident","ndiff", "diff")){
if(missing(.data)) {expect_nodes(); .data <- .G()}
if(is.null(mark)) mark <- node_is_core(.data)
- out <- stats::cor(c(as_matrix(.data)),
- c(as_matrix(create_core(.data, mark = mark))))
+
+ method <- match.arg(method)
+ if(method == "correlation"){
+ out <- stats::cor(c(as_matrix(.data)),
+ c(as_matrix(create_core(.data, mark = mark))))
+ } else if(method == "ident"){
+ out <- sqrt(sum((as_matrix(.data) -
+ as_matrix(create_core(.data, mark = mark)))^2))
+ } else if(method %in% c("ndiff","diff")){
+ # Sort nodes by coreness
+ c_scores <- node_coreness(.data)
+ core <- c_scores[mark]
+ periphery <- c_scores[!mark]
+
+ min_core <- min(core)
+ max_periphery <- max(periphery)
+
+ diff1 <- sum(min_core - periphery)
+ diff2 <- sum(core - max_periphery)
+
+ if(method == "ndiff"){
+ out <- (diff1 + diff2) / length(c_scores) # Normalize
+ } else if(method == "diff"){
+ out <- (diff1 + diff2) * sqrt(sum(mark))
+ }
+ } else snet_unavailable(method)
make_network_measure(out, .data, call = deparse(sys.call()))
}
diff --git a/R/member_components.R b/R/member_components.R
index f4eea507..fec09ea6 100644
--- a/R/member_components.R
+++ b/R/member_components.R
@@ -36,8 +36,7 @@ NULL
#' @importFrom igraph components
# #' @examples
# #' ison_monks %>% to_uniplex("esteem") %>%
-# #' mutate_nodes(comp = node_in_component()) %>%
-# #' graphr(node_color = "comp")
+# #' mutate_nodes(comp = node_in_component())
#' @export
node_in_component <- function(.data){
if(missing(.data)) {expect_nodes(); .data <- .G()}
diff --git a/R/member_core.R b/R/member_core.R
index 8139593c..8080a14f 100644
--- a/R/member_core.R
+++ b/R/member_core.R
@@ -2,8 +2,13 @@
#' @description
#' These functions identify nodes belonging to (some level of) the core of a network:
#'
-#' - `node_is_core()` assigns nodes to either the core or periphery.
-#' - `node_coreness()` assigns nodes to their level of k-coreness.
+#' - `node_is_universal()` identifies whether nodes are adjacent to all other
+#' nodes in the network.
+#' - `node_is_core()` identifies whether nodes belong to the core of the
+#' network, as opposed to the periphery.
+#' - `node_coreness()` returns a continuous measure of how closely each node
+#' resembles a typical core node.
+#' - `node_kcoreness()` assigns nodes to their level of k-coreness.
#'
#' @inheritParams mark_is
#' @param method Which method to use to identify cores and periphery.
@@ -90,12 +95,33 @@ node_is_core <- function(.data, method = c("degree", "eigenvector")){
#' @rdname mark_core
#' @examples
-#' node_coreness(ison_adolescents)
+#' node_kcoreness(ison_adolescents)
#' @export
-node_coreness <- function(.data){
+node_kcoreness <- function(.data){
if(missing(.data)) {expect_nodes(); .data <- .G()}
if(!manynet::is_graph(.data)) .data <- manynet::as_igraph(.data)
out <- igraph::coreness(.data)
make_node_measure(out, .data)
}
+#' @rdname mark_core
+#' @examples
+#' node_coreness(ison_adolescents)
+#' @export
+node_coreness <- function(.data) {
+ A <- as_matrix(.data)
+ n <- nrow(A)
+ obj_fun <- function(c) {
+ ideal <- outer(c, c)
+ val <- suppressWarnings(cor(as.vector(A), as.vector(ideal)))
+ if (!is.finite(val)) return(1e6) # Penalize non-finite values
+ return(-val) # Negative for maximization
+ }
+ # Initial guess: all nodes have coreness 0.5
+ init <- rep(0.5, n)
+ result <- stats::optim(init, obj_fun, method = "L-BFGS-B",
+ lower = 0, upper = 1)
+ make_node_measure(result$par, .data)
+}
+
+
diff --git a/R/reexports_ggplot2.R b/R/reexports_ggplot2.R
deleted file mode 100644
index c65073a9..00000000
--- a/R/reexports_ggplot2.R
+++ /dev/null
@@ -1,36 +0,0 @@
-#' @importFrom ggplot2 ggplot
-#' @export
-ggplot2::ggplot
-
-#' @importFrom ggplot2 ggtitle
-#' @export
-ggplot2::ggtitle
-
-#' @importFrom ggplot2 guides
-#' @export
-ggplot2::guides
-
-#' @importFrom ggplot2 labs
-#' @export
-ggplot2::labs
-
-#' @importFrom ggplot2 xlab
-#' @export
-ggplot2::xlab
-
-#' @importFrom ggplot2 ylab
-#' @export
-ggplot2::ylab
-
-#' @importFrom ggplot2 aes
-#' @export
-ggplot2::aes
-
-#' @importFrom ggplot2 ggsave
-#' @export
-ggplot2::ggsave
-
-#' @importFrom ggplot2 scale_size
-#' @export
-ggplot2::scale_size
-
diff --git a/README.Rmd b/README.Rmd
index aafa223a..743d0c85 100644
--- a/README.Rmd
+++ b/README.Rmd
@@ -51,8 +51,9 @@ which can mess up your pretty presentation or paper.
This can make learning and using network analysis tools in R challenging.
By contrast, `{manynet}` offers _many_ analytic tools that work on _many_ (if not most) types and kinds of networks.
-It helps researchers make, modify, map, mark, measure, and identify nodes' motifs and memberships in networks.
-For further testing and modelling capabilities,
+It helps researchers make, modify, mark, measure, and identify nodes' motifs and memberships in networks.
+For graph drawing see [`{autograph}`](https://stocnet.github.io/autograph/),
+and for further testing and modelling capabilities
see [`{migraph}`](https://stocnet.github.io/migraph/) and the other [stocnet](https://github.com/stocnet) packages.
- [Making](#making)
@@ -65,13 +66,6 @@ see [`{migraph}`](https://stocnet.github.io/migraph/) and the other [stocnet](ht
- [Transforming](#transforming)
- [Splitting and Joining](#splitting-and-joining)
- [Extracting](#extracting)
-- [Mapping](#mapping)
- - [Graphing](#graphing)
- - [More options](#more-options)
- - [More layouts](#more-layouts)
- - [More themes and scales](#more-themes-and-scales)
- - [graphs](#graphs)
- - [grapht](#grapht)
- [Marking](#marking)
- [Motifs](#motifs)
- [Memberships](#memberships)
@@ -286,146 +280,6 @@ Split data can be rejoined using the `from_*()` family of functions.
See also `r list_functions("^to_")` and `r list_functions("^from_")`.
-## Mapping
-
-`{manynet}` includes three one-line graphing functions with sensible defaults based on
-the network's properties.
-
-### Graphing
-
-First, `graphr()` is used to graph networks in any of the `{manynet}` formats.
-It includes sensible defaults so that researchers can view their network's structure
-or distribution quickly with a minimum of fuss.
-Compare the output from `{manynet}` with a similar default from `{igraph}`:
-
-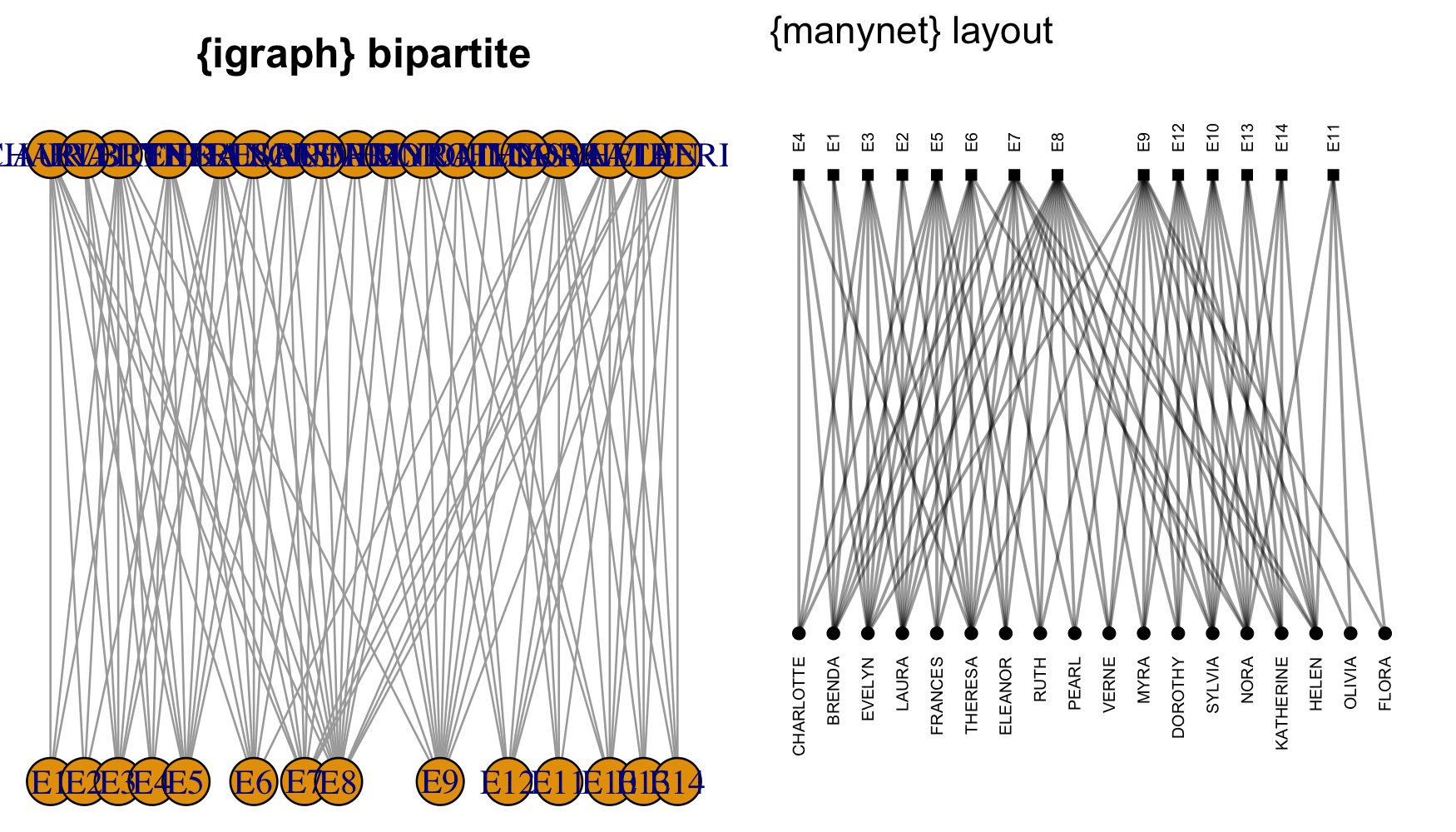 -
-```{r layout-comparison, echo = FALSE, message=FALSE, dpi = 250, fig.height=4, eval = FALSE}
-library(manynet)
-library(igraph)
-library(gridBase)
-library(grid)
-
-par(mfrow=c(1, 2), mai = c(0,0,0.5,0))
-plot(as_igraph(ison_southern_women), layout = layout_as_bipartite, main = "{igraph} bipartite")
-## the last one is the current plot
-plot.new() ## suggested by @Josh
-vps <- baseViewports()
-pushViewport(vps$figure) ## I am in the space of the autocorrelation plot
-vp1 <-plotViewport(c(1.8,1,0,1)) ## create new vp with margins, you play with this values
-p <- graphr(ison_southern_women) + ggtitle("{manynet} twomode")
-print(p,vp = vp1)
-```
-
-Here the `{manynet}` function recognises that the network is a two-mode network
-and uses a bipartite layout by default,
-and recognises that the network contains names for the nodes and
-prints them vertically so that they are legible in this layout.
-Other 'clever' features include automatic node sizing and more.
-By contrast, `{igraph}` requires the bipartite layout to be specified, has cumbersome node size defaults for all but the smallest graphs, and labels also very often need resizing and adjustment to avoid overlap.
-All of `{manynet}`'s adjustments can be overridden, however...
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as
-specifying the function's relevant argument with a replacement,
-or indicating from which attribute it should inherit this information.
-
-
-
-```{r layout-comparison, echo = FALSE, message=FALSE, dpi = 250, fig.height=4, eval = FALSE}
-library(manynet)
-library(igraph)
-library(gridBase)
-library(grid)
-
-par(mfrow=c(1, 2), mai = c(0,0,0.5,0))
-plot(as_igraph(ison_southern_women), layout = layout_as_bipartite, main = "{igraph} bipartite")
-## the last one is the current plot
-plot.new() ## suggested by @Josh
-vps <- baseViewports()
-pushViewport(vps$figure) ## I am in the space of the autocorrelation plot
-vp1 <-plotViewport(c(1.8,1,0,1)) ## create new vp with margins, you play with this values
-p <- graphr(ison_southern_women) + ggtitle("{manynet} twomode")
-print(p,vp = vp1)
-```
-
-Here the `{manynet}` function recognises that the network is a two-mode network
-and uses a bipartite layout by default,
-and recognises that the network contains names for the nodes and
-prints them vertically so that they are legible in this layout.
-Other 'clever' features include automatic node sizing and more.
-By contrast, `{igraph}` requires the bipartite layout to be specified, has cumbersome node size defaults for all but the smallest graphs, and labels also very often need resizing and adjustment to avoid overlap.
-All of `{manynet}`'s adjustments can be overridden, however...
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as
-specifying the function's relevant argument with a replacement,
-or indicating from which attribute it should inherit this information.
-
-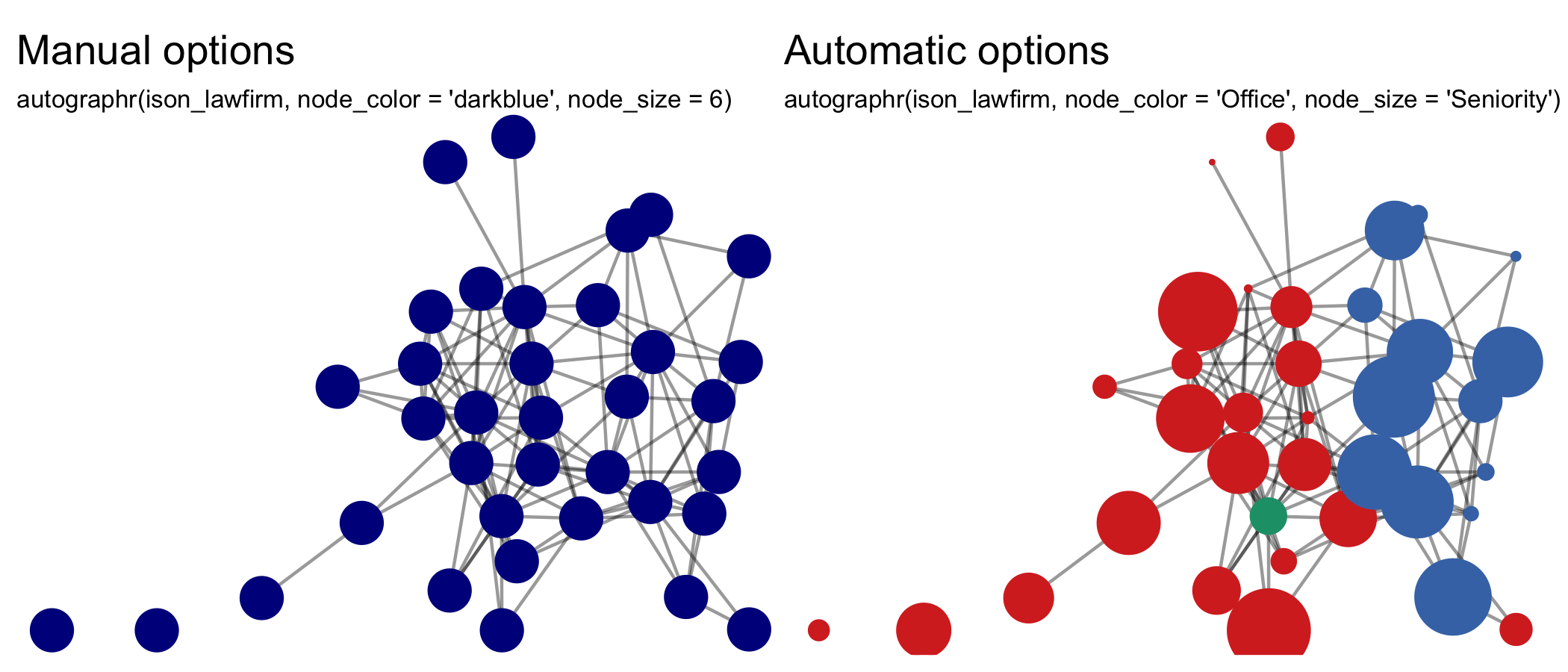 -
-```{r more-options, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-graphr(ison_lawfirm, node_color = "darkblue", node_size = 6) +
- ggtitle("Manual options",
- subtitle = "graphr(ison_lawfirm, node_color = 'darkblue', node_size = 6)") +
-graphr(mutate(ison_lawfirm, Seniority = Seniority/3), node_color = "Office", node_size = "Seniority") +
- ggtitle("Automatic options",
- subtitle = "graphr(ison_lawfirm, node_color = 'Office', node_size = 'Seniority')") &
- theme(plot.subtitle = element_text(size = 8))
-```
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such as `{igraph}`, `{ggraph}`, and `{graphlayouts}`,
-and offers some additional layout algorithms for
-snapping layouts to a grid,
-visualising partitions horizontally, vertically, or concentrically,
-or conforming to configurational coordinates.
-
-
-
-```{r more-options, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-graphr(ison_lawfirm, node_color = "darkblue", node_size = 6) +
- ggtitle("Manual options",
- subtitle = "graphr(ison_lawfirm, node_color = 'darkblue', node_size = 6)") +
-graphr(mutate(ison_lawfirm, Seniority = Seniority/3), node_color = "Office", node_size = "Seniority") +
- ggtitle("Automatic options",
- subtitle = "graphr(ison_lawfirm, node_color = 'Office', node_size = 'Seniority')") &
- theme(plot.subtitle = element_text(size = 8))
-```
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such as `{igraph}`, `{ggraph}`, and `{graphlayouts}`,
-and offers some additional layout algorithms for
-snapping layouts to a grid,
-visualising partitions horizontally, vertically, or concentrically,
-or conforming to configurational coordinates.
-
-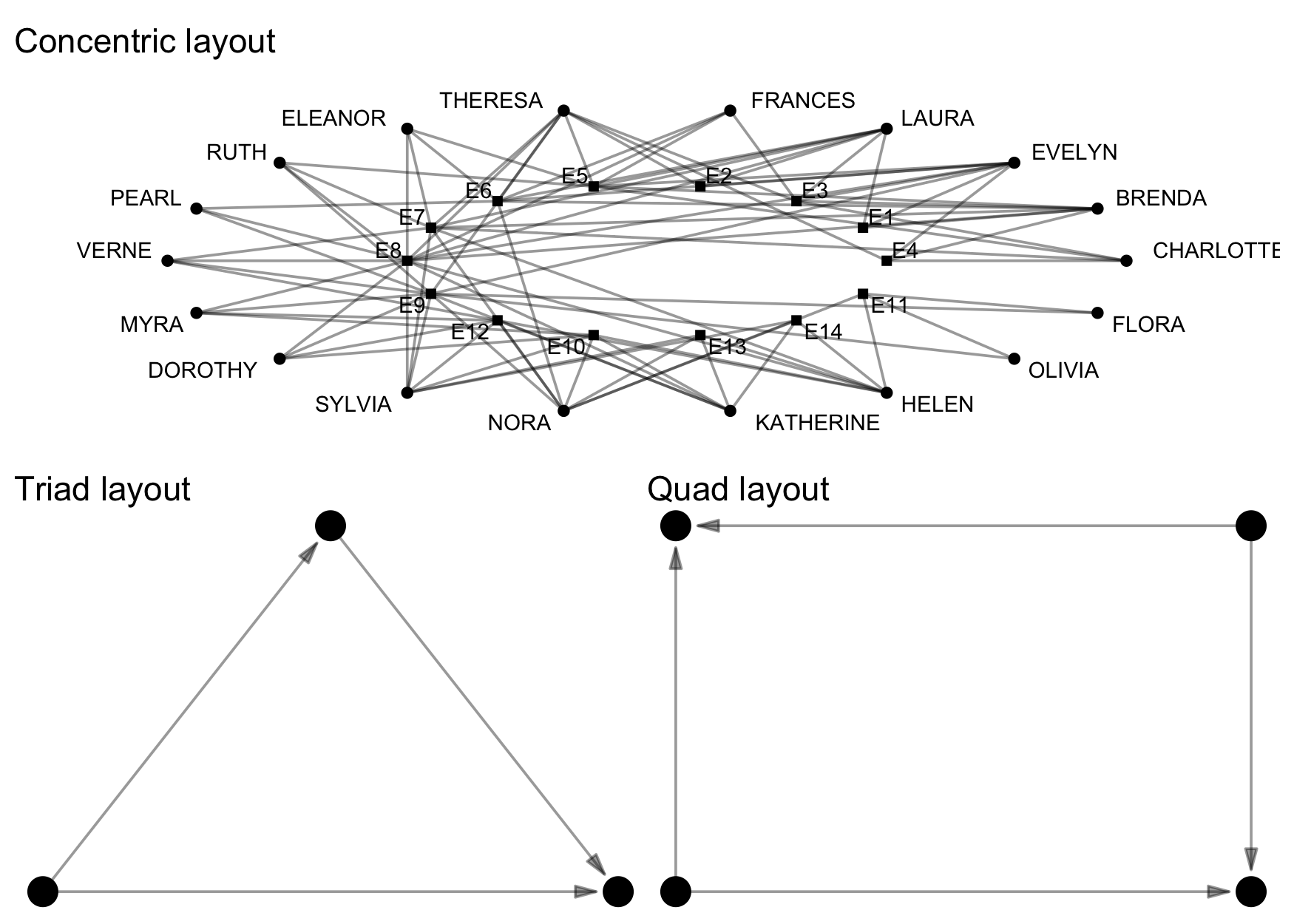 -
-```{r more-layouts, echo = FALSE, message=FALSE, dpi = 250, eval=FALSE}
-(graphr(ison_southern_women, layout = "concentric") + ggtitle("Concentric layout")) /
- ((graphr(to_unnamed(create_explicit(A-+B-+C, A-+C))) + ggtitle("Triad layout")) |
- (graphr(to_unnamed(create_explicit(A-+C, A-+D, B-+C, B-+D))) + ggtitle("Quad layout")))
-```
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of your plots.
-`{manynet}` uses the excellent `{ggraph}` package (and thus `{ggplot2}`)
-as a plotting engine.
-This enables alterations such as the application of themes to be applied upon the defaults.
-If you want to quickly make sure your plots conform to your institution or taste,
-then it is easy to do with themes and scales that update the basic look and color palette
-used in your plots.
-
-
-
-```{r more-layouts, echo = FALSE, message=FALSE, dpi = 250, eval=FALSE}
-(graphr(ison_southern_women, layout = "concentric") + ggtitle("Concentric layout")) /
- ((graphr(to_unnamed(create_explicit(A-+B-+C, A-+C))) + ggtitle("Triad layout")) |
- (graphr(to_unnamed(create_explicit(A-+C, A-+D, B-+C, B-+D))) + ggtitle("Quad layout")))
-```
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of your plots.
-`{manynet}` uses the excellent `{ggraph}` package (and thus `{ggplot2}`)
-as a plotting engine.
-This enables alterations such as the application of themes to be applied upon the defaults.
-If you want to quickly make sure your plots conform to your institution or taste,
-then it is easy to do with themes and scales that update the basic look and color palette
-used in your plots.
-
-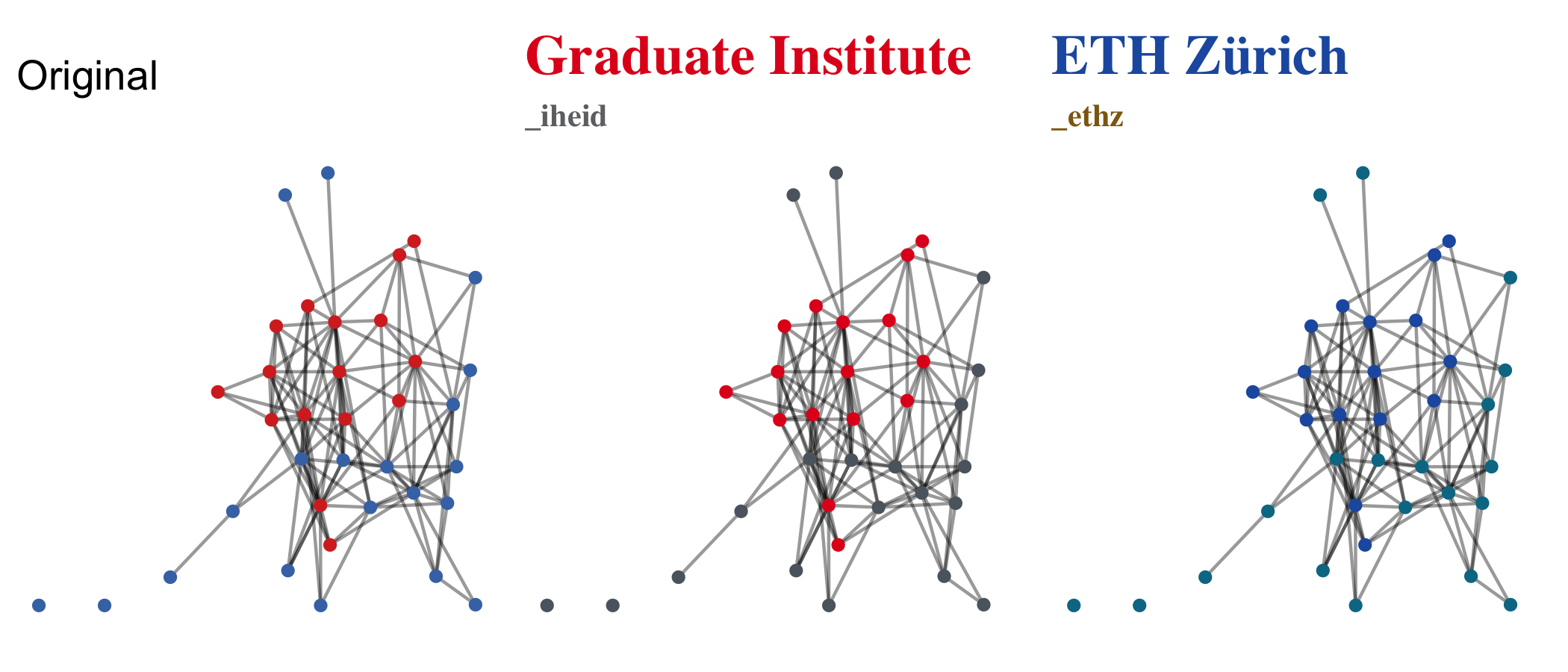 -
-```{r more-themes, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-p <- graphr(ison_lawfirm, node_color = "Practice") +
- ggtitle("Original")
-p + p + theme_iheid() + scale_color_iheid() +
- ggtitle("Graduate Institute", subtitle = "_iheid") +
- p + theme_ethz() + scale_color_ethz() +
- ggtitle("ETH Zürich", subtitle = "_ethz") & ggplot2::theme(legend.position = "none")
-```
-
-More themes are on their way, and we're happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together,
-which can be useful for ego networks or network panels.
-`{patchwork}` is used to help arrange individual plots together.
-
-
-
-```{r more-themes, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-p <- graphr(ison_lawfirm, node_color = "Practice") +
- ggtitle("Original")
-p + p + theme_iheid() + scale_color_iheid() +
- ggtitle("Graduate Institute", subtitle = "_iheid") +
- p + theme_ethz() + scale_color_ethz() +
- ggtitle("ETH Zürich", subtitle = "_ethz") & ggplot2::theme(legend.position = "none")
-```
-
-More themes are on their way, and we're happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together,
-which can be useful for ego networks or network panels.
-`{patchwork}` is used to help arrange individual plots together.
-
-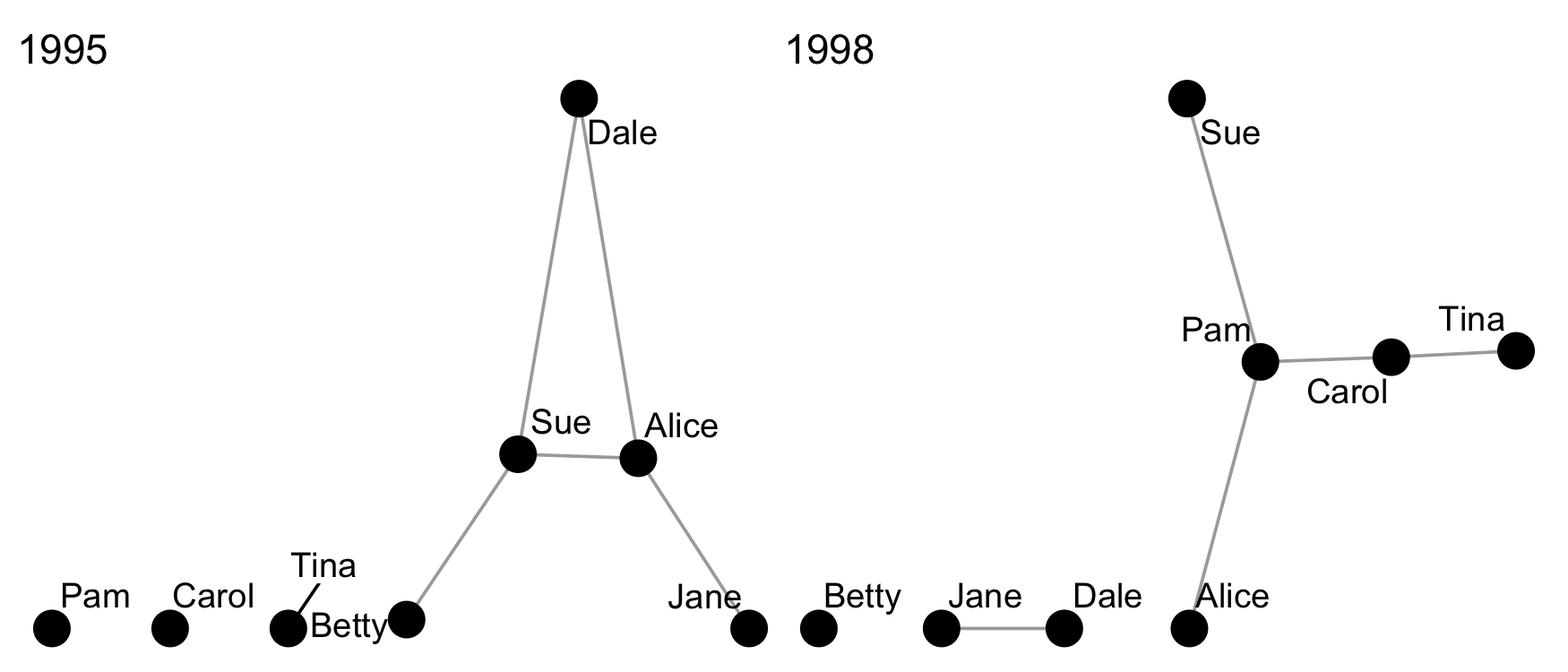 -
-```{r autographs, echo = FALSE, dpi = 250, fig.height=3, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(wave = c(rep(1995, 5), rep(1998, 5))) %>%
- to_waves(attribute = "wave", panels = c(1995, 1998)) %>%
- graphs()
-```
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks.
-It uses `{gganimate}` and `{gifski}` to create a gif that
-visualises network changes over time.
-It really couldn't be easier.
-
-
-
-```{r autographs, echo = FALSE, dpi = 250, fig.height=3, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(wave = c(rep(1995, 5), rep(1998, 5))) %>%
- to_waves(attribute = "wave", panels = c(1995, 1998)) %>%
- graphs()
-```
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks.
-It uses `{gganimate}` and `{gifski}` to create a gif that
-visualises network changes over time.
-It really couldn't be easier.
-
-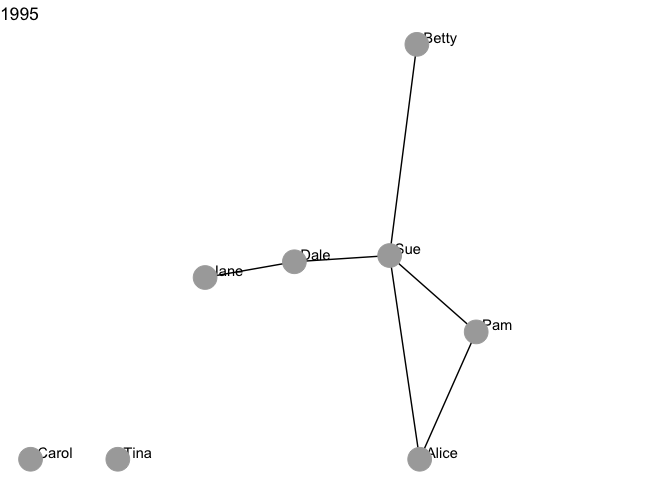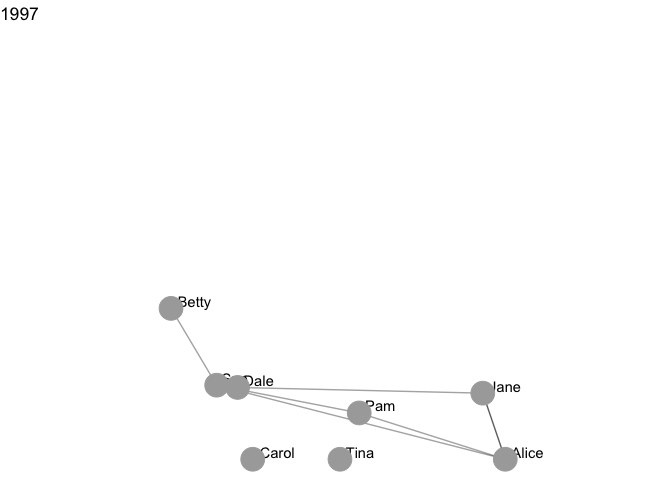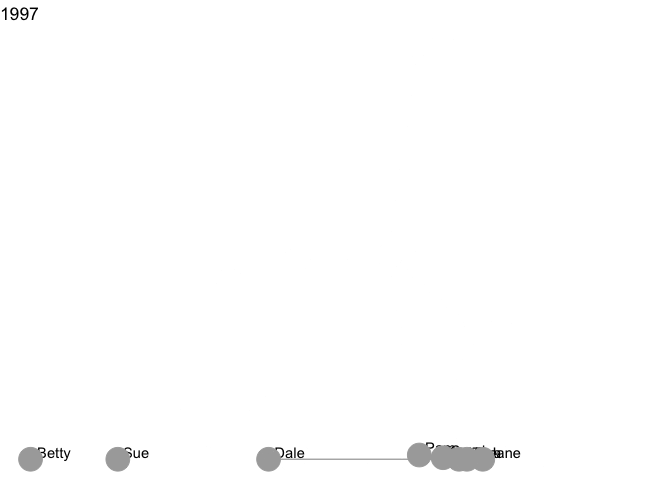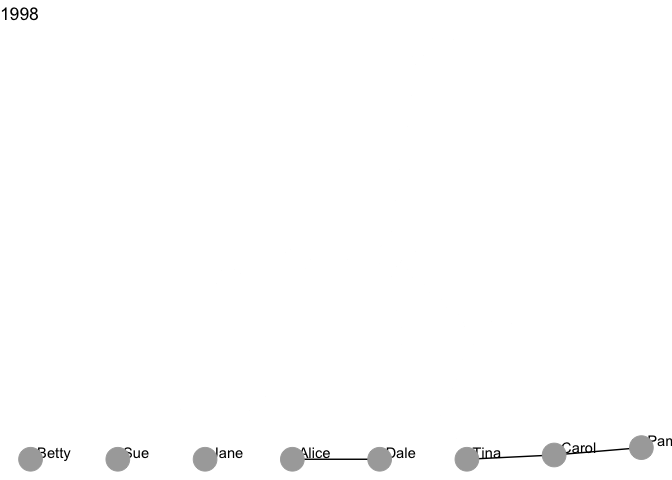 -
-```{r autographd, echo = FALSE, dpi = 250, fig.height=3.5, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) %>%
- to_waves(attribute = "year") %>%
- grapht()
-```
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions,
diff --git a/README.md b/README.md
index a41d6f4a..68a18c7b 100644
--- a/README.md
+++ b/README.md
@@ -38,9 +38,11 @@ make learning and using network analysis tools in R challenging.
By contrast, `{manynet}` offers *many* analytic tools that work on
*many* (if not most) types and kinds of networks. It helps researchers
-make, modify, map, mark, measure, and identify nodes’ motifs and
-memberships in networks. For further testing and modelling capabilities,
-see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
+make, modify, mark, measure, and identify nodes’ motifs and memberships
+in networks. For graph drawing see
+[`{autograph}`](https://stocnet.github.io/autograph/), and for further
+testing and modelling capabilities see
+[`{migraph}`](https://stocnet.github.io/migraph/) and the other
[stocnet](https://github.com/stocnet) packages.
- [Making](#making)
@@ -53,13 +55,6 @@ see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
- [Transforming](#transforming)
- [Splitting and Joining](#splitting-and-joining)
- [Extracting](#extracting)
-- [Mapping](#mapping)
- - [Graphing](#graphing)
- - [More options](#more-options)
- - [More layouts](#more-layouts)
- - [More themes and scales](#more-themes-and-scales)
- - [graphs](#graphs)
- - [grapht](#grapht)
- [Marking](#marking)
- [Motifs](#motifs)
- [Memberships](#memberships)
@@ -237,99 +232,6 @@ See also `to_acyclic()`, `to_anti()`, `to_blocks()`, `to_components()`,
`to_unweighted()`, `to_waves()`, `to_weighted()` and `from_egos()`,
`from_slices()`, `from_subgraphs()`, `from_ties()`, `from_waves()`.
-## Mapping
-
-`{manynet}` includes three one-line graphing functions with sensible
-defaults based on the network’s properties.
-
-### Graphing
-
-First, `graphr()` is used to graph networks in any of the `{manynet}`
-formats. It includes sensible defaults so that researchers can view
-their network’s structure or distribution quickly with a minimum of
-fuss. Compare the output from `{manynet}` with a similar default from
-`{igraph}`:
-
-
-
-```{r autographd, echo = FALSE, dpi = 250, fig.height=3.5, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) %>%
- to_waves(attribute = "year") %>%
- grapht()
-```
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions,
diff --git a/README.md b/README.md
index a41d6f4a..68a18c7b 100644
--- a/README.md
+++ b/README.md
@@ -38,9 +38,11 @@ make learning and using network analysis tools in R challenging.
By contrast, `{manynet}` offers *many* analytic tools that work on
*many* (if not most) types and kinds of networks. It helps researchers
-make, modify, map, mark, measure, and identify nodes’ motifs and
-memberships in networks. For further testing and modelling capabilities,
-see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
+make, modify, mark, measure, and identify nodes’ motifs and memberships
+in networks. For graph drawing see
+[`{autograph}`](https://stocnet.github.io/autograph/), and for further
+testing and modelling capabilities see
+[`{migraph}`](https://stocnet.github.io/migraph/) and the other
[stocnet](https://github.com/stocnet) packages.
- [Making](#making)
@@ -53,13 +55,6 @@ see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
- [Transforming](#transforming)
- [Splitting and Joining](#splitting-and-joining)
- [Extracting](#extracting)
-- [Mapping](#mapping)
- - [Graphing](#graphing)
- - [More options](#more-options)
- - [More layouts](#more-layouts)
- - [More themes and scales](#more-themes-and-scales)
- - [graphs](#graphs)
- - [grapht](#grapht)
- [Marking](#marking)
- [Motifs](#motifs)
- [Memberships](#memberships)
@@ -237,99 +232,6 @@ See also `to_acyclic()`, `to_anti()`, `to_blocks()`, `to_components()`,
`to_unweighted()`, `to_waves()`, `to_weighted()` and `from_egos()`,
`from_slices()`, `from_subgraphs()`, `from_ties()`, `from_waves()`.
-## Mapping
-
-`{manynet}` includes three one-line graphing functions with sensible
-defaults based on the network’s properties.
-
-### Graphing
-
-First, `graphr()` is used to graph networks in any of the `{manynet}`
-formats. It includes sensible defaults so that researchers can view
-their network’s structure or distribution quickly with a minimum of
-fuss. Compare the output from `{manynet}` with a similar default from
-`{igraph}`:
-
- -
-Here the `{manynet}` function recognises that the network is a two-mode
-network and uses a bipartite layout by default, and recognises that the
-network contains names for the nodes and prints them vertically so that
-they are legible in this layout. Other ‘clever’ features include
-automatic node sizing and more. By contrast, `{igraph}` requires the
-bipartite layout to be specified, has cumbersome node size defaults for
-all but the smallest graphs, and labels also very often need resizing
-and adjustment to avoid overlap. All of `{manynet}`’s adjustments can be
-overridden, however…
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as specifying
-the function’s relevant argument with a replacement, or indicating from
-which attribute it should inherit this information.
-
-
-
-Here the `{manynet}` function recognises that the network is a two-mode
-network and uses a bipartite layout by default, and recognises that the
-network contains names for the nodes and prints them vertically so that
-they are legible in this layout. Other ‘clever’ features include
-automatic node sizing and more. By contrast, `{igraph}` requires the
-bipartite layout to be specified, has cumbersome node size defaults for
-all but the smallest graphs, and labels also very often need resizing
-and adjustment to avoid overlap. All of `{manynet}`’s adjustments can be
-overridden, however…
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as specifying
-the function’s relevant argument with a replacement, or indicating from
-which attribute it should inherit this information.
-
- -
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such
-as `{igraph}`, `{ggraph}`, and `{graphlayouts}`, and offers some
-additional layout algorithms for snapping layouts to a grid, visualising
-partitions horizontally, vertically, or concentrically, or conforming to
-configurational coordinates.
-
-
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such
-as `{igraph}`, `{ggraph}`, and `{graphlayouts}`, and offers some
-additional layout algorithms for snapping layouts to a grid, visualising
-partitions horizontally, vertically, or concentrically, or conforming to
-configurational coordinates.
-
- -
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of
-your plots. `{manynet}` uses the excellent `{ggraph}` package (and thus
-`{ggplot2}`) as a plotting engine. This enables alterations such as the
-application of themes to be applied upon the defaults. If you want to
-quickly make sure your plots conform to your institution or taste, then
-it is easy to do with themes and scales that update the basic look and
-color palette used in your plots.
-
-
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of
-your plots. `{manynet}` uses the excellent `{ggraph}` package (and thus
-`{ggplot2}`) as a plotting engine. This enables alterations such as the
-application of themes to be applied upon the defaults. If you want to
-quickly make sure your plots conform to your institution or taste, then
-it is easy to do with themes and scales that update the basic look and
-color palette used in your plots.
-
- -
-More themes are on their way, and we’re happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together, which
-can be useful for ego networks or network panels. `{patchwork}` is used
-to help arrange individual plots together.
-
-
-
-More themes are on their way, and we’re happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together, which
-can be useful for ego networks or network panels. `{patchwork}` is used
-to help arrange individual plots together.
-
- -
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks. It uses
-`{gganimate}` and `{gifski}` to create a gif that visualises network
-changes over time. It really couldn’t be easier.
-
-
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks. It uses
-`{gganimate}` and `{gifski}` to create a gif that visualises network
-changes over time. It really couldn’t be easier.
-
- -
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions, each with their
diff --git a/cran-comments.md b/cran-comments.md
index 2827d039..634afb8c 100644
--- a/cran-comments.md
+++ b/cran-comments.md
@@ -9,4 +9,4 @@
0 errors | 0 warnings | 0 notes
-- Removed failing URLs from previous submission (v1.5.0)
\ No newline at end of file
+- Required for ggplot2 v4.0.0
\ No newline at end of file
diff --git a/man/defunct.Rd b/man/defunct.Rd
index 7258166b..1a5285e1 100644
--- a/man/defunct.Rd
+++ b/man/defunct.Rd
@@ -4,9 +4,6 @@
\alias{defunct}
\alias{pkg_data}
\alias{node_mode}
-\alias{autographr}
-\alias{autographs}
-\alias{autographd}
\alias{node_optimal}
\alias{node_kernighanlin}
\alias{node_edge_betweenness}
@@ -89,7 +86,6 @@
\alias{node_core}
\alias{node_by_quad}
\alias{net_by_quad}
-\alias{layout_tbl_graph_quad}
\alias{net_hazard}
\title{Functions that have been renamed, superseded, or are no longer working}
\usage{
@@ -97,34 +93,6 @@ pkg_data(pkg = "manynet")
node_mode(.data)
-autographr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- ...
-)
-
-autographs(netlist, waves, based_on = c("first", "last", "both"), ...)
-
-autographd(
- tlist,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- keep_isolates = TRUE,
- ...
-)
-
node_optimal(.data)
node_kernighanlin(.data)
@@ -295,8 +263,6 @@ node_by_quad(.data)
net_by_quad(.data)
-layout_tbl_graph_quad(.data, circular = FALSE, times = 1000)
-
net_hazard(.data)
}
\description{
@@ -314,12 +280,6 @@ wherever possible and update your scripts accordingly.
\item \code{node_mode()}: Deprecated on 2024-06-17.
-\item \code{autographr()}: Deprecated on 2024-06-17.
-
-\item \code{autographs()}: Deprecated on 2024-06-17.
-
-\item \code{autographd()}: Deprecated on 2024-06-17.
-
\item \code{node_optimal()}: Deprecated on 2024-06-14.
\item \code{node_kernighanlin()}: Deprecated on 2024-06-14.
@@ -486,8 +446,6 @@ wherever possible and update your scripts accordingly.
\item \code{net_by_quad()}: Deprecated on 2024-10-10.
-\item \code{layout_tbl_graph_quad()}: Deprecated on 2024-10-10.
-
\item \code{net_hazard()}: Deprecated on 2024-12-30.
}}
diff --git a/man/depth_first_recursive_search.Rd b/man/depth_first_recursive_search.Rd
deleted file mode 100644
index 9642a068..00000000
--- a/man/depth_first_recursive_search.Rd
+++ /dev/null
@@ -1,20 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{depth_first_recursive_search}
-\alias{depth_first_recursive_search}
-\title{Layouts for snapping layouts to a grid}
-\usage{
-depth_first_recursive_search(layout)
-}
-\description{
-The function uses approximate pattern matching
-to redistribute coarse layouts on square grid points, while
-preserving the topological relationships among the nodes (see Inoue et al. 2012).
-}
-\references{
-Inoue, Kentaro, Shinichi Shimozono, Hideaki Yoshida, and Hiroyuki Kurata. 2012.
-“Application of Approximate Pattern Matching in Two Dimensional Spaces to Grid Layout for Biochemical Network Maps” edited by J. Bourdon.
-\emph{PLoS ONE} 7(6):e37739.
-\doi{https://doi.org/10.1371/journal.pone.0037739}.
-}
-\keyword{internal}
diff --git a/man/make_cran.Rd b/man/make_cran.Rd
index b49d70b3..16557e6d 100644
--- a/man/make_cran.Rd
+++ b/man/make_cran.Rd
@@ -37,8 +37,6 @@ by \href{https://github.com/stocnet/manynet/issues}{raising an issue on Github}.
\examples{
# mnet <- read_cran()
# mnet <- to_ego(mnet, "manynet", max_dist = 2)
-# graphr(mnet, layout = "hierarchy",
-# edge_color = "type", node_color = "Compilation")
}
\seealso{
\link{as}
diff --git a/man/make_learning.Rd b/man/make_learning.Rd
index b06808d4..6f08083a 100644
--- a/man/make_learning.Rd
+++ b/man/make_learning.Rd
@@ -87,9 +87,6 @@ from other nodes whose beliefs are not too dissimilar from their own.
latticeEg <- add_node_attribute(latticeEg, "startValues", startValues)
latticeEg
play_segregation(latticeEg, "startValues", 0.5)
- # graphr(latticeEg, node_color = "startValues", node_size = 5) +
- # graphr(play_segregation(latticeEg, "startValues", 0.2),
- # node_color = "startValues", node_size = 5)
}
\references{
DeGroot, Morris H. 1974.
diff --git a/man/make_play.Rd b/man/make_play.Rd
index fe1b0621..7d1aa8c7 100644
--- a/man/make_play.Rd
+++ b/man/make_play.Rd
@@ -185,8 +185,6 @@ This can be used in in SEI, SEIS, SEIR, and SEIRS models.
\examples{
smeg <- generate_smallworld(15, 0.025)
- plot(play_diffusion(smeg, recovery = 0.4))
- #graphr(play_diffusion(ison_karateka))
}
\seealso{
Other makes:
diff --git a/man/make_random.Rd b/man/make_random.Rd
index 0146a541..e8471363 100644
--- a/man/make_random.Rd
+++ b/man/make_random.Rd
@@ -105,8 +105,8 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_random(12, 0.4))
-# graphr(generate_random(c(6, 6), 0.4))
+generate_random(12, 0.4)
+# generate_random(c(6, 6), 0.4)
}
\references{
\subsection{On random networks}{
diff --git a/man/make_stochastic.Rd b/man/make_stochastic.Rd
index f2930b77..0a977e6b 100644
--- a/man/make_stochastic.Rd
+++ b/man/make_stochastic.Rd
@@ -104,10 +104,10 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_smallworld(12, 0.025))
-graphr(generate_smallworld(12, 0.25))
-graphr(generate_scalefree(12, 0.25))
-graphr(generate_scalefree(12, 1.25))
+generate_smallworld(12, 0.025)
+generate_smallworld(12, 0.25)
+generate_scalefree(12, 0.25)
+generate_scalefree(12, 1.25)
generate_fire(10)
generate_islands(10)
generate_citations(10)
diff --git a/man/manip_paths.Rd b/man/manip_paths.Rd
index f62c282e..ca35715c 100644
--- a/man/manip_paths.Rd
+++ b/man/manip_paths.Rd
@@ -109,10 +109,7 @@ This is, however, computationally slower.
\examples{
to_matching(ison_southern_women)
-#graphr(to_matching(ison_southern_women))
-graphr(to_mentoring(ison_adolescents))
to_eulerian(delete_nodes(ison_koenigsberg, "Lomse"))
- #graphr(to_eulerian(delete_nodes(ison_koenigsberg, "Lomse")))
}
\references{
\subsection{On matching}{
@@ -124,7 +121,7 @@ Gale, David, and Lloyd Stowell Shapley. 1962.
Goldberg, Andrew V., and Robert E. Tarjan. 1986.
"A new approach to the maximum flow problem".
-\emph{Proceedings of the eighteenth annual ACM symposium on Theory of computing – STOC '86}.
+\emph{Proceedings of the 18th Annual ACM Symposium on Theory of Computing}.
136-146.
\doi{10.1145/12130.12144}
}
diff --git a/man/manip_permutation.Rd b/man/manip_permutation.Rd
index 0578c343..32a6df8b 100644
--- a/man/manip_permutation.Rd
+++ b/man/manip_permutation.Rd
@@ -26,10 +26,6 @@ By default TRUE.}
on both the rows and columns (for a one-mode network)
or on each of the rows and columns (for a two-mode network).
}
-\examples{
-graphr(ison_adolescents, node_size = 4)
-graphr(to_permuted(ison_adolescents), node_size = 4)
-}
\seealso{
Other modifications:
\code{\link{manip_as}},
diff --git a/man/manip_project.Rd b/man/manip_project.Rd
index 0b59bb28..04f6e5ec 100644
--- a/man/manip_project.Rd
+++ b/man/manip_project.Rd
@@ -71,10 +71,7 @@ Below are the currently implemented S3 methods:\tabular{lrrrrr}{
\examples{
to_mode1(ison_southern_women)
to_mode2(ison_southern_women)
-#graphr(to_mode1(ison_southern_women))
-#graphr(to_mode2(ison_southern_women))
to_ties(ison_adolescents)
-#graphr(to_ties(ison_adolescents))
}
\seealso{
Other modifications:
diff --git a/man/manip_reformat.Rd b/man/manip_reformat.Rd
index 464f1716..17185089 100644
--- a/man/manip_reformat.Rd
+++ b/man/manip_reformat.Rd
@@ -70,7 +70,6 @@ This essentially has no effect on undirected networks or reciprocated ties.
}}
\examples{
to_anti(ison_southern_women)
-#graphr(to_anti(ison_southern_women))
}
\seealso{
Other modifications:
diff --git a/man/manip_ties.Rd b/man/manip_ties.Rd
index 531efcc0..c53695bf 100644
--- a/man/manip_ties.Rd
+++ b/man/manip_ties.Rd
@@ -83,7 +83,7 @@ Note that while \verb{add_*()}/\verb{delete_*()} functions operate similarly as
other <- create_filled(4) \%>\% mutate(name = c("A", "B", "C", "D"))
mutate_ties(other, form = 1:6) \%>\% filter_ties(form < 4)
add_tie_attribute(other, "weight", c(1, 2, 2, 2, 1, 2))
-ison_adolescents \%>\% add_ties(c("Betty","Tina")) \%>\% graphr()
+ison_adolescents \%>\% add_ties(c("Betty","Tina"))
delete_ties(ison_adolescents, 3)
delete_ties(ison_adolescents, "Alice|Sue")
}
diff --git a/man/map_graphr.Rd b/man/map_graphr.Rd
deleted file mode 100644
index 474f3d4b..00000000
--- a/man/map_graphr.Rd
+++ /dev/null
@@ -1,141 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphr}
-\alias{map_graphr}
-\alias{graphr}
-\title{Easily graph networks with sensible defaults}
-\usage{
-graphr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- snap = FALSE,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{.data}{A manynet-consistent object.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{node_group}{Node variable to be used for grouping the nodes.
-It is easiest if this is added as a hull over
-groups before plotting.
-Group variables should have a minimum of 3 nodes,
-if less, number groups will be reduced by
-merging categories with lower counts into one called "other".}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{snap}{Logical scalar, whether the layout should be snapped to a grid.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-A \code{ggplot2::ggplot()} object.
-The last plot can be saved to the file system using \code{ggplot2::ggsave()}.
-}
-\description{
-This function provides users with an easy way to graph
-(m)any network data for exploration, investigation, inspiration,
-and communication.
-
-It builds upon \code{{ggplot2}} and \code{{ggraph}} to offer
-pretty and extensible graphing solutions.
-However, compared to those solutions,
-\code{graphr()} contains various algorithms to provide better looking
-graphs by default.
-This means that just passing the function some network data
-will often be sufficient to return a reasonable-looking graph.
-
-The function also makes it easy to modify many of the most
-commonly adapted aspects of a graph,
-including node and edge size, colour, and shape,
-as arguments rather than additional functions that you need to remember.
-These can be defined outright, e.g. \code{node_size = 8}, or
-in reference to an attribute of the network, e.g. \code{node_size = "wealth"}.
-
-Lastly, \code{graphr()} uses \code{{ggplot2}}-related theme information, so
-it is easy to make colour palette and fonts institution-specific and consistent.
-See e.g. \code{theme_iheid()} for more.
-
-To learn more about what can be done visually,
-try \code{run_tute("Visualisation")}.
-}
-\examples{
-graphr(ison_adolescents)
-ison_adolescents \%>\%
- mutate(color = rep(c("introvert","extrovert"), times = 4),
- size = ifelse(node_is_cutpoint(ison_adolescents), 6, 3)) \%>\%
- mutate_ties(ecolor = rep(c("friends", "acquaintances"), times = 5)) \%>\%
- graphr(node_color = "color", node_size = "size",
- edge_size = 1.5, edge_color = "ecolor")
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_graphs.Rd b/man/map_graphs.Rd
deleted file mode 100644
index 549d1128..00000000
--- a/man/map_graphs.Rd
+++ /dev/null
@@ -1,60 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphs}
-\alias{map_graphs}
-\alias{graphs}
-\title{Easily graph a set of networks with sensible defaults}
-\usage{
-graphs(netlist, waves, based_on = c("first", "last", "both"), ...)
-}
-\arguments{
-\item{netlist}{A list of manynet-compatible networks.}
-
-\item{waves}{Numeric, the number of plots to be displayed side-by-side.
-If missing, the number of plots will be reduced to the first and last
-when there are more than four plots.
-This argument can also be passed a vector selecting the waves to plot.}
-
-\item{based_on}{Whether the layout of the joint plots should
-be based on the "first" or the "last" network, or "both".}
-
-\item{...}{Additional arguments passed to \code{graphr()}.}
-}
-\value{
-Multiple \code{ggplot2::ggplot()} objects displayed side-by-side.
-}
-\description{
-This function provides users with an easy way to graph
-lists of network data for comparison.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{patchwork}} package to plot the graphs
-side by side and, if necessary, in successive rows.
-This is useful for lists of networks that represent, for example,
-ego or component subgraphs of a network,
-or a list of a network's different types of tie or across time.
-By default just the first and last network will be plotted,
-but this can be overridden by the "waves" parameter.
-
-Where the graphs are of the same network (same nodes),
-the graphs may share a layout to facilitate comparison.
-By default, successive graphs will use the layout calculated for
-the "first" network, but other options include the "last" layout,
-or a mix, "both", of them.
-}
-\examples{
-#graphs(to_egos(ison_adolescents))
-#graphs(to_egos(ison_adolescents), waves = 8)
-#graphs(to_egos(ison_adolescents), waves = c(2, 4, 6))
-#graphs(play_diffusion(ison_adolescents))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_grapht.Rd b/man/map_grapht.Rd
deleted file mode 100644
index ff0cce5a..00000000
--- a/man/map_grapht.Rd
+++ /dev/null
@@ -1,137 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_grapht}
-\alias{map_grapht}
-\alias{grapht}
-\title{Easily animate dynamic networks with sensible defaults}
-\source{
-https://blog.schochastics.net/posts/2021-09-15_animating-network-evolutions-with-gganimate/
-}
-\usage{
-grapht(
- tlist,
- keep_isolates = TRUE,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{tlist}{The same migraph-compatible network listed according to
-a time attribute, waves, or slices.}
-
-\item{keep_isolates}{Logical, whether to keep isolate nodes in the graph.
-TRUE by default.
-If FALSE, removes nodes from each frame they are isolated in.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-Shows a .gif image. Assigning the result of the function
-saves the gif to a temporary folder and the object holds the path to this file.
-}
-\description{
-This function provides users with an easy way to graph
-dynamic network data for exploration and presentation.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{gganimate}} package to animate the changes
-between successive iterations of a network.
-This is useful for networks in which the ties and/or the node or tie
-attributes are changing.
-
-A progress bar is shown if it takes some time to encoding all the
-.png files into a .gif.
-}
-\examples{
-#ison_adolescents \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year", cumulative = TRUE) \%>\%
-# grapht()
-#ison_adolescents \%>\%
-# mutate(gender = rep(c("male", "female"), times = 4),
-# hair = rep(c("black", "brown"), times = 4),
-# age = sample(11:16, 8, replace = TRUE)) \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE),
-# links = sample(c("friends", "not_friends"), 10, replace = TRUE),
-# weekly_meetings = sample(c(3, 5, 7), 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year") \%>\%
-# grapht(layout = "concentric", membership = "gender",
-# node_shape = "gender", node_color = "hair",
-# node_size = "age", edge_color = "links",
-# edge_size = "weekly_meetings")
-#grapht(play_diffusion(ison_adolescents, seeds = 5))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_configuration.Rd b/man/map_layout_configuration.Rd
deleted file mode 100644
index 8f8e0f33..00000000
--- a/man/map_layout_configuration.Rd
+++ /dev/null
@@ -1,54 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_configuration}
-\alias{map_layout_configuration}
-\alias{layout_tbl_graph_configuration}
-\alias{layout_tbl_graph_dyad}
-\alias{layout_tbl_graph_triad}
-\alias{layout_tbl_graph_tetrad}
-\alias{layout_tbl_graph_pentad}
-\alias{layout_tbl_graph_hexad}
-\title{Layout algorithms based on configurational positions}
-\usage{
-layout_tbl_graph_configuration(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_dyad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_triad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_tetrad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_pentad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_hexad(.data, circular = FALSE, times = 1000)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-}
-\description{
-Configurational layouts locate nodes at symmetric coordinates
-to help illustrate particular configurations.
-Currently "triad" and "quad" layouts are available.
-The "configuration" layout will choose the appropriate configurational
-layout automatically.
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_partition.Rd b/man/map_layout_partition.Rd
deleted file mode 100644
index c30b378a..00000000
--- a/man/map_layout_partition.Rd
+++ /dev/null
@@ -1,130 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_partition}
-\alias{map_layout_partition}
-\alias{layout_tbl_graph_hierarchy}
-\alias{layout_tbl_graph_alluvial}
-\alias{layout_tbl_graph_railway}
-\alias{layout_tbl_graph_ladder}
-\alias{layout_tbl_graph_concentric}
-\alias{layout_tbl_graph_multilevel}
-\alias{layout_tbl_graph_lineage}
-\title{Layout algorithms based on bi- or other partitions}
-\source{
-Diego Diez, Andrew P. Hutchins and Diego Miranda-Saavedra. 2014.
-"Systematic identification of transcriptional regulatory modules from
-protein-protein interaction networks".
-\emph{Nucleic Acids Research}, 42 (1) e6.
-}
-\usage{
-layout_tbl_graph_hierarchy(
- .data,
- center = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_alluvial(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_railway(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_ladder(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_concentric(
- .data,
- membership,
- radius = NULL,
- order.by = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_multilevel(.data, level, circular = FALSE)
-
-layout_tbl_graph_lineage(.data, rank, circular = FALSE)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{center}{Further split "hierarchical" layouts by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name in hierarchy layout.
-Defaults to NULL.}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-
-\item{membership}{A node attribute or a vector to draw concentric circles
-for "concentric" layout.}
-
-\item{radius}{A vector of radii at which the concentric circles
-should be located for "concentric" layout.
-By default this is equal placement around an empty centre,
-unless one (the core) is a single node,
-in which case this node occupies the centre of the graph.}
-
-\item{order.by}{An attribute label indicating the (decreasing) order
-for the nodes around the circles for "concentric" layout.
-By default ordering is given by a bipartite placement that reduces
-the number of edge crossings.}
-
-\item{level}{A node attribute or a vector to hierarchically order levels for
-"multilevel" layout.}
-
-\item{rank}{A numerical node attribute to place nodes in Y axis
-according to values for "lineage" layout.}
-}
-\description{
-These algorithms layout networks based on two or more partitions,
-and are recommended for use with \code{graphr()} or \code{{ggraph}}.
-Note that these layout algorithms use \code{{Rgraphviz}},
-a package that is only available on Bioconductor.
-It will first need to be downloaded using \code{BiocManager::install("Rgraphviz")}.
-If it has not already been installed, there is a prompt the first time
-these functions are used though.
-
-The "hierarchy" layout layers the first node set along the bottom,
-and the second node set along the top,
-sequenced and spaced as necessary to minimise edge overlap.
-The "alluvial" layout is similar to "hierarchy",
-but places successive layers horizontally rather than vertically.
-The "railway" layout is similar to "hierarchy",
-but nodes are aligned across the layers.
-The "ladder" layout is similar to "railway",
-but places successive layers horizontally rather than vertically.
-The "concentric" layout places a "hierarchy" layout
-around a circle, with successive layers appearing as concentric circles.
-The "multilevel" layout places successive layers as multiple levels.
-The "lineage" layout ranks nodes in Y axis according to values.
-}
-\examples{
-#graphr(ison_southern_women, layout = "hierarchy", center = "events",
-# node_color = "type", node_size = 3)
-#graphr(ison_southern_women, layout = "alluvial")
-#graphr(ison_southern_women, layout = "concentric", membership = "type",
-# node_color = "type", node_size = 3)
-#graphr(ison_lotr, layout = "multilevel",
-# node_color = "Race", level = "Race", node_size = 3)
-# ison_adolescents \%>\%
-# mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2),
-# cut = node_is_cutpoint(ison_adolescents)) \%>\%
-# graphr(layout = "lineage", rank = "year", node_color = "cut",
-# node_size = migraph::node_degree(ison_adolescents)*10)
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}}
-}
-\concept{mapping}
diff --git a/man/map_palettes.Rd b/man/map_palettes.Rd
deleted file mode 100644
index 8c086ea9..00000000
--- a/man/map_palettes.Rd
+++ /dev/null
@@ -1,32 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_palettes.R
-\name{map_palettes}
-\alias{map_palettes}
-\alias{many_palettes}
-\title{Many palettes generator}
-\source{
-Adapted from \url{https://github.com/karthik/wesanderson/blob/master/R/colors.R}
-}
-\usage{
-many_palettes(palette, n, type = c("discrete", "continuous"))
-}
-\arguments{
-\item{palette}{Name of desired palette. Current choices are:
-\code{IHEID}, \code{Centres}, \code{SDGs}, \code{ETHZ}, \code{RUG},
-and \code{UZH}.}
-
-\item{n}{Number of colors desired. If omitted, uses all colours.}
-
-\item{type}{Either "continuous" or "discrete". Use continuous if you want
-to automatically interpolate between colours.}
-}
-\value{
-A graphic display of colours in palette.
-}
-\description{
-Many palettes generator
-}
-\examples{
-many_palettes()
-#many_palettes("IHEID")
-}
diff --git a/man/map_scales.Rd b/man/map_scales.Rd
deleted file mode 100644
index bdee1bc6..00000000
--- a/man/map_scales.Rd
+++ /dev/null
@@ -1,112 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_scales}
-\alias{map_scales}
-\alias{scale_fill_iheid}
-\alias{scale_colour_iheid}
-\alias{scale_color_iheid}
-\alias{scale_edge_colour_iheid}
-\alias{scale_edge_color_iheid}
-\alias{scale_fill_centres}
-\alias{scale_colour_centres}
-\alias{scale_color_centres}
-\alias{scale_edge_colour_centres}
-\alias{scale_edge_color_centres}
-\alias{scale_fill_sdgs}
-\alias{scale_colour_sdgs}
-\alias{scale_color_sdgs}
-\alias{scale_edge_colour_sdgs}
-\alias{scale_edge_color_sdgs}
-\alias{scale_fill_ethz}
-\alias{scale_colour_ethz}
-\alias{scale_color_ethz}
-\alias{scale_edge_colour_ethz}
-\alias{scale_edge_color_ethz}
-\alias{scale_fill_uzh}
-\alias{scale_colour_uzh}
-\alias{scale_color_uzh}
-\alias{scale_edge_colour_uzh}
-\alias{scale_edge_color_uzh}
-\alias{scale_fill_rug}
-\alias{scale_colour_rug}
-\alias{scale_color_rug}
-\alias{scale_edge_colour_rug}
-\alias{scale_edge_color_rug}
-\title{Many scales}
-\usage{
-scale_fill_iheid(direction = 1, ...)
-
-scale_colour_iheid(direction = 1, ...)
-
-scale_color_iheid(direction = 1, ...)
-
-scale_edge_colour_iheid(direction = 1, ...)
-
-scale_edge_color_iheid(direction = 1, ...)
-
-scale_fill_centres(direction = 1, ...)
-
-scale_colour_centres(direction = 1, ...)
-
-scale_color_centres(direction = 1, ...)
-
-scale_edge_colour_centres(direction = 1, ...)
-
-scale_edge_color_centres(direction = 1, ...)
-
-scale_fill_sdgs(direction = 1, ...)
-
-scale_colour_sdgs(direction = 1, ...)
-
-scale_color_sdgs(direction = 1, ...)
-
-scale_edge_colour_sdgs(direction = 1, ...)
-
-scale_edge_color_sdgs(direction = 1, ...)
-
-scale_fill_ethz(direction = 1, ...)
-
-scale_colour_ethz(direction = 1, ...)
-
-scale_color_ethz(direction = 1, ...)
-
-scale_edge_colour_ethz(direction = 1, ...)
-
-scale_edge_color_ethz(direction = 1, ...)
-
-scale_fill_uzh(direction = 1, ...)
-
-scale_colour_uzh(direction = 1, ...)
-
-scale_color_uzh(direction = 1, ...)
-
-scale_edge_colour_uzh(direction = 1, ...)
-
-scale_edge_color_uzh(direction = 1, ...)
-
-scale_fill_rug(direction = 1, ...)
-
-scale_colour_rug(direction = 1, ...)
-
-scale_color_rug(direction = 1, ...)
-
-scale_edge_colour_rug(direction = 1, ...)
-
-scale_edge_color_rug(direction = 1, ...)
-}
-\arguments{
-\item{direction}{Direction for using palette colors.}
-
-\item{...}{Extra arguments passed to \code{ggplot2::discrete_scale()}.}
-}
-\description{
-These functions enable to add color scales to be graphs.
-}
-\examples{
-#ison_brandes \%>\%
-#mutate(core = migraph::node_is_core(ison_brandes)) \%>\%
-#graphr(node_color = "core") +
-#scale_color_iheid()
-#graphr(ison_physicians[[1]], edge_color = "type") +
-#scale_edge_color_ethz()
-}
diff --git a/man/map_themes.Rd b/man/map_themes.Rd
deleted file mode 100644
index 288176ee..00000000
--- a/man/map_themes.Rd
+++ /dev/null
@@ -1,41 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_themes}
-\alias{map_themes}
-\alias{set_manynet_theme}
-\alias{theme_iheid}
-\alias{theme_ethz}
-\alias{theme_uzh}
-\alias{theme_rug}
-\title{Many themes}
-\usage{
-set_manynet_theme(theme = "default")
-
-theme_iheid(base_size = 12, base_family = "serif")
-
-theme_ethz(base_size = 12, base_family = "sans")
-
-theme_uzh(base_size = 12, base_family = "sans")
-
-theme_rug(base_size = 12, base_family = "mono")
-}
-\arguments{
-\item{theme}{String naming a theme.
-By default "default".}
-
-\item{base_size}{Font size, by default 12.}
-
-\item{base_family}{Font family, by default "sans".}
-}
-\description{
-These functions enable graphs to be easily and quickly themed,
-e.g. changing the default colour of the graph's vertices and edges.
-}
-\examples{
-to_mentoring(ison_brandes) \%>\%
- mutate(color = c(rep(c(1,2,3), 3), 3)) \%>\%
- graphr(node_color = "color") +
- labs(title = "Who leads and who follows?") +
- scale_color_iheid() +
- theme_iheid()
-}
diff --git a/man/mark_core.Rd b/man/mark_core.Rd
index 6143dee3..7f3c805e 100644
--- a/man/mark_core.Rd
+++ b/man/mark_core.Rd
@@ -4,6 +4,7 @@
\alias{mark_core}
\alias{node_is_universal}
\alias{node_is_core}
+\alias{node_kcoreness}
\alias{node_coreness}
\title{Core-periphery clustering algorithms}
\usage{
@@ -11,6 +12,8 @@ node_is_universal(.data)
node_is_core(.data, method = c("degree", "eigenvector"))
+node_kcoreness(.data)
+
node_coreness(.data)
}
\arguments{
@@ -33,8 +36,13 @@ can be added if there is interest.}
\description{
These functions identify nodes belonging to (some level of) the core of a network:
\itemize{
-\item \code{node_is_core()} assigns nodes to either the core or periphery.
-\item \code{node_coreness()} assigns nodes to their level of k-coreness.
+\item \code{node_is_universal()} identifies whether nodes are adjacent to all other
+nodes in the network.
+\item \code{node_is_core()} identifies whether nodes belong to the core of the
+network, as opposed to the periphery.
+\item \code{node_coreness()} returns a continuous measure of how closely each node
+resembles a typical core node.
+\item \code{node_kcoreness()} assigns nodes to their level of k-coreness.
}
}
\section{Universal/dominating node}{
@@ -68,6 +76,7 @@ node_is_core(ison_adolescents)
#ison_adolescents \%>\%
# mutate(corep = node_is_core()) \%>\%
# graphr(node_color = "corep")
+node_kcoreness(ison_adolescents)
node_coreness(ison_adolescents)
}
\references{
diff --git a/man/mark_ties.Rd b/man/mark_ties.Rd
index 8979a1aa..180bee9f 100644
--- a/man/mark_ties.Rd
+++ b/man/mark_ties.Rd
@@ -61,8 +61,8 @@ tie_is_reciprocated(ison_algebra)
tie_is_feedback(ison_algebra)
tie_is_bridge(ison_brandes)
ison_adolescents \%>\%
- mutate_ties(route = tie_is_path(from = "Jane", to = 7)) \%>\%
- graphr(edge_colour = "route")
+ mutate_ties(route = tie_is_path(from = "Jane", to = 7))
+ #graphr(edge_colour = "route")
}
\seealso{
Other marks:
diff --git a/man/mark_triangles.Rd b/man/mark_triangles.Rd
index aeb301cb..edb37409 100644
--- a/man/mark_triangles.Rd
+++ b/man/mark_triangles.Rd
@@ -52,23 +52,23 @@ are cohesively connected.
}
\examples{
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(tri = tie_is_triangular()) \%>\%
- graphr(edge_color = "tri")
+ mutate_ties(tri = tie_is_triangular())
+ #graphr(edge_color = "tri")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trans = tie_is_transitive()) \%>\%
- graphr(edge_color = "trans")
+ mutate_ties(trans = tie_is_transitive())
+ #graphr(edge_color = "trans")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trip = tie_is_triplet()) \%>\%
- graphr(edge_color = "trip")
+ mutate_ties(trip = tie_is_triplet())
+ #graphr(edge_color = "trip")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(cyc = tie_is_cyclical()) \%>\%
- graphr(edge_color = "cyc")
+ mutate_ties(cyc = tie_is_cyclical())
+ #graphr(edge_color = "cyc")
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(simmel = tie_is_simmelian()) \%>\%
- graphr(edge_color = "simmel")
+ mutate_ties(simmel = tie_is_simmelian())
+ #graphr(edge_color = "simmel")
generate_random(8, directed = TRUE) \%>\%
- mutate_ties(forbid = tie_is_forbidden()) \%>\%
- graphr(edge_color = "forbid")
+ mutate_ties(forbid = tie_is_forbidden())
+ #graphr(edge_color = "forbid")
tie_is_imbalanced(ison_marvel_relationships)
}
\seealso{
diff --git a/man/measure_central_between.Rd b/man/measure_central_between.Rd
index b50e1b57..35a86c42 100644
--- a/man/measure_central_between.Rd
+++ b/man/measure_central_between.Rd
@@ -107,9 +107,7 @@ nodes, and thus associated with bridging or spanning boundaries.
node_betweenness(ison_southern_women)
node_induced(ison_adolescents)
(tb <- tie_betweenness(ison_adolescents))
-plot(tb)
-ison_adolescents \%>\% mutate_ties(weight = tb) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = tb)
net_betweenness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_central_close.Rd b/man/measure_central_close.Rd
index ac2a6e3a..1992e688 100644
--- a/man/measure_central_close.Rd
+++ b/man/measure_central_close.Rd
@@ -176,9 +176,7 @@ removing that node would disconnect the network.
node_closeness(ison_southern_women)
node_reach(ison_adolescents)
(ec <- tie_closeness(ison_adolescents))
-plot(ec)
-ison_adolescents \%>\% mutate_ties(weight = ec) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = ec)
net_closeness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_features.Rd b/man/measure_features.Rd
index f119f997..05666148 100644
--- a/man/measure_features.Rd
+++ b/man/measure_features.Rd
@@ -14,7 +14,11 @@
\code{{signnet}} by David Schoch
}
\usage{
-net_core(.data, mark = NULL)
+net_core(
+ .data,
+ mark = NULL,
+ method = c("correlation", "ident", "ndiff", "diff")
+)
net_richclub(.data)
@@ -41,13 +45,6 @@ net_balance(.data)
\item{mark}{A logical vector the length of the nodes in the network.
This can be created by, among other things, any \verb{node_is_*()} function.}
-\item{membership}{A vector of partition membership.}
-
-\item{resolution}{A proportion indicating the resolution scale.
-By default 1, which returns the original definition of modularity.
-The higher this parameter, the more smaller communities will be privileged.
-The lower this parameter, the fewer larger communities are likely to be found.}
-
\item{method}{There are three small-world measures implemented:
\itemize{
\item "sigma" is the original equation from Watts and Strogatz (1998),
@@ -72,6 +69,13 @@ with the same dimensions.
but where there may not be a network for which \eqn{SWI = 1}.
}}
+\item{membership}{A vector of partition membership.}
+
+\item{resolution}{A proportion indicating the resolution scale.
+By default 1, which returns the original definition of modularity.
+The higher this parameter, the more smaller communities will be privileged.
+The lower this parameter, the fewer larger communities are likely to be found.}
+
\item{times}{Integer of number of simulations.}
}
\description{
@@ -100,6 +104,14 @@ ranging between \code{0} if all triangles are imbalanced and
These \verb{net_*()} functions return a single numeric scalar or value.
}
+\section{Core-Periphery}{
+
+\code{net_core()} calculates the Pearson correlation between the given network,
+where the nodes in the core are assigned by some given mark, and an ideal
+typical core-periphery network with the same number of nodes in the core
+and the periphery.
+}
+
\section{Modularity}{
Modularity measures the difference between the number of ties within each community
diff --git a/man/reexports.Rd b/man/reexports.Rd
index 8efbe88d..7cc10505 100644
--- a/man/reexports.Rd
+++ b/man/reexports.Rd
@@ -1,5 +1,5 @@
% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/reexports_classes.R, R/reexports_ggplot2.R
+% Please edit documentation in R/reexports_classes.R
\docType{import}
\name{reexports}
\alias{reexports}
@@ -10,15 +10,6 @@
\alias{.G}
\alias{.N}
\alias{.E}
-\alias{ggplot}
-\alias{ggtitle}
-\alias{guides}
-\alias{labs}
-\alias{xlab}
-\alias{ylab}
-\alias{aes}
-\alias{ggsave}
-\alias{scale_size}
\title{Objects exported from other packages}
\keyword{internal}
\description{
@@ -28,8 +19,6 @@ below to see their documentation.
\describe{
\item{dplyr}{\code{\link[dplyr:reexports]{\%>\%}}}
- \item{ggplot2}{\code{\link[ggplot2]{aes}}, \code{\link[ggplot2]{ggplot}}, \code{\link[ggplot2]{ggsave}}, \code{\link[ggplot2:labs]{ggtitle}}, \code{\link[ggplot2]{guides}}, \code{\link[ggplot2]{labs}}, \code{\link[ggplot2]{scale_size}}, \code{\link[ggplot2:labs]{xlab}}, \code{\link[ggplot2:labs]{ylab}}}
-
\item{igraph}{\code{\link[igraph]{is_igraph}}}
\item{tidygraph}{\code{\link[tidygraph:context_accessors]{.E}}, \code{\link[tidygraph:context_accessors]{.G}}, \code{\link[tidygraph:context_accessors]{.N}}, \code{\link[tidygraph:tbl_graph]{is.tbl_graph}}, \code{\link[tidygraph]{with_graph}}}
diff --git a/tests/testthat/test-manip_reformed.R b/tests/testthat/test-manip_reformed.R
index 5afe68dd..d0f77e1e 100644
--- a/tests/testthat/test-manip_reformed.R
+++ b/tests/testthat/test-manip_reformed.R
@@ -1,3 +1,15 @@
+match_net <- to_matching(ison_southern_women)
+
+test_that("to_matching works with two-mode networks", {
+ expect_true(net_ties(match_net) == min(net_dims(ison_southern_women)))
+})
+
+test_that("to_matching is idempotent", {
+ match_twice <- to_matching(match_net)
+ expect_equal(as_edgelist(match_net),
+ as_edgelist(match_twice))
+})
+
test_that("to_no_missing.tbl_graph removes nodes with missing values", {
# Create a tbl_graph with some missing values
graph <- tbl_graph(
diff --git a/tests/testthat/test-map_autograph.R b/tests/testthat/test-map_autograph.R
deleted file mode 100644
index e44590be..00000000
--- a/tests/testthat/test-map_autograph.R
+++ /dev/null
@@ -1,187 +0,0 @@
-test_that("unweighted, unsigned, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, undirected network
- test_brandes <- graphr(ison_brandes)
- # Node position
- expect_equal(round(test_brandes[["data"]][["x"]][[1]]), 3)
- expect_equal(round(test_brandes[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- # Node parameters
- expect_equal(round(test_brandes[["layers"]][[2]][["aes_params"]][["size"]]), 11)
- expect_equal(as.character(test_brandes[["layers"]][[2]][["aes_params"]][["shape"]]), "circle")
-})
-
-test_that("unweighted, signed, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, signed, undirected network
- test_marvel <- graphr(to_giant(ison_marvel_relationships))
- # Node position
- expect_equal(round(test_marvel[["data"]][["x"]][[1]]), -1)
- expect_equal(round(test_marvel[["data"]][["y"]][[1]]), 1)
- # Edge parameters
- expect_equal(test_marvel[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- # Node parameters
- expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["size"]], 3)
- #expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("unweighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, directed network
- test_algebra <- graphr(ison_algebra)
- # Node position
- expect_equal(round(test_algebra[["data"]][["x"]][[1]]), 0)
- expect_equal(round(test_algebra[["data"]][["y"]][[1]]), 0)
- # Edge parameters
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_algebra[["layers"]][[1]][["mapping"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_algebra[["layers"]][[2]][["aes_params"]][["size"]]), 8)
- expect_equal(test_algebra[["layers"]][[2]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("weighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # Weighted, unsigned, directed network
- test_networkers <- graphr(ison_networkers)
- # Node position
- expect_equal(round(test_networkers[["data"]][["x"]][[1]]), 9)
- expect_equal(round(test_networkers[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_networkers[["layers"]][[3]][["aes_params"]][["size"]]), 3)
- #expect_equal(test_networkers[["layers"]][[3]][["aes_params"]][["shape"]], "circle")
-})
-
-# Testing the node_color, node_size, and node_shape args by specifying a node attribute
-test_that("fancy node mods graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # one-mode network
- ison_marvel_relationships <- dplyr::mutate(ison_marvel_relationships,
- nodesize = Appearances/1000)
- testcolnodes <- graphr(ison_marvel_relationships, node_color = "Gender",
- node_size = "nodesize", node_shape = "Attractive")
- expect_s3_class(testcolnodes, c("ggraph","gg","ggplot"))
- expect_equal(round(testcolnodes$data$x[1]), 4)
- expect_equal(round(testcolnodes$data$y[1]), 3)
- expect_equal(nrow(testcolnodes[["plot_env"]][["lo"]]),
- c(net_nodes(ison_marvel_relationships)))
- # two-mode network
- ison_southern_women <- add_node_attribute(ison_southern_women, "group",
- c(sample(c("a", "b"),
- length(ison_southern_women),
- replace = TRUE)))
- test2 <- graphr(ison_southern_women, node_color = "type")
- expect_s3_class(test2, c("ggraph","gg","ggplot"))
- expect_equal(round(test2$data$x[1]), 0)
- expect_equal(round(test2$data$y[1]), 0)
- expect_equal(nrow(test2[["plot_env"]][["lo"]]),
- c(net_nodes(ison_southern_women)))
-})
-
-test_that("edge colours and edge size graph correctly", {
- skip_on_cran()
- ison_brandes2 <- ison_brandes %>%
- add_tie_attribute("tiecolour",
- c("A", "B", "A", "B", "B", "B", "B", "B", "B", "B", "B", "B")) %>%
- add_tie_attribute("weight", c(rep(1:6, 2)))
- test_brandes2 <- graphr(ison_brandes2, edge_color = "tiecolour", edge_size = "weight")
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_colour))
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_width))
-})
-
-# Named networks
-test_that("named networks plot correctly", {
- skip_on_cran()
- skip_on_ci()
- onemode <- graphr(ison_adolescents)
- twomode <- graphr(ison_southern_women)
- expect_equal(onemode[["data"]][["name"]], node_names(ison_adolescents))
- expect_equal(twomode[["data"]][["name"]], node_names(ison_southern_women))
-})
-
-# Test that autographr() works with arguments without quotes
-test_that("node_group works correctly", {
- skip_on_cran()
- testthat::skip_if_not_installed("concaveman")
- expect_equal(graphr(ison_lawfirm, node_group = gender),
- graphr(ison_lawfirm, node_group = "gender"))
-})
-
-test_that("unquoted arguments plot correctly", {
- skip_on_cran()
- expect_equal(graphr(ison_lawfirm, node_color = "gender"),
- graphr(ison_lawfirm, node_color = gender))
-})
-
-# Layouts
-test_that("concentric and circular layouts graph correctly", {
- skip_on_cran()
- test_circle <- graphr(to_giant(ison_marvel_relationships),
- layout = "circle")
- test_conc <- graphr(to_giant(ison_marvel_relationships),
- layout = "concentric", membership = "Gender")
- expect_equal(test_circle$plot_env$layout, "circle")
- expect_equal(test_conc$plot_env$layout, "concentric")
- expect_equal(eval(quote(pairlist(...)),
- envir = test_conc$plot_env)$membership,
- "Gender")
-})
-
-test_that("hierarchy and lineage layouts graph correctly", {
- skip_on_cran()
- skip_on_ci()
- test_lin <- ison_adolescents %>%
- mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2)) %>%
- graphr(layout = "lineage", rank = "year")
- test_hie <- graphr(ison_southern_women,
- layout = "hierarchy", center = "events")
- expect_equal(test_lin$plot_env$layout, "lineage")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_lin[["plot_env"]])[["rank"]]),
- "year")
- expect_equal(test_hie$plot_env$layout, "hierarchy")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_hie[["plot_env"]])[["center"]]),
- "events")
-})
-
-# test_that("graphr works for diff_model objects", {
-# skip_on_cran()
-# skip_on_ci()
-# test_diff <- graphr(play_diffusion(ison_brandes, old_version = TRUE))
-# if (inherits(test_diff$guides, "Guides")) {
-# expect_s3_class(test_diff[["guides"]][["guides"]][["shape"]], "GuideLegend")
-# expect_s3_class(test_diff[["guides"]][["guides"]][["colour"]], "GuideColourbar")
-# } else {
-# expect_equal(test_diff[["guides"]][["shape"]][["name"]], "legend")
-# expect_equal(test_diff[["guides"]][["colour"]][["name"]], "colorbar")
-# }
-# })
-
-test_that("concentric layout works when node names are missing", {
- skip_on_cran()
- skip_on_ci()
- llabel <- ison_southern_women %>%
- mutate(name = ifelse(type == TRUE, "", name)) %>%
- graphr(layout = "concentric")
- expect_true(any(llabel$data$name == ""))
-})
-
-test_that("hierarchy layout works for two mode networks", {
- skip_on_cran()
- skip_on_ci()
- tm <- ison_brandes %>%
- mutate(type = twomode_type, name = LETTERS[1:11]) %>%
- graphr()
- expect_length(unique(tm$data[tm$data$type == TRUE, "y"]), 1)
- expect_length(unique(tm$data[tm$data$type == FALSE, "y"]), 1)
-})
diff --git a/tests/testthat/test-map_theme.R b/tests/testthat/test-map_theme.R
deleted file mode 100644
index 6e6ffe6f..00000000
--- a/tests/testthat/test-map_theme.R
+++ /dev/null
@@ -1,58 +0,0 @@
-##### test themes
-test_that("themes graph correctly", {
- test_iheid <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_iheid()
- test_ethz <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_ethz()
- test_uzh <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_uzh()
- expect_equal(names(test_iheid[["theme"]][["title"]][["colour"]]), "IHEIDRed")
- expect_null(names(test_ethz[["theme"]][["title"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["title"]][["colour"]]), "UZH_Orange")
- expect_equal(names(test_iheid[["theme"]][["plot.subtitle"]][["colour"]]),
- "IHEIDGrey")
- expect_null(names(test_ethz[["theme"]][["plot.subtitle"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["plot.subtitle"]][["colour"]]),
- "UZH_Blue")
- expect_equal(test_iheid[["theme"]][["plot.caption"]][["family"]], "serif")
- expect_equal(test_ethz[["theme"]][["plot.caption"]][["family"]], "sans")
- expect_equal(test_uzh[["theme"]][["plot.caption"]][["family"]], "sans")
-})
-
-##### test scales
-test_that("scales graph correctly", {
- test_sdg <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3,4,5), 2), 1)) %>%
- graphr(node_color = color) +
- scale_color_sdgs()
- test_iheid <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 5), 3)) %>%
- graphr(node_color = color) +
- scale_color_iheid()
- test_ethz <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 4, 5)) %>%
- graphr(node_color = color) +
- scale_color_ethz()
- test_uzh <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 1, 2)) %>%
- graphr(node_color = color) +
- scale_color_uzh()
- test_rug <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 4), 1, 2, 1)) %>%
- graphr(node_color = color) +
- scale_color_rug()
- expect_equal(as.character(test_sdg[["scales"]][["scales"]][[2]][["call"]]), "scale_color_sdgs")
- expect_equal(as.character(test_iheid[["scales"]][["scales"]][[2]][["call"]]), "scale_color_iheid")
- expect_equal(as.character(test_ethz[["scales"]][["scales"]][[2]][["call"]]), "scale_color_ethz")
- expect_equal(as.character(test_uzh[["scales"]][["scales"]][[2]][["call"]]), "scale_color_uzh")
- expect_equal(as.character(test_rug[["scales"]][["scales"]][[2]][["call"]]), "scale_color_rug")
-})
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions, each with their
diff --git a/cran-comments.md b/cran-comments.md
index 2827d039..634afb8c 100644
--- a/cran-comments.md
+++ b/cran-comments.md
@@ -9,4 +9,4 @@
0 errors | 0 warnings | 0 notes
-- Removed failing URLs from previous submission (v1.5.0)
\ No newline at end of file
+- Required for ggplot2 v4.0.0
\ No newline at end of file
diff --git a/man/defunct.Rd b/man/defunct.Rd
index 7258166b..1a5285e1 100644
--- a/man/defunct.Rd
+++ b/man/defunct.Rd
@@ -4,9 +4,6 @@
\alias{defunct}
\alias{pkg_data}
\alias{node_mode}
-\alias{autographr}
-\alias{autographs}
-\alias{autographd}
\alias{node_optimal}
\alias{node_kernighanlin}
\alias{node_edge_betweenness}
@@ -89,7 +86,6 @@
\alias{node_core}
\alias{node_by_quad}
\alias{net_by_quad}
-\alias{layout_tbl_graph_quad}
\alias{net_hazard}
\title{Functions that have been renamed, superseded, or are no longer working}
\usage{
@@ -97,34 +93,6 @@ pkg_data(pkg = "manynet")
node_mode(.data)
-autographr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- ...
-)
-
-autographs(netlist, waves, based_on = c("first", "last", "both"), ...)
-
-autographd(
- tlist,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- keep_isolates = TRUE,
- ...
-)
-
node_optimal(.data)
node_kernighanlin(.data)
@@ -295,8 +263,6 @@ node_by_quad(.data)
net_by_quad(.data)
-layout_tbl_graph_quad(.data, circular = FALSE, times = 1000)
-
net_hazard(.data)
}
\description{
@@ -314,12 +280,6 @@ wherever possible and update your scripts accordingly.
\item \code{node_mode()}: Deprecated on 2024-06-17.
-\item \code{autographr()}: Deprecated on 2024-06-17.
-
-\item \code{autographs()}: Deprecated on 2024-06-17.
-
-\item \code{autographd()}: Deprecated on 2024-06-17.
-
\item \code{node_optimal()}: Deprecated on 2024-06-14.
\item \code{node_kernighanlin()}: Deprecated on 2024-06-14.
@@ -486,8 +446,6 @@ wherever possible and update your scripts accordingly.
\item \code{net_by_quad()}: Deprecated on 2024-10-10.
-\item \code{layout_tbl_graph_quad()}: Deprecated on 2024-10-10.
-
\item \code{net_hazard()}: Deprecated on 2024-12-30.
}}
diff --git a/man/depth_first_recursive_search.Rd b/man/depth_first_recursive_search.Rd
deleted file mode 100644
index 9642a068..00000000
--- a/man/depth_first_recursive_search.Rd
+++ /dev/null
@@ -1,20 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{depth_first_recursive_search}
-\alias{depth_first_recursive_search}
-\title{Layouts for snapping layouts to a grid}
-\usage{
-depth_first_recursive_search(layout)
-}
-\description{
-The function uses approximate pattern matching
-to redistribute coarse layouts on square grid points, while
-preserving the topological relationships among the nodes (see Inoue et al. 2012).
-}
-\references{
-Inoue, Kentaro, Shinichi Shimozono, Hideaki Yoshida, and Hiroyuki Kurata. 2012.
-“Application of Approximate Pattern Matching in Two Dimensional Spaces to Grid Layout for Biochemical Network Maps” edited by J. Bourdon.
-\emph{PLoS ONE} 7(6):e37739.
-\doi{https://doi.org/10.1371/journal.pone.0037739}.
-}
-\keyword{internal}
diff --git a/man/make_cran.Rd b/man/make_cran.Rd
index b49d70b3..16557e6d 100644
--- a/man/make_cran.Rd
+++ b/man/make_cran.Rd
@@ -37,8 +37,6 @@ by \href{https://github.com/stocnet/manynet/issues}{raising an issue on Github}.
\examples{
# mnet <- read_cran()
# mnet <- to_ego(mnet, "manynet", max_dist = 2)
-# graphr(mnet, layout = "hierarchy",
-# edge_color = "type", node_color = "Compilation")
}
\seealso{
\link{as}
diff --git a/man/make_learning.Rd b/man/make_learning.Rd
index b06808d4..6f08083a 100644
--- a/man/make_learning.Rd
+++ b/man/make_learning.Rd
@@ -87,9 +87,6 @@ from other nodes whose beliefs are not too dissimilar from their own.
latticeEg <- add_node_attribute(latticeEg, "startValues", startValues)
latticeEg
play_segregation(latticeEg, "startValues", 0.5)
- # graphr(latticeEg, node_color = "startValues", node_size = 5) +
- # graphr(play_segregation(latticeEg, "startValues", 0.2),
- # node_color = "startValues", node_size = 5)
}
\references{
DeGroot, Morris H. 1974.
diff --git a/man/make_play.Rd b/man/make_play.Rd
index fe1b0621..7d1aa8c7 100644
--- a/man/make_play.Rd
+++ b/man/make_play.Rd
@@ -185,8 +185,6 @@ This can be used in in SEI, SEIS, SEIR, and SEIRS models.
\examples{
smeg <- generate_smallworld(15, 0.025)
- plot(play_diffusion(smeg, recovery = 0.4))
- #graphr(play_diffusion(ison_karateka))
}
\seealso{
Other makes:
diff --git a/man/make_random.Rd b/man/make_random.Rd
index 0146a541..e8471363 100644
--- a/man/make_random.Rd
+++ b/man/make_random.Rd
@@ -105,8 +105,8 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_random(12, 0.4))
-# graphr(generate_random(c(6, 6), 0.4))
+generate_random(12, 0.4)
+# generate_random(c(6, 6), 0.4)
}
\references{
\subsection{On random networks}{
diff --git a/man/make_stochastic.Rd b/man/make_stochastic.Rd
index f2930b77..0a977e6b 100644
--- a/man/make_stochastic.Rd
+++ b/man/make_stochastic.Rd
@@ -104,10 +104,10 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_smallworld(12, 0.025))
-graphr(generate_smallworld(12, 0.25))
-graphr(generate_scalefree(12, 0.25))
-graphr(generate_scalefree(12, 1.25))
+generate_smallworld(12, 0.025)
+generate_smallworld(12, 0.25)
+generate_scalefree(12, 0.25)
+generate_scalefree(12, 1.25)
generate_fire(10)
generate_islands(10)
generate_citations(10)
diff --git a/man/manip_paths.Rd b/man/manip_paths.Rd
index f62c282e..ca35715c 100644
--- a/man/manip_paths.Rd
+++ b/man/manip_paths.Rd
@@ -109,10 +109,7 @@ This is, however, computationally slower.
\examples{
to_matching(ison_southern_women)
-#graphr(to_matching(ison_southern_women))
-graphr(to_mentoring(ison_adolescents))
to_eulerian(delete_nodes(ison_koenigsberg, "Lomse"))
- #graphr(to_eulerian(delete_nodes(ison_koenigsberg, "Lomse")))
}
\references{
\subsection{On matching}{
@@ -124,7 +121,7 @@ Gale, David, and Lloyd Stowell Shapley. 1962.
Goldberg, Andrew V., and Robert E. Tarjan. 1986.
"A new approach to the maximum flow problem".
-\emph{Proceedings of the eighteenth annual ACM symposium on Theory of computing – STOC '86}.
+\emph{Proceedings of the 18th Annual ACM Symposium on Theory of Computing}.
136-146.
\doi{10.1145/12130.12144}
}
diff --git a/man/manip_permutation.Rd b/man/manip_permutation.Rd
index 0578c343..32a6df8b 100644
--- a/man/manip_permutation.Rd
+++ b/man/manip_permutation.Rd
@@ -26,10 +26,6 @@ By default TRUE.}
on both the rows and columns (for a one-mode network)
or on each of the rows and columns (for a two-mode network).
}
-\examples{
-graphr(ison_adolescents, node_size = 4)
-graphr(to_permuted(ison_adolescents), node_size = 4)
-}
\seealso{
Other modifications:
\code{\link{manip_as}},
diff --git a/man/manip_project.Rd b/man/manip_project.Rd
index 0b59bb28..04f6e5ec 100644
--- a/man/manip_project.Rd
+++ b/man/manip_project.Rd
@@ -71,10 +71,7 @@ Below are the currently implemented S3 methods:\tabular{lrrrrr}{
\examples{
to_mode1(ison_southern_women)
to_mode2(ison_southern_women)
-#graphr(to_mode1(ison_southern_women))
-#graphr(to_mode2(ison_southern_women))
to_ties(ison_adolescents)
-#graphr(to_ties(ison_adolescents))
}
\seealso{
Other modifications:
diff --git a/man/manip_reformat.Rd b/man/manip_reformat.Rd
index 464f1716..17185089 100644
--- a/man/manip_reformat.Rd
+++ b/man/manip_reformat.Rd
@@ -70,7 +70,6 @@ This essentially has no effect on undirected networks or reciprocated ties.
}}
\examples{
to_anti(ison_southern_women)
-#graphr(to_anti(ison_southern_women))
}
\seealso{
Other modifications:
diff --git a/man/manip_ties.Rd b/man/manip_ties.Rd
index 531efcc0..c53695bf 100644
--- a/man/manip_ties.Rd
+++ b/man/manip_ties.Rd
@@ -83,7 +83,7 @@ Note that while \verb{add_*()}/\verb{delete_*()} functions operate similarly as
other <- create_filled(4) \%>\% mutate(name = c("A", "B", "C", "D"))
mutate_ties(other, form = 1:6) \%>\% filter_ties(form < 4)
add_tie_attribute(other, "weight", c(1, 2, 2, 2, 1, 2))
-ison_adolescents \%>\% add_ties(c("Betty","Tina")) \%>\% graphr()
+ison_adolescents \%>\% add_ties(c("Betty","Tina"))
delete_ties(ison_adolescents, 3)
delete_ties(ison_adolescents, "Alice|Sue")
}
diff --git a/man/map_graphr.Rd b/man/map_graphr.Rd
deleted file mode 100644
index 474f3d4b..00000000
--- a/man/map_graphr.Rd
+++ /dev/null
@@ -1,141 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphr}
-\alias{map_graphr}
-\alias{graphr}
-\title{Easily graph networks with sensible defaults}
-\usage{
-graphr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- snap = FALSE,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{.data}{A manynet-consistent object.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{node_group}{Node variable to be used for grouping the nodes.
-It is easiest if this is added as a hull over
-groups before plotting.
-Group variables should have a minimum of 3 nodes,
-if less, number groups will be reduced by
-merging categories with lower counts into one called "other".}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{snap}{Logical scalar, whether the layout should be snapped to a grid.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-A \code{ggplot2::ggplot()} object.
-The last plot can be saved to the file system using \code{ggplot2::ggsave()}.
-}
-\description{
-This function provides users with an easy way to graph
-(m)any network data for exploration, investigation, inspiration,
-and communication.
-
-It builds upon \code{{ggplot2}} and \code{{ggraph}} to offer
-pretty and extensible graphing solutions.
-However, compared to those solutions,
-\code{graphr()} contains various algorithms to provide better looking
-graphs by default.
-This means that just passing the function some network data
-will often be sufficient to return a reasonable-looking graph.
-
-The function also makes it easy to modify many of the most
-commonly adapted aspects of a graph,
-including node and edge size, colour, and shape,
-as arguments rather than additional functions that you need to remember.
-These can be defined outright, e.g. \code{node_size = 8}, or
-in reference to an attribute of the network, e.g. \code{node_size = "wealth"}.
-
-Lastly, \code{graphr()} uses \code{{ggplot2}}-related theme information, so
-it is easy to make colour palette and fonts institution-specific and consistent.
-See e.g. \code{theme_iheid()} for more.
-
-To learn more about what can be done visually,
-try \code{run_tute("Visualisation")}.
-}
-\examples{
-graphr(ison_adolescents)
-ison_adolescents \%>\%
- mutate(color = rep(c("introvert","extrovert"), times = 4),
- size = ifelse(node_is_cutpoint(ison_adolescents), 6, 3)) \%>\%
- mutate_ties(ecolor = rep(c("friends", "acquaintances"), times = 5)) \%>\%
- graphr(node_color = "color", node_size = "size",
- edge_size = 1.5, edge_color = "ecolor")
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_graphs.Rd b/man/map_graphs.Rd
deleted file mode 100644
index 549d1128..00000000
--- a/man/map_graphs.Rd
+++ /dev/null
@@ -1,60 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphs}
-\alias{map_graphs}
-\alias{graphs}
-\title{Easily graph a set of networks with sensible defaults}
-\usage{
-graphs(netlist, waves, based_on = c("first", "last", "both"), ...)
-}
-\arguments{
-\item{netlist}{A list of manynet-compatible networks.}
-
-\item{waves}{Numeric, the number of plots to be displayed side-by-side.
-If missing, the number of plots will be reduced to the first and last
-when there are more than four plots.
-This argument can also be passed a vector selecting the waves to plot.}
-
-\item{based_on}{Whether the layout of the joint plots should
-be based on the "first" or the "last" network, or "both".}
-
-\item{...}{Additional arguments passed to \code{graphr()}.}
-}
-\value{
-Multiple \code{ggplot2::ggplot()} objects displayed side-by-side.
-}
-\description{
-This function provides users with an easy way to graph
-lists of network data for comparison.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{patchwork}} package to plot the graphs
-side by side and, if necessary, in successive rows.
-This is useful for lists of networks that represent, for example,
-ego or component subgraphs of a network,
-or a list of a network's different types of tie or across time.
-By default just the first and last network will be plotted,
-but this can be overridden by the "waves" parameter.
-
-Where the graphs are of the same network (same nodes),
-the graphs may share a layout to facilitate comparison.
-By default, successive graphs will use the layout calculated for
-the "first" network, but other options include the "last" layout,
-or a mix, "both", of them.
-}
-\examples{
-#graphs(to_egos(ison_adolescents))
-#graphs(to_egos(ison_adolescents), waves = 8)
-#graphs(to_egos(ison_adolescents), waves = c(2, 4, 6))
-#graphs(play_diffusion(ison_adolescents))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_grapht.Rd b/man/map_grapht.Rd
deleted file mode 100644
index ff0cce5a..00000000
--- a/man/map_grapht.Rd
+++ /dev/null
@@ -1,137 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_grapht}
-\alias{map_grapht}
-\alias{grapht}
-\title{Easily animate dynamic networks with sensible defaults}
-\source{
-https://blog.schochastics.net/posts/2021-09-15_animating-network-evolutions-with-gganimate/
-}
-\usage{
-grapht(
- tlist,
- keep_isolates = TRUE,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{tlist}{The same migraph-compatible network listed according to
-a time attribute, waves, or slices.}
-
-\item{keep_isolates}{Logical, whether to keep isolate nodes in the graph.
-TRUE by default.
-If FALSE, removes nodes from each frame they are isolated in.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-Shows a .gif image. Assigning the result of the function
-saves the gif to a temporary folder and the object holds the path to this file.
-}
-\description{
-This function provides users with an easy way to graph
-dynamic network data for exploration and presentation.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{gganimate}} package to animate the changes
-between successive iterations of a network.
-This is useful for networks in which the ties and/or the node or tie
-attributes are changing.
-
-A progress bar is shown if it takes some time to encoding all the
-.png files into a .gif.
-}
-\examples{
-#ison_adolescents \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year", cumulative = TRUE) \%>\%
-# grapht()
-#ison_adolescents \%>\%
-# mutate(gender = rep(c("male", "female"), times = 4),
-# hair = rep(c("black", "brown"), times = 4),
-# age = sample(11:16, 8, replace = TRUE)) \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE),
-# links = sample(c("friends", "not_friends"), 10, replace = TRUE),
-# weekly_meetings = sample(c(3, 5, 7), 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year") \%>\%
-# grapht(layout = "concentric", membership = "gender",
-# node_shape = "gender", node_color = "hair",
-# node_size = "age", edge_color = "links",
-# edge_size = "weekly_meetings")
-#grapht(play_diffusion(ison_adolescents, seeds = 5))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_configuration.Rd b/man/map_layout_configuration.Rd
deleted file mode 100644
index 8f8e0f33..00000000
--- a/man/map_layout_configuration.Rd
+++ /dev/null
@@ -1,54 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_configuration}
-\alias{map_layout_configuration}
-\alias{layout_tbl_graph_configuration}
-\alias{layout_tbl_graph_dyad}
-\alias{layout_tbl_graph_triad}
-\alias{layout_tbl_graph_tetrad}
-\alias{layout_tbl_graph_pentad}
-\alias{layout_tbl_graph_hexad}
-\title{Layout algorithms based on configurational positions}
-\usage{
-layout_tbl_graph_configuration(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_dyad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_triad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_tetrad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_pentad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_hexad(.data, circular = FALSE, times = 1000)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-}
-\description{
-Configurational layouts locate nodes at symmetric coordinates
-to help illustrate particular configurations.
-Currently "triad" and "quad" layouts are available.
-The "configuration" layout will choose the appropriate configurational
-layout automatically.
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_partition.Rd b/man/map_layout_partition.Rd
deleted file mode 100644
index c30b378a..00000000
--- a/man/map_layout_partition.Rd
+++ /dev/null
@@ -1,130 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_partition}
-\alias{map_layout_partition}
-\alias{layout_tbl_graph_hierarchy}
-\alias{layout_tbl_graph_alluvial}
-\alias{layout_tbl_graph_railway}
-\alias{layout_tbl_graph_ladder}
-\alias{layout_tbl_graph_concentric}
-\alias{layout_tbl_graph_multilevel}
-\alias{layout_tbl_graph_lineage}
-\title{Layout algorithms based on bi- or other partitions}
-\source{
-Diego Diez, Andrew P. Hutchins and Diego Miranda-Saavedra. 2014.
-"Systematic identification of transcriptional regulatory modules from
-protein-protein interaction networks".
-\emph{Nucleic Acids Research}, 42 (1) e6.
-}
-\usage{
-layout_tbl_graph_hierarchy(
- .data,
- center = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_alluvial(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_railway(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_ladder(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_concentric(
- .data,
- membership,
- radius = NULL,
- order.by = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_multilevel(.data, level, circular = FALSE)
-
-layout_tbl_graph_lineage(.data, rank, circular = FALSE)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{center}{Further split "hierarchical" layouts by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name in hierarchy layout.
-Defaults to NULL.}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-
-\item{membership}{A node attribute or a vector to draw concentric circles
-for "concentric" layout.}
-
-\item{radius}{A vector of radii at which the concentric circles
-should be located for "concentric" layout.
-By default this is equal placement around an empty centre,
-unless one (the core) is a single node,
-in which case this node occupies the centre of the graph.}
-
-\item{order.by}{An attribute label indicating the (decreasing) order
-for the nodes around the circles for "concentric" layout.
-By default ordering is given by a bipartite placement that reduces
-the number of edge crossings.}
-
-\item{level}{A node attribute or a vector to hierarchically order levels for
-"multilevel" layout.}
-
-\item{rank}{A numerical node attribute to place nodes in Y axis
-according to values for "lineage" layout.}
-}
-\description{
-These algorithms layout networks based on two or more partitions,
-and are recommended for use with \code{graphr()} or \code{{ggraph}}.
-Note that these layout algorithms use \code{{Rgraphviz}},
-a package that is only available on Bioconductor.
-It will first need to be downloaded using \code{BiocManager::install("Rgraphviz")}.
-If it has not already been installed, there is a prompt the first time
-these functions are used though.
-
-The "hierarchy" layout layers the first node set along the bottom,
-and the second node set along the top,
-sequenced and spaced as necessary to minimise edge overlap.
-The "alluvial" layout is similar to "hierarchy",
-but places successive layers horizontally rather than vertically.
-The "railway" layout is similar to "hierarchy",
-but nodes are aligned across the layers.
-The "ladder" layout is similar to "railway",
-but places successive layers horizontally rather than vertically.
-The "concentric" layout places a "hierarchy" layout
-around a circle, with successive layers appearing as concentric circles.
-The "multilevel" layout places successive layers as multiple levels.
-The "lineage" layout ranks nodes in Y axis according to values.
-}
-\examples{
-#graphr(ison_southern_women, layout = "hierarchy", center = "events",
-# node_color = "type", node_size = 3)
-#graphr(ison_southern_women, layout = "alluvial")
-#graphr(ison_southern_women, layout = "concentric", membership = "type",
-# node_color = "type", node_size = 3)
-#graphr(ison_lotr, layout = "multilevel",
-# node_color = "Race", level = "Race", node_size = 3)
-# ison_adolescents \%>\%
-# mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2),
-# cut = node_is_cutpoint(ison_adolescents)) \%>\%
-# graphr(layout = "lineage", rank = "year", node_color = "cut",
-# node_size = migraph::node_degree(ison_adolescents)*10)
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}}
-}
-\concept{mapping}
diff --git a/man/map_palettes.Rd b/man/map_palettes.Rd
deleted file mode 100644
index 8c086ea9..00000000
--- a/man/map_palettes.Rd
+++ /dev/null
@@ -1,32 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_palettes.R
-\name{map_palettes}
-\alias{map_palettes}
-\alias{many_palettes}
-\title{Many palettes generator}
-\source{
-Adapted from \url{https://github.com/karthik/wesanderson/blob/master/R/colors.R}
-}
-\usage{
-many_palettes(palette, n, type = c("discrete", "continuous"))
-}
-\arguments{
-\item{palette}{Name of desired palette. Current choices are:
-\code{IHEID}, \code{Centres}, \code{SDGs}, \code{ETHZ}, \code{RUG},
-and \code{UZH}.}
-
-\item{n}{Number of colors desired. If omitted, uses all colours.}
-
-\item{type}{Either "continuous" or "discrete". Use continuous if you want
-to automatically interpolate between colours.}
-}
-\value{
-A graphic display of colours in palette.
-}
-\description{
-Many palettes generator
-}
-\examples{
-many_palettes()
-#many_palettes("IHEID")
-}
diff --git a/man/map_scales.Rd b/man/map_scales.Rd
deleted file mode 100644
index bdee1bc6..00000000
--- a/man/map_scales.Rd
+++ /dev/null
@@ -1,112 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_scales}
-\alias{map_scales}
-\alias{scale_fill_iheid}
-\alias{scale_colour_iheid}
-\alias{scale_color_iheid}
-\alias{scale_edge_colour_iheid}
-\alias{scale_edge_color_iheid}
-\alias{scale_fill_centres}
-\alias{scale_colour_centres}
-\alias{scale_color_centres}
-\alias{scale_edge_colour_centres}
-\alias{scale_edge_color_centres}
-\alias{scale_fill_sdgs}
-\alias{scale_colour_sdgs}
-\alias{scale_color_sdgs}
-\alias{scale_edge_colour_sdgs}
-\alias{scale_edge_color_sdgs}
-\alias{scale_fill_ethz}
-\alias{scale_colour_ethz}
-\alias{scale_color_ethz}
-\alias{scale_edge_colour_ethz}
-\alias{scale_edge_color_ethz}
-\alias{scale_fill_uzh}
-\alias{scale_colour_uzh}
-\alias{scale_color_uzh}
-\alias{scale_edge_colour_uzh}
-\alias{scale_edge_color_uzh}
-\alias{scale_fill_rug}
-\alias{scale_colour_rug}
-\alias{scale_color_rug}
-\alias{scale_edge_colour_rug}
-\alias{scale_edge_color_rug}
-\title{Many scales}
-\usage{
-scale_fill_iheid(direction = 1, ...)
-
-scale_colour_iheid(direction = 1, ...)
-
-scale_color_iheid(direction = 1, ...)
-
-scale_edge_colour_iheid(direction = 1, ...)
-
-scale_edge_color_iheid(direction = 1, ...)
-
-scale_fill_centres(direction = 1, ...)
-
-scale_colour_centres(direction = 1, ...)
-
-scale_color_centres(direction = 1, ...)
-
-scale_edge_colour_centres(direction = 1, ...)
-
-scale_edge_color_centres(direction = 1, ...)
-
-scale_fill_sdgs(direction = 1, ...)
-
-scale_colour_sdgs(direction = 1, ...)
-
-scale_color_sdgs(direction = 1, ...)
-
-scale_edge_colour_sdgs(direction = 1, ...)
-
-scale_edge_color_sdgs(direction = 1, ...)
-
-scale_fill_ethz(direction = 1, ...)
-
-scale_colour_ethz(direction = 1, ...)
-
-scale_color_ethz(direction = 1, ...)
-
-scale_edge_colour_ethz(direction = 1, ...)
-
-scale_edge_color_ethz(direction = 1, ...)
-
-scale_fill_uzh(direction = 1, ...)
-
-scale_colour_uzh(direction = 1, ...)
-
-scale_color_uzh(direction = 1, ...)
-
-scale_edge_colour_uzh(direction = 1, ...)
-
-scale_edge_color_uzh(direction = 1, ...)
-
-scale_fill_rug(direction = 1, ...)
-
-scale_colour_rug(direction = 1, ...)
-
-scale_color_rug(direction = 1, ...)
-
-scale_edge_colour_rug(direction = 1, ...)
-
-scale_edge_color_rug(direction = 1, ...)
-}
-\arguments{
-\item{direction}{Direction for using palette colors.}
-
-\item{...}{Extra arguments passed to \code{ggplot2::discrete_scale()}.}
-}
-\description{
-These functions enable to add color scales to be graphs.
-}
-\examples{
-#ison_brandes \%>\%
-#mutate(core = migraph::node_is_core(ison_brandes)) \%>\%
-#graphr(node_color = "core") +
-#scale_color_iheid()
-#graphr(ison_physicians[[1]], edge_color = "type") +
-#scale_edge_color_ethz()
-}
diff --git a/man/map_themes.Rd b/man/map_themes.Rd
deleted file mode 100644
index 288176ee..00000000
--- a/man/map_themes.Rd
+++ /dev/null
@@ -1,41 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_themes}
-\alias{map_themes}
-\alias{set_manynet_theme}
-\alias{theme_iheid}
-\alias{theme_ethz}
-\alias{theme_uzh}
-\alias{theme_rug}
-\title{Many themes}
-\usage{
-set_manynet_theme(theme = "default")
-
-theme_iheid(base_size = 12, base_family = "serif")
-
-theme_ethz(base_size = 12, base_family = "sans")
-
-theme_uzh(base_size = 12, base_family = "sans")
-
-theme_rug(base_size = 12, base_family = "mono")
-}
-\arguments{
-\item{theme}{String naming a theme.
-By default "default".}
-
-\item{base_size}{Font size, by default 12.}
-
-\item{base_family}{Font family, by default "sans".}
-}
-\description{
-These functions enable graphs to be easily and quickly themed,
-e.g. changing the default colour of the graph's vertices and edges.
-}
-\examples{
-to_mentoring(ison_brandes) \%>\%
- mutate(color = c(rep(c(1,2,3), 3), 3)) \%>\%
- graphr(node_color = "color") +
- labs(title = "Who leads and who follows?") +
- scale_color_iheid() +
- theme_iheid()
-}
diff --git a/man/mark_core.Rd b/man/mark_core.Rd
index 6143dee3..7f3c805e 100644
--- a/man/mark_core.Rd
+++ b/man/mark_core.Rd
@@ -4,6 +4,7 @@
\alias{mark_core}
\alias{node_is_universal}
\alias{node_is_core}
+\alias{node_kcoreness}
\alias{node_coreness}
\title{Core-periphery clustering algorithms}
\usage{
@@ -11,6 +12,8 @@ node_is_universal(.data)
node_is_core(.data, method = c("degree", "eigenvector"))
+node_kcoreness(.data)
+
node_coreness(.data)
}
\arguments{
@@ -33,8 +36,13 @@ can be added if there is interest.}
\description{
These functions identify nodes belonging to (some level of) the core of a network:
\itemize{
-\item \code{node_is_core()} assigns nodes to either the core or periphery.
-\item \code{node_coreness()} assigns nodes to their level of k-coreness.
+\item \code{node_is_universal()} identifies whether nodes are adjacent to all other
+nodes in the network.
+\item \code{node_is_core()} identifies whether nodes belong to the core of the
+network, as opposed to the periphery.
+\item \code{node_coreness()} returns a continuous measure of how closely each node
+resembles a typical core node.
+\item \code{node_kcoreness()} assigns nodes to their level of k-coreness.
}
}
\section{Universal/dominating node}{
@@ -68,6 +76,7 @@ node_is_core(ison_adolescents)
#ison_adolescents \%>\%
# mutate(corep = node_is_core()) \%>\%
# graphr(node_color = "corep")
+node_kcoreness(ison_adolescents)
node_coreness(ison_adolescents)
}
\references{
diff --git a/man/mark_ties.Rd b/man/mark_ties.Rd
index 8979a1aa..180bee9f 100644
--- a/man/mark_ties.Rd
+++ b/man/mark_ties.Rd
@@ -61,8 +61,8 @@ tie_is_reciprocated(ison_algebra)
tie_is_feedback(ison_algebra)
tie_is_bridge(ison_brandes)
ison_adolescents \%>\%
- mutate_ties(route = tie_is_path(from = "Jane", to = 7)) \%>\%
- graphr(edge_colour = "route")
+ mutate_ties(route = tie_is_path(from = "Jane", to = 7))
+ #graphr(edge_colour = "route")
}
\seealso{
Other marks:
diff --git a/man/mark_triangles.Rd b/man/mark_triangles.Rd
index aeb301cb..edb37409 100644
--- a/man/mark_triangles.Rd
+++ b/man/mark_triangles.Rd
@@ -52,23 +52,23 @@ are cohesively connected.
}
\examples{
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(tri = tie_is_triangular()) \%>\%
- graphr(edge_color = "tri")
+ mutate_ties(tri = tie_is_triangular())
+ #graphr(edge_color = "tri")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trans = tie_is_transitive()) \%>\%
- graphr(edge_color = "trans")
+ mutate_ties(trans = tie_is_transitive())
+ #graphr(edge_color = "trans")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trip = tie_is_triplet()) \%>\%
- graphr(edge_color = "trip")
+ mutate_ties(trip = tie_is_triplet())
+ #graphr(edge_color = "trip")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(cyc = tie_is_cyclical()) \%>\%
- graphr(edge_color = "cyc")
+ mutate_ties(cyc = tie_is_cyclical())
+ #graphr(edge_color = "cyc")
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(simmel = tie_is_simmelian()) \%>\%
- graphr(edge_color = "simmel")
+ mutate_ties(simmel = tie_is_simmelian())
+ #graphr(edge_color = "simmel")
generate_random(8, directed = TRUE) \%>\%
- mutate_ties(forbid = tie_is_forbidden()) \%>\%
- graphr(edge_color = "forbid")
+ mutate_ties(forbid = tie_is_forbidden())
+ #graphr(edge_color = "forbid")
tie_is_imbalanced(ison_marvel_relationships)
}
\seealso{
diff --git a/man/measure_central_between.Rd b/man/measure_central_between.Rd
index b50e1b57..35a86c42 100644
--- a/man/measure_central_between.Rd
+++ b/man/measure_central_between.Rd
@@ -107,9 +107,7 @@ nodes, and thus associated with bridging or spanning boundaries.
node_betweenness(ison_southern_women)
node_induced(ison_adolescents)
(tb <- tie_betweenness(ison_adolescents))
-plot(tb)
-ison_adolescents \%>\% mutate_ties(weight = tb) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = tb)
net_betweenness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_central_close.Rd b/man/measure_central_close.Rd
index ac2a6e3a..1992e688 100644
--- a/man/measure_central_close.Rd
+++ b/man/measure_central_close.Rd
@@ -176,9 +176,7 @@ removing that node would disconnect the network.
node_closeness(ison_southern_women)
node_reach(ison_adolescents)
(ec <- tie_closeness(ison_adolescents))
-plot(ec)
-ison_adolescents \%>\% mutate_ties(weight = ec) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = ec)
net_closeness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_features.Rd b/man/measure_features.Rd
index f119f997..05666148 100644
--- a/man/measure_features.Rd
+++ b/man/measure_features.Rd
@@ -14,7 +14,11 @@
\code{{signnet}} by David Schoch
}
\usage{
-net_core(.data, mark = NULL)
+net_core(
+ .data,
+ mark = NULL,
+ method = c("correlation", "ident", "ndiff", "diff")
+)
net_richclub(.data)
@@ -41,13 +45,6 @@ net_balance(.data)
\item{mark}{A logical vector the length of the nodes in the network.
This can be created by, among other things, any \verb{node_is_*()} function.}
-\item{membership}{A vector of partition membership.}
-
-\item{resolution}{A proportion indicating the resolution scale.
-By default 1, which returns the original definition of modularity.
-The higher this parameter, the more smaller communities will be privileged.
-The lower this parameter, the fewer larger communities are likely to be found.}
-
\item{method}{There are three small-world measures implemented:
\itemize{
\item "sigma" is the original equation from Watts and Strogatz (1998),
@@ -72,6 +69,13 @@ with the same dimensions.
but where there may not be a network for which \eqn{SWI = 1}.
}}
+\item{membership}{A vector of partition membership.}
+
+\item{resolution}{A proportion indicating the resolution scale.
+By default 1, which returns the original definition of modularity.
+The higher this parameter, the more smaller communities will be privileged.
+The lower this parameter, the fewer larger communities are likely to be found.}
+
\item{times}{Integer of number of simulations.}
}
\description{
@@ -100,6 +104,14 @@ ranging between \code{0} if all triangles are imbalanced and
These \verb{net_*()} functions return a single numeric scalar or value.
}
+\section{Core-Periphery}{
+
+\code{net_core()} calculates the Pearson correlation between the given network,
+where the nodes in the core are assigned by some given mark, and an ideal
+typical core-periphery network with the same number of nodes in the core
+and the periphery.
+}
+
\section{Modularity}{
Modularity measures the difference between the number of ties within each community
diff --git a/man/reexports.Rd b/man/reexports.Rd
index 8efbe88d..7cc10505 100644
--- a/man/reexports.Rd
+++ b/man/reexports.Rd
@@ -1,5 +1,5 @@
% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/reexports_classes.R, R/reexports_ggplot2.R
+% Please edit documentation in R/reexports_classes.R
\docType{import}
\name{reexports}
\alias{reexports}
@@ -10,15 +10,6 @@
\alias{.G}
\alias{.N}
\alias{.E}
-\alias{ggplot}
-\alias{ggtitle}
-\alias{guides}
-\alias{labs}
-\alias{xlab}
-\alias{ylab}
-\alias{aes}
-\alias{ggsave}
-\alias{scale_size}
\title{Objects exported from other packages}
\keyword{internal}
\description{
@@ -28,8 +19,6 @@ below to see their documentation.
\describe{
\item{dplyr}{\code{\link[dplyr:reexports]{\%>\%}}}
- \item{ggplot2}{\code{\link[ggplot2]{aes}}, \code{\link[ggplot2]{ggplot}}, \code{\link[ggplot2]{ggsave}}, \code{\link[ggplot2:labs]{ggtitle}}, \code{\link[ggplot2]{guides}}, \code{\link[ggplot2]{labs}}, \code{\link[ggplot2]{scale_size}}, \code{\link[ggplot2:labs]{xlab}}, \code{\link[ggplot2:labs]{ylab}}}
-
\item{igraph}{\code{\link[igraph]{is_igraph}}}
\item{tidygraph}{\code{\link[tidygraph:context_accessors]{.E}}, \code{\link[tidygraph:context_accessors]{.G}}, \code{\link[tidygraph:context_accessors]{.N}}, \code{\link[tidygraph:tbl_graph]{is.tbl_graph}}, \code{\link[tidygraph]{with_graph}}}
diff --git a/tests/testthat/test-manip_reformed.R b/tests/testthat/test-manip_reformed.R
index 5afe68dd..d0f77e1e 100644
--- a/tests/testthat/test-manip_reformed.R
+++ b/tests/testthat/test-manip_reformed.R
@@ -1,3 +1,15 @@
+match_net <- to_matching(ison_southern_women)
+
+test_that("to_matching works with two-mode networks", {
+ expect_true(net_ties(match_net) == min(net_dims(ison_southern_women)))
+})
+
+test_that("to_matching is idempotent", {
+ match_twice <- to_matching(match_net)
+ expect_equal(as_edgelist(match_net),
+ as_edgelist(match_twice))
+})
+
test_that("to_no_missing.tbl_graph removes nodes with missing values", {
# Create a tbl_graph with some missing values
graph <- tbl_graph(
diff --git a/tests/testthat/test-map_autograph.R b/tests/testthat/test-map_autograph.R
deleted file mode 100644
index e44590be..00000000
--- a/tests/testthat/test-map_autograph.R
+++ /dev/null
@@ -1,187 +0,0 @@
-test_that("unweighted, unsigned, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, undirected network
- test_brandes <- graphr(ison_brandes)
- # Node position
- expect_equal(round(test_brandes[["data"]][["x"]][[1]]), 3)
- expect_equal(round(test_brandes[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- # Node parameters
- expect_equal(round(test_brandes[["layers"]][[2]][["aes_params"]][["size"]]), 11)
- expect_equal(as.character(test_brandes[["layers"]][[2]][["aes_params"]][["shape"]]), "circle")
-})
-
-test_that("unweighted, signed, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, signed, undirected network
- test_marvel <- graphr(to_giant(ison_marvel_relationships))
- # Node position
- expect_equal(round(test_marvel[["data"]][["x"]][[1]]), -1)
- expect_equal(round(test_marvel[["data"]][["y"]][[1]]), 1)
- # Edge parameters
- expect_equal(test_marvel[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- # Node parameters
- expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["size"]], 3)
- #expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("unweighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, directed network
- test_algebra <- graphr(ison_algebra)
- # Node position
- expect_equal(round(test_algebra[["data"]][["x"]][[1]]), 0)
- expect_equal(round(test_algebra[["data"]][["y"]][[1]]), 0)
- # Edge parameters
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_algebra[["layers"]][[1]][["mapping"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_algebra[["layers"]][[2]][["aes_params"]][["size"]]), 8)
- expect_equal(test_algebra[["layers"]][[2]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("weighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # Weighted, unsigned, directed network
- test_networkers <- graphr(ison_networkers)
- # Node position
- expect_equal(round(test_networkers[["data"]][["x"]][[1]]), 9)
- expect_equal(round(test_networkers[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_networkers[["layers"]][[3]][["aes_params"]][["size"]]), 3)
- #expect_equal(test_networkers[["layers"]][[3]][["aes_params"]][["shape"]], "circle")
-})
-
-# Testing the node_color, node_size, and node_shape args by specifying a node attribute
-test_that("fancy node mods graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # one-mode network
- ison_marvel_relationships <- dplyr::mutate(ison_marvel_relationships,
- nodesize = Appearances/1000)
- testcolnodes <- graphr(ison_marvel_relationships, node_color = "Gender",
- node_size = "nodesize", node_shape = "Attractive")
- expect_s3_class(testcolnodes, c("ggraph","gg","ggplot"))
- expect_equal(round(testcolnodes$data$x[1]), 4)
- expect_equal(round(testcolnodes$data$y[1]), 3)
- expect_equal(nrow(testcolnodes[["plot_env"]][["lo"]]),
- c(net_nodes(ison_marvel_relationships)))
- # two-mode network
- ison_southern_women <- add_node_attribute(ison_southern_women, "group",
- c(sample(c("a", "b"),
- length(ison_southern_women),
- replace = TRUE)))
- test2 <- graphr(ison_southern_women, node_color = "type")
- expect_s3_class(test2, c("ggraph","gg","ggplot"))
- expect_equal(round(test2$data$x[1]), 0)
- expect_equal(round(test2$data$y[1]), 0)
- expect_equal(nrow(test2[["plot_env"]][["lo"]]),
- c(net_nodes(ison_southern_women)))
-})
-
-test_that("edge colours and edge size graph correctly", {
- skip_on_cran()
- ison_brandes2 <- ison_brandes %>%
- add_tie_attribute("tiecolour",
- c("A", "B", "A", "B", "B", "B", "B", "B", "B", "B", "B", "B")) %>%
- add_tie_attribute("weight", c(rep(1:6, 2)))
- test_brandes2 <- graphr(ison_brandes2, edge_color = "tiecolour", edge_size = "weight")
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_colour))
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_width))
-})
-
-# Named networks
-test_that("named networks plot correctly", {
- skip_on_cran()
- skip_on_ci()
- onemode <- graphr(ison_adolescents)
- twomode <- graphr(ison_southern_women)
- expect_equal(onemode[["data"]][["name"]], node_names(ison_adolescents))
- expect_equal(twomode[["data"]][["name"]], node_names(ison_southern_women))
-})
-
-# Test that autographr() works with arguments without quotes
-test_that("node_group works correctly", {
- skip_on_cran()
- testthat::skip_if_not_installed("concaveman")
- expect_equal(graphr(ison_lawfirm, node_group = gender),
- graphr(ison_lawfirm, node_group = "gender"))
-})
-
-test_that("unquoted arguments plot correctly", {
- skip_on_cran()
- expect_equal(graphr(ison_lawfirm, node_color = "gender"),
- graphr(ison_lawfirm, node_color = gender))
-})
-
-# Layouts
-test_that("concentric and circular layouts graph correctly", {
- skip_on_cran()
- test_circle <- graphr(to_giant(ison_marvel_relationships),
- layout = "circle")
- test_conc <- graphr(to_giant(ison_marvel_relationships),
- layout = "concentric", membership = "Gender")
- expect_equal(test_circle$plot_env$layout, "circle")
- expect_equal(test_conc$plot_env$layout, "concentric")
- expect_equal(eval(quote(pairlist(...)),
- envir = test_conc$plot_env)$membership,
- "Gender")
-})
-
-test_that("hierarchy and lineage layouts graph correctly", {
- skip_on_cran()
- skip_on_ci()
- test_lin <- ison_adolescents %>%
- mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2)) %>%
- graphr(layout = "lineage", rank = "year")
- test_hie <- graphr(ison_southern_women,
- layout = "hierarchy", center = "events")
- expect_equal(test_lin$plot_env$layout, "lineage")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_lin[["plot_env"]])[["rank"]]),
- "year")
- expect_equal(test_hie$plot_env$layout, "hierarchy")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_hie[["plot_env"]])[["center"]]),
- "events")
-})
-
-# test_that("graphr works for diff_model objects", {
-# skip_on_cran()
-# skip_on_ci()
-# test_diff <- graphr(play_diffusion(ison_brandes, old_version = TRUE))
-# if (inherits(test_diff$guides, "Guides")) {
-# expect_s3_class(test_diff[["guides"]][["guides"]][["shape"]], "GuideLegend")
-# expect_s3_class(test_diff[["guides"]][["guides"]][["colour"]], "GuideColourbar")
-# } else {
-# expect_equal(test_diff[["guides"]][["shape"]][["name"]], "legend")
-# expect_equal(test_diff[["guides"]][["colour"]][["name"]], "colorbar")
-# }
-# })
-
-test_that("concentric layout works when node names are missing", {
- skip_on_cran()
- skip_on_ci()
- llabel <- ison_southern_women %>%
- mutate(name = ifelse(type == TRUE, "", name)) %>%
- graphr(layout = "concentric")
- expect_true(any(llabel$data$name == ""))
-})
-
-test_that("hierarchy layout works for two mode networks", {
- skip_on_cran()
- skip_on_ci()
- tm <- ison_brandes %>%
- mutate(type = twomode_type, name = LETTERS[1:11]) %>%
- graphr()
- expect_length(unique(tm$data[tm$data$type == TRUE, "y"]), 1)
- expect_length(unique(tm$data[tm$data$type == FALSE, "y"]), 1)
-})
diff --git a/tests/testthat/test-map_theme.R b/tests/testthat/test-map_theme.R
deleted file mode 100644
index 6e6ffe6f..00000000
--- a/tests/testthat/test-map_theme.R
+++ /dev/null
@@ -1,58 +0,0 @@
-##### test themes
-test_that("themes graph correctly", {
- test_iheid <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_iheid()
- test_ethz <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_ethz()
- test_uzh <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_uzh()
- expect_equal(names(test_iheid[["theme"]][["title"]][["colour"]]), "IHEIDRed")
- expect_null(names(test_ethz[["theme"]][["title"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["title"]][["colour"]]), "UZH_Orange")
- expect_equal(names(test_iheid[["theme"]][["plot.subtitle"]][["colour"]]),
- "IHEIDGrey")
- expect_null(names(test_ethz[["theme"]][["plot.subtitle"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["plot.subtitle"]][["colour"]]),
- "UZH_Blue")
- expect_equal(test_iheid[["theme"]][["plot.caption"]][["family"]], "serif")
- expect_equal(test_ethz[["theme"]][["plot.caption"]][["family"]], "sans")
- expect_equal(test_uzh[["theme"]][["plot.caption"]][["family"]], "sans")
-})
-
-##### test scales
-test_that("scales graph correctly", {
- test_sdg <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3,4,5), 2), 1)) %>%
- graphr(node_color = color) +
- scale_color_sdgs()
- test_iheid <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 5), 3)) %>%
- graphr(node_color = color) +
- scale_color_iheid()
- test_ethz <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 4, 5)) %>%
- graphr(node_color = color) +
- scale_color_ethz()
- test_uzh <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 1, 2)) %>%
- graphr(node_color = color) +
- scale_color_uzh()
- test_rug <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 4), 1, 2, 1)) %>%
- graphr(node_color = color) +
- scale_color_rug()
- expect_equal(as.character(test_sdg[["scales"]][["scales"]][[2]][["call"]]), "scale_color_sdgs")
- expect_equal(as.character(test_iheid[["scales"]][["scales"]][[2]][["call"]]), "scale_color_iheid")
- expect_equal(as.character(test_ethz[["scales"]][["scales"]][[2]][["call"]]), "scale_color_ethz")
- expect_equal(as.character(test_uzh[["scales"]][["scales"]][[2]][["call"]]), "scale_color_uzh")
- expect_equal(as.character(test_rug[["scales"]][["scales"]][[2]][["call"]]), "scale_color_rug")
-})
 -
-```{r layout-comparison, echo = FALSE, message=FALSE, dpi = 250, fig.height=4, eval = FALSE}
-library(manynet)
-library(igraph)
-library(gridBase)
-library(grid)
-
-par(mfrow=c(1, 2), mai = c(0,0,0.5,0))
-plot(as_igraph(ison_southern_women), layout = layout_as_bipartite, main = "{igraph} bipartite")
-## the last one is the current plot
-plot.new() ## suggested by @Josh
-vps <- baseViewports()
-pushViewport(vps$figure) ## I am in the space of the autocorrelation plot
-vp1 <-plotViewport(c(1.8,1,0,1)) ## create new vp with margins, you play with this values
-p <- graphr(ison_southern_women) + ggtitle("{manynet} twomode")
-print(p,vp = vp1)
-```
-
-Here the `{manynet}` function recognises that the network is a two-mode network
-and uses a bipartite layout by default,
-and recognises that the network contains names for the nodes and
-prints them vertically so that they are legible in this layout.
-Other 'clever' features include automatic node sizing and more.
-By contrast, `{igraph}` requires the bipartite layout to be specified, has cumbersome node size defaults for all but the smallest graphs, and labels also very often need resizing and adjustment to avoid overlap.
-All of `{manynet}`'s adjustments can be overridden, however...
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as
-specifying the function's relevant argument with a replacement,
-or indicating from which attribute it should inherit this information.
-
-
-
-```{r layout-comparison, echo = FALSE, message=FALSE, dpi = 250, fig.height=4, eval = FALSE}
-library(manynet)
-library(igraph)
-library(gridBase)
-library(grid)
-
-par(mfrow=c(1, 2), mai = c(0,0,0.5,0))
-plot(as_igraph(ison_southern_women), layout = layout_as_bipartite, main = "{igraph} bipartite")
-## the last one is the current plot
-plot.new() ## suggested by @Josh
-vps <- baseViewports()
-pushViewport(vps$figure) ## I am in the space of the autocorrelation plot
-vp1 <-plotViewport(c(1.8,1,0,1)) ## create new vp with margins, you play with this values
-p <- graphr(ison_southern_women) + ggtitle("{manynet} twomode")
-print(p,vp = vp1)
-```
-
-Here the `{manynet}` function recognises that the network is a two-mode network
-and uses a bipartite layout by default,
-and recognises that the network contains names for the nodes and
-prints them vertically so that they are legible in this layout.
-Other 'clever' features include automatic node sizing and more.
-By contrast, `{igraph}` requires the bipartite layout to be specified, has cumbersome node size defaults for all but the smallest graphs, and labels also very often need resizing and adjustment to avoid overlap.
-All of `{manynet}`'s adjustments can be overridden, however...
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as
-specifying the function's relevant argument with a replacement,
-or indicating from which attribute it should inherit this information.
-
- -
-```{r more-options, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-graphr(ison_lawfirm, node_color = "darkblue", node_size = 6) +
- ggtitle("Manual options",
- subtitle = "graphr(ison_lawfirm, node_color = 'darkblue', node_size = 6)") +
-graphr(mutate(ison_lawfirm, Seniority = Seniority/3), node_color = "Office", node_size = "Seniority") +
- ggtitle("Automatic options",
- subtitle = "graphr(ison_lawfirm, node_color = 'Office', node_size = 'Seniority')") &
- theme(plot.subtitle = element_text(size = 8))
-```
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such as `{igraph}`, `{ggraph}`, and `{graphlayouts}`,
-and offers some additional layout algorithms for
-snapping layouts to a grid,
-visualising partitions horizontally, vertically, or concentrically,
-or conforming to configurational coordinates.
-
-
-
-```{r more-options, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-graphr(ison_lawfirm, node_color = "darkblue", node_size = 6) +
- ggtitle("Manual options",
- subtitle = "graphr(ison_lawfirm, node_color = 'darkblue', node_size = 6)") +
-graphr(mutate(ison_lawfirm, Seniority = Seniority/3), node_color = "Office", node_size = "Seniority") +
- ggtitle("Automatic options",
- subtitle = "graphr(ison_lawfirm, node_color = 'Office', node_size = 'Seniority')") &
- theme(plot.subtitle = element_text(size = 8))
-```
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such as `{igraph}`, `{ggraph}`, and `{graphlayouts}`,
-and offers some additional layout algorithms for
-snapping layouts to a grid,
-visualising partitions horizontally, vertically, or concentrically,
-or conforming to configurational coordinates.
-
- -
-```{r more-layouts, echo = FALSE, message=FALSE, dpi = 250, eval=FALSE}
-(graphr(ison_southern_women, layout = "concentric") + ggtitle("Concentric layout")) /
- ((graphr(to_unnamed(create_explicit(A-+B-+C, A-+C))) + ggtitle("Triad layout")) |
- (graphr(to_unnamed(create_explicit(A-+C, A-+D, B-+C, B-+D))) + ggtitle("Quad layout")))
-```
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of your plots.
-`{manynet}` uses the excellent `{ggraph}` package (and thus `{ggplot2}`)
-as a plotting engine.
-This enables alterations such as the application of themes to be applied upon the defaults.
-If you want to quickly make sure your plots conform to your institution or taste,
-then it is easy to do with themes and scales that update the basic look and color palette
-used in your plots.
-
-
-
-```{r more-layouts, echo = FALSE, message=FALSE, dpi = 250, eval=FALSE}
-(graphr(ison_southern_women, layout = "concentric") + ggtitle("Concentric layout")) /
- ((graphr(to_unnamed(create_explicit(A-+B-+C, A-+C))) + ggtitle("Triad layout")) |
- (graphr(to_unnamed(create_explicit(A-+C, A-+D, B-+C, B-+D))) + ggtitle("Quad layout")))
-```
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of your plots.
-`{manynet}` uses the excellent `{ggraph}` package (and thus `{ggplot2}`)
-as a plotting engine.
-This enables alterations such as the application of themes to be applied upon the defaults.
-If you want to quickly make sure your plots conform to your institution or taste,
-then it is easy to do with themes and scales that update the basic look and color palette
-used in your plots.
-
- -
-```{r more-themes, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-p <- graphr(ison_lawfirm, node_color = "Practice") +
- ggtitle("Original")
-p + p + theme_iheid() + scale_color_iheid() +
- ggtitle("Graduate Institute", subtitle = "_iheid") +
- p + theme_ethz() + scale_color_ethz() +
- ggtitle("ETH Zürich", subtitle = "_ethz") & ggplot2::theme(legend.position = "none")
-```
-
-More themes are on their way, and we're happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together,
-which can be useful for ego networks or network panels.
-`{patchwork}` is used to help arrange individual plots together.
-
-
-
-```{r more-themes, echo = FALSE, message=FALSE, dpi = 300, fig.height=3, eval=FALSE}
-p <- graphr(ison_lawfirm, node_color = "Practice") +
- ggtitle("Original")
-p + p + theme_iheid() + scale_color_iheid() +
- ggtitle("Graduate Institute", subtitle = "_iheid") +
- p + theme_ethz() + scale_color_ethz() +
- ggtitle("ETH Zürich", subtitle = "_ethz") & ggplot2::theme(legend.position = "none")
-```
-
-More themes are on their way, and we're happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together,
-which can be useful for ego networks or network panels.
-`{patchwork}` is used to help arrange individual plots together.
-
- -
-```{r autographs, echo = FALSE, dpi = 250, fig.height=3, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(wave = c(rep(1995, 5), rep(1998, 5))) %>%
- to_waves(attribute = "wave", panels = c(1995, 1998)) %>%
- graphs()
-```
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks.
-It uses `{gganimate}` and `{gifski}` to create a gif that
-visualises network changes over time.
-It really couldn't be easier.
-
-
-
-```{r autographs, echo = FALSE, dpi = 250, fig.height=3, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(wave = c(rep(1995, 5), rep(1998, 5))) %>%
- to_waves(attribute = "wave", panels = c(1995, 1998)) %>%
- graphs()
-```
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks.
-It uses `{gganimate}` and `{gifski}` to create a gif that
-visualises network changes over time.
-It really couldn't be easier.
-
- -
-```{r autographd, echo = FALSE, dpi = 250, fig.height=3.5, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) %>%
- to_waves(attribute = "year") %>%
- grapht()
-```
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions,
diff --git a/README.md b/README.md
index a41d6f4a..68a18c7b 100644
--- a/README.md
+++ b/README.md
@@ -38,9 +38,11 @@ make learning and using network analysis tools in R challenging.
By contrast, `{manynet}` offers *many* analytic tools that work on
*many* (if not most) types and kinds of networks. It helps researchers
-make, modify, map, mark, measure, and identify nodes’ motifs and
-memberships in networks. For further testing and modelling capabilities,
-see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
+make, modify, mark, measure, and identify nodes’ motifs and memberships
+in networks. For graph drawing see
+[`{autograph}`](https://stocnet.github.io/autograph/), and for further
+testing and modelling capabilities see
+[`{migraph}`](https://stocnet.github.io/migraph/) and the other
[stocnet](https://github.com/stocnet) packages.
- [Making](#making)
@@ -53,13 +55,6 @@ see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
- [Transforming](#transforming)
- [Splitting and Joining](#splitting-and-joining)
- [Extracting](#extracting)
-- [Mapping](#mapping)
- - [Graphing](#graphing)
- - [More options](#more-options)
- - [More layouts](#more-layouts)
- - [More themes and scales](#more-themes-and-scales)
- - [graphs](#graphs)
- - [grapht](#grapht)
- [Marking](#marking)
- [Motifs](#motifs)
- [Memberships](#memberships)
@@ -237,99 +232,6 @@ See also `to_acyclic()`, `to_anti()`, `to_blocks()`, `to_components()`,
`to_unweighted()`, `to_waves()`, `to_weighted()` and `from_egos()`,
`from_slices()`, `from_subgraphs()`, `from_ties()`, `from_waves()`.
-## Mapping
-
-`{manynet}` includes three one-line graphing functions with sensible
-defaults based on the network’s properties.
-
-### Graphing
-
-First, `graphr()` is used to graph networks in any of the `{manynet}`
-formats. It includes sensible defaults so that researchers can view
-their network’s structure or distribution quickly with a minimum of
-fuss. Compare the output from `{manynet}` with a similar default from
-`{igraph}`:
-
-
-
-```{r autographd, echo = FALSE, dpi = 250, fig.height=3.5, eval=FALSE}
-ison_adolescents %>%
- mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) %>%
- to_waves(attribute = "year") %>%
- grapht()
-```
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions,
diff --git a/README.md b/README.md
index a41d6f4a..68a18c7b 100644
--- a/README.md
+++ b/README.md
@@ -38,9 +38,11 @@ make learning and using network analysis tools in R challenging.
By contrast, `{manynet}` offers *many* analytic tools that work on
*many* (if not most) types and kinds of networks. It helps researchers
-make, modify, map, mark, measure, and identify nodes’ motifs and
-memberships in networks. For further testing and modelling capabilities,
-see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
+make, modify, mark, measure, and identify nodes’ motifs and memberships
+in networks. For graph drawing see
+[`{autograph}`](https://stocnet.github.io/autograph/), and for further
+testing and modelling capabilities see
+[`{migraph}`](https://stocnet.github.io/migraph/) and the other
[stocnet](https://github.com/stocnet) packages.
- [Making](#making)
@@ -53,13 +55,6 @@ see [`{migraph}`](https://stocnet.github.io/migraph/) and the other
- [Transforming](#transforming)
- [Splitting and Joining](#splitting-and-joining)
- [Extracting](#extracting)
-- [Mapping](#mapping)
- - [Graphing](#graphing)
- - [More options](#more-options)
- - [More layouts](#more-layouts)
- - [More themes and scales](#more-themes-and-scales)
- - [graphs](#graphs)
- - [grapht](#grapht)
- [Marking](#marking)
- [Motifs](#motifs)
- [Memberships](#memberships)
@@ -237,99 +232,6 @@ See also `to_acyclic()`, `to_anti()`, `to_blocks()`, `to_components()`,
`to_unweighted()`, `to_waves()`, `to_weighted()` and `from_egos()`,
`from_slices()`, `from_subgraphs()`, `from_ties()`, `from_waves()`.
-## Mapping
-
-`{manynet}` includes three one-line graphing functions with sensible
-defaults based on the network’s properties.
-
-### Graphing
-
-First, `graphr()` is used to graph networks in any of the `{manynet}`
-formats. It includes sensible defaults so that researchers can view
-their network’s structure or distribution quickly with a minimum of
-fuss. Compare the output from `{manynet}` with a similar default from
-`{igraph}`:
-
- -
-Here the `{manynet}` function recognises that the network is a two-mode
-network and uses a bipartite layout by default, and recognises that the
-network contains names for the nodes and prints them vertically so that
-they are legible in this layout. Other ‘clever’ features include
-automatic node sizing and more. By contrast, `{igraph}` requires the
-bipartite layout to be specified, has cumbersome node size defaults for
-all but the smallest graphs, and labels also very often need resizing
-and adjustment to avoid overlap. All of `{manynet}`’s adjustments can be
-overridden, however…
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as specifying
-the function’s relevant argument with a replacement, or indicating from
-which attribute it should inherit this information.
-
-
-
-Here the `{manynet}` function recognises that the network is a two-mode
-network and uses a bipartite layout by default, and recognises that the
-network contains names for the nodes and prints them vertically so that
-they are legible in this layout. Other ‘clever’ features include
-automatic node sizing and more. By contrast, `{igraph}` requires the
-bipartite layout to be specified, has cumbersome node size defaults for
-all but the smallest graphs, and labels also very often need resizing
-and adjustment to avoid overlap. All of `{manynet}`’s adjustments can be
-overridden, however…
-
-#### More options
-
-Changing the size and colors of nodes and ties is as easy as specifying
-the function’s relevant argument with a replacement, or indicating from
-which attribute it should inherit this information.
-
- -
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such
-as `{igraph}`, `{ggraph}`, and `{graphlayouts}`, and offers some
-additional layout algorithms for snapping layouts to a grid, visualising
-partitions horizontally, vertically, or concentrically, or conforming to
-configurational coordinates.
-
-
-
-#### More layouts
-
-`{manynet}` can use all the layout algorithms offered by packages such
-as `{igraph}`, `{ggraph}`, and `{graphlayouts}`, and offers some
-additional layout algorithms for snapping layouts to a grid, visualising
-partitions horizontally, vertically, or concentrically, or conforming to
-configurational coordinates.
-
- -
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of
-your plots. `{manynet}` uses the excellent `{ggraph}` package (and thus
-`{ggplot2}`) as a plotting engine. This enables alterations such as the
-application of themes to be applied upon the defaults. If you want to
-quickly make sure your plots conform to your institution or taste, then
-it is easy to do with themes and scales that update the basic look and
-color palette used in your plots.
-
-
-
-#### More themes and scales
-
-Lastly, `graphr()` is highly extensible in terms of the overall look of
-your plots. `{manynet}` uses the excellent `{ggraph}` package (and thus
-`{ggplot2}`) as a plotting engine. This enables alterations such as the
-application of themes to be applied upon the defaults. If you want to
-quickly make sure your plots conform to your institution or taste, then
-it is easy to do with themes and scales that update the basic look and
-color palette used in your plots.
-
- -
-More themes are on their way, and we’re happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together, which
-can be useful for ego networks or network panels. `{patchwork}` is used
-to help arrange individual plots together.
-
-
-
-More themes are on their way, and we’re happy to take suggestions.
-
-### graphs
-
-Second, `graphs()` is used to graph multiple networks together, which
-can be useful for ego networks or network panels. `{patchwork}` is used
-to help arrange individual plots together.
-
- -
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks. It uses
-`{gganimate}` and `{gifski}` to create a gif that visualises network
-changes over time. It really couldn’t be easier.
-
-
-
-### grapht
-
-Third, `grapht()` is used to visualise dynamic networks. It uses
-`{gganimate}` and `{gifski}` to create a gif that visualises network
-changes over time. It really couldn’t be easier.
-
- -
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions, each with their
diff --git a/cran-comments.md b/cran-comments.md
index 2827d039..634afb8c 100644
--- a/cran-comments.md
+++ b/cran-comments.md
@@ -9,4 +9,4 @@
0 errors | 0 warnings | 0 notes
-- Removed failing URLs from previous submission (v1.5.0)
\ No newline at end of file
+- Required for ggplot2 v4.0.0
\ No newline at end of file
diff --git a/man/defunct.Rd b/man/defunct.Rd
index 7258166b..1a5285e1 100644
--- a/man/defunct.Rd
+++ b/man/defunct.Rd
@@ -4,9 +4,6 @@
\alias{defunct}
\alias{pkg_data}
\alias{node_mode}
-\alias{autographr}
-\alias{autographs}
-\alias{autographd}
\alias{node_optimal}
\alias{node_kernighanlin}
\alias{node_edge_betweenness}
@@ -89,7 +86,6 @@
\alias{node_core}
\alias{node_by_quad}
\alias{net_by_quad}
-\alias{layout_tbl_graph_quad}
\alias{net_hazard}
\title{Functions that have been renamed, superseded, or are no longer working}
\usage{
@@ -97,34 +93,6 @@ pkg_data(pkg = "manynet")
node_mode(.data)
-autographr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- ...
-)
-
-autographs(netlist, waves, based_on = c("first", "last", "both"), ...)
-
-autographd(
- tlist,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- keep_isolates = TRUE,
- ...
-)
-
node_optimal(.data)
node_kernighanlin(.data)
@@ -295,8 +263,6 @@ node_by_quad(.data)
net_by_quad(.data)
-layout_tbl_graph_quad(.data, circular = FALSE, times = 1000)
-
net_hazard(.data)
}
\description{
@@ -314,12 +280,6 @@ wherever possible and update your scripts accordingly.
\item \code{node_mode()}: Deprecated on 2024-06-17.
-\item \code{autographr()}: Deprecated on 2024-06-17.
-
-\item \code{autographs()}: Deprecated on 2024-06-17.
-
-\item \code{autographd()}: Deprecated on 2024-06-17.
-
\item \code{node_optimal()}: Deprecated on 2024-06-14.
\item \code{node_kernighanlin()}: Deprecated on 2024-06-14.
@@ -486,8 +446,6 @@ wherever possible and update your scripts accordingly.
\item \code{net_by_quad()}: Deprecated on 2024-10-10.
-\item \code{layout_tbl_graph_quad()}: Deprecated on 2024-10-10.
-
\item \code{net_hazard()}: Deprecated on 2024-12-30.
}}
diff --git a/man/depth_first_recursive_search.Rd b/man/depth_first_recursive_search.Rd
deleted file mode 100644
index 9642a068..00000000
--- a/man/depth_first_recursive_search.Rd
+++ /dev/null
@@ -1,20 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{depth_first_recursive_search}
-\alias{depth_first_recursive_search}
-\title{Layouts for snapping layouts to a grid}
-\usage{
-depth_first_recursive_search(layout)
-}
-\description{
-The function uses approximate pattern matching
-to redistribute coarse layouts on square grid points, while
-preserving the topological relationships among the nodes (see Inoue et al. 2012).
-}
-\references{
-Inoue, Kentaro, Shinichi Shimozono, Hideaki Yoshida, and Hiroyuki Kurata. 2012.
-“Application of Approximate Pattern Matching in Two Dimensional Spaces to Grid Layout for Biochemical Network Maps” edited by J. Bourdon.
-\emph{PLoS ONE} 7(6):e37739.
-\doi{https://doi.org/10.1371/journal.pone.0037739}.
-}
-\keyword{internal}
diff --git a/man/make_cran.Rd b/man/make_cran.Rd
index b49d70b3..16557e6d 100644
--- a/man/make_cran.Rd
+++ b/man/make_cran.Rd
@@ -37,8 +37,6 @@ by \href{https://github.com/stocnet/manynet/issues}{raising an issue on Github}.
\examples{
# mnet <- read_cran()
# mnet <- to_ego(mnet, "manynet", max_dist = 2)
-# graphr(mnet, layout = "hierarchy",
-# edge_color = "type", node_color = "Compilation")
}
\seealso{
\link{as}
diff --git a/man/make_learning.Rd b/man/make_learning.Rd
index b06808d4..6f08083a 100644
--- a/man/make_learning.Rd
+++ b/man/make_learning.Rd
@@ -87,9 +87,6 @@ from other nodes whose beliefs are not too dissimilar from their own.
latticeEg <- add_node_attribute(latticeEg, "startValues", startValues)
latticeEg
play_segregation(latticeEg, "startValues", 0.5)
- # graphr(latticeEg, node_color = "startValues", node_size = 5) +
- # graphr(play_segregation(latticeEg, "startValues", 0.2),
- # node_color = "startValues", node_size = 5)
}
\references{
DeGroot, Morris H. 1974.
diff --git a/man/make_play.Rd b/man/make_play.Rd
index fe1b0621..7d1aa8c7 100644
--- a/man/make_play.Rd
+++ b/man/make_play.Rd
@@ -185,8 +185,6 @@ This can be used in in SEI, SEIS, SEIR, and SEIRS models.
\examples{
smeg <- generate_smallworld(15, 0.025)
- plot(play_diffusion(smeg, recovery = 0.4))
- #graphr(play_diffusion(ison_karateka))
}
\seealso{
Other makes:
diff --git a/man/make_random.Rd b/man/make_random.Rd
index 0146a541..e8471363 100644
--- a/man/make_random.Rd
+++ b/man/make_random.Rd
@@ -105,8 +105,8 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_random(12, 0.4))
-# graphr(generate_random(c(6, 6), 0.4))
+generate_random(12, 0.4)
+# generate_random(c(6, 6), 0.4)
}
\references{
\subsection{On random networks}{
diff --git a/man/make_stochastic.Rd b/man/make_stochastic.Rd
index f2930b77..0a977e6b 100644
--- a/man/make_stochastic.Rd
+++ b/man/make_stochastic.Rd
@@ -104,10 +104,10 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_smallworld(12, 0.025))
-graphr(generate_smallworld(12, 0.25))
-graphr(generate_scalefree(12, 0.25))
-graphr(generate_scalefree(12, 1.25))
+generate_smallworld(12, 0.025)
+generate_smallworld(12, 0.25)
+generate_scalefree(12, 0.25)
+generate_scalefree(12, 1.25)
generate_fire(10)
generate_islands(10)
generate_citations(10)
diff --git a/man/manip_paths.Rd b/man/manip_paths.Rd
index f62c282e..ca35715c 100644
--- a/man/manip_paths.Rd
+++ b/man/manip_paths.Rd
@@ -109,10 +109,7 @@ This is, however, computationally slower.
\examples{
to_matching(ison_southern_women)
-#graphr(to_matching(ison_southern_women))
-graphr(to_mentoring(ison_adolescents))
to_eulerian(delete_nodes(ison_koenigsberg, "Lomse"))
- #graphr(to_eulerian(delete_nodes(ison_koenigsberg, "Lomse")))
}
\references{
\subsection{On matching}{
@@ -124,7 +121,7 @@ Gale, David, and Lloyd Stowell Shapley. 1962.
Goldberg, Andrew V., and Robert E. Tarjan. 1986.
"A new approach to the maximum flow problem".
-\emph{Proceedings of the eighteenth annual ACM symposium on Theory of computing – STOC '86}.
+\emph{Proceedings of the 18th Annual ACM Symposium on Theory of Computing}.
136-146.
\doi{10.1145/12130.12144}
}
diff --git a/man/manip_permutation.Rd b/man/manip_permutation.Rd
index 0578c343..32a6df8b 100644
--- a/man/manip_permutation.Rd
+++ b/man/manip_permutation.Rd
@@ -26,10 +26,6 @@ By default TRUE.}
on both the rows and columns (for a one-mode network)
or on each of the rows and columns (for a two-mode network).
}
-\examples{
-graphr(ison_adolescents, node_size = 4)
-graphr(to_permuted(ison_adolescents), node_size = 4)
-}
\seealso{
Other modifications:
\code{\link{manip_as}},
diff --git a/man/manip_project.Rd b/man/manip_project.Rd
index 0b59bb28..04f6e5ec 100644
--- a/man/manip_project.Rd
+++ b/man/manip_project.Rd
@@ -71,10 +71,7 @@ Below are the currently implemented S3 methods:\tabular{lrrrrr}{
\examples{
to_mode1(ison_southern_women)
to_mode2(ison_southern_women)
-#graphr(to_mode1(ison_southern_women))
-#graphr(to_mode2(ison_southern_women))
to_ties(ison_adolescents)
-#graphr(to_ties(ison_adolescents))
}
\seealso{
Other modifications:
diff --git a/man/manip_reformat.Rd b/man/manip_reformat.Rd
index 464f1716..17185089 100644
--- a/man/manip_reformat.Rd
+++ b/man/manip_reformat.Rd
@@ -70,7 +70,6 @@ This essentially has no effect on undirected networks or reciprocated ties.
}}
\examples{
to_anti(ison_southern_women)
-#graphr(to_anti(ison_southern_women))
}
\seealso{
Other modifications:
diff --git a/man/manip_ties.Rd b/man/manip_ties.Rd
index 531efcc0..c53695bf 100644
--- a/man/manip_ties.Rd
+++ b/man/manip_ties.Rd
@@ -83,7 +83,7 @@ Note that while \verb{add_*()}/\verb{delete_*()} functions operate similarly as
other <- create_filled(4) \%>\% mutate(name = c("A", "B", "C", "D"))
mutate_ties(other, form = 1:6) \%>\% filter_ties(form < 4)
add_tie_attribute(other, "weight", c(1, 2, 2, 2, 1, 2))
-ison_adolescents \%>\% add_ties(c("Betty","Tina")) \%>\% graphr()
+ison_adolescents \%>\% add_ties(c("Betty","Tina"))
delete_ties(ison_adolescents, 3)
delete_ties(ison_adolescents, "Alice|Sue")
}
diff --git a/man/map_graphr.Rd b/man/map_graphr.Rd
deleted file mode 100644
index 474f3d4b..00000000
--- a/man/map_graphr.Rd
+++ /dev/null
@@ -1,141 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphr}
-\alias{map_graphr}
-\alias{graphr}
-\title{Easily graph networks with sensible defaults}
-\usage{
-graphr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- snap = FALSE,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{.data}{A manynet-consistent object.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{node_group}{Node variable to be used for grouping the nodes.
-It is easiest if this is added as a hull over
-groups before plotting.
-Group variables should have a minimum of 3 nodes,
-if less, number groups will be reduced by
-merging categories with lower counts into one called "other".}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{snap}{Logical scalar, whether the layout should be snapped to a grid.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-A \code{ggplot2::ggplot()} object.
-The last plot can be saved to the file system using \code{ggplot2::ggsave()}.
-}
-\description{
-This function provides users with an easy way to graph
-(m)any network data for exploration, investigation, inspiration,
-and communication.
-
-It builds upon \code{{ggplot2}} and \code{{ggraph}} to offer
-pretty and extensible graphing solutions.
-However, compared to those solutions,
-\code{graphr()} contains various algorithms to provide better looking
-graphs by default.
-This means that just passing the function some network data
-will often be sufficient to return a reasonable-looking graph.
-
-The function also makes it easy to modify many of the most
-commonly adapted aspects of a graph,
-including node and edge size, colour, and shape,
-as arguments rather than additional functions that you need to remember.
-These can be defined outright, e.g. \code{node_size = 8}, or
-in reference to an attribute of the network, e.g. \code{node_size = "wealth"}.
-
-Lastly, \code{graphr()} uses \code{{ggplot2}}-related theme information, so
-it is easy to make colour palette and fonts institution-specific and consistent.
-See e.g. \code{theme_iheid()} for more.
-
-To learn more about what can be done visually,
-try \code{run_tute("Visualisation")}.
-}
-\examples{
-graphr(ison_adolescents)
-ison_adolescents \%>\%
- mutate(color = rep(c("introvert","extrovert"), times = 4),
- size = ifelse(node_is_cutpoint(ison_adolescents), 6, 3)) \%>\%
- mutate_ties(ecolor = rep(c("friends", "acquaintances"), times = 5)) \%>\%
- graphr(node_color = "color", node_size = "size",
- edge_size = 1.5, edge_color = "ecolor")
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_graphs.Rd b/man/map_graphs.Rd
deleted file mode 100644
index 549d1128..00000000
--- a/man/map_graphs.Rd
+++ /dev/null
@@ -1,60 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphs}
-\alias{map_graphs}
-\alias{graphs}
-\title{Easily graph a set of networks with sensible defaults}
-\usage{
-graphs(netlist, waves, based_on = c("first", "last", "both"), ...)
-}
-\arguments{
-\item{netlist}{A list of manynet-compatible networks.}
-
-\item{waves}{Numeric, the number of plots to be displayed side-by-side.
-If missing, the number of plots will be reduced to the first and last
-when there are more than four plots.
-This argument can also be passed a vector selecting the waves to plot.}
-
-\item{based_on}{Whether the layout of the joint plots should
-be based on the "first" or the "last" network, or "both".}
-
-\item{...}{Additional arguments passed to \code{graphr()}.}
-}
-\value{
-Multiple \code{ggplot2::ggplot()} objects displayed side-by-side.
-}
-\description{
-This function provides users with an easy way to graph
-lists of network data for comparison.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{patchwork}} package to plot the graphs
-side by side and, if necessary, in successive rows.
-This is useful for lists of networks that represent, for example,
-ego or component subgraphs of a network,
-or a list of a network's different types of tie or across time.
-By default just the first and last network will be plotted,
-but this can be overridden by the "waves" parameter.
-
-Where the graphs are of the same network (same nodes),
-the graphs may share a layout to facilitate comparison.
-By default, successive graphs will use the layout calculated for
-the "first" network, but other options include the "last" layout,
-or a mix, "both", of them.
-}
-\examples{
-#graphs(to_egos(ison_adolescents))
-#graphs(to_egos(ison_adolescents), waves = 8)
-#graphs(to_egos(ison_adolescents), waves = c(2, 4, 6))
-#graphs(play_diffusion(ison_adolescents))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_grapht.Rd b/man/map_grapht.Rd
deleted file mode 100644
index ff0cce5a..00000000
--- a/man/map_grapht.Rd
+++ /dev/null
@@ -1,137 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_grapht}
-\alias{map_grapht}
-\alias{grapht}
-\title{Easily animate dynamic networks with sensible defaults}
-\source{
-https://blog.schochastics.net/posts/2021-09-15_animating-network-evolutions-with-gganimate/
-}
-\usage{
-grapht(
- tlist,
- keep_isolates = TRUE,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{tlist}{The same migraph-compatible network listed according to
-a time attribute, waves, or slices.}
-
-\item{keep_isolates}{Logical, whether to keep isolate nodes in the graph.
-TRUE by default.
-If FALSE, removes nodes from each frame they are isolated in.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-Shows a .gif image. Assigning the result of the function
-saves the gif to a temporary folder and the object holds the path to this file.
-}
-\description{
-This function provides users with an easy way to graph
-dynamic network data for exploration and presentation.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{gganimate}} package to animate the changes
-between successive iterations of a network.
-This is useful for networks in which the ties and/or the node or tie
-attributes are changing.
-
-A progress bar is shown if it takes some time to encoding all the
-.png files into a .gif.
-}
-\examples{
-#ison_adolescents \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year", cumulative = TRUE) \%>\%
-# grapht()
-#ison_adolescents \%>\%
-# mutate(gender = rep(c("male", "female"), times = 4),
-# hair = rep(c("black", "brown"), times = 4),
-# age = sample(11:16, 8, replace = TRUE)) \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE),
-# links = sample(c("friends", "not_friends"), 10, replace = TRUE),
-# weekly_meetings = sample(c(3, 5, 7), 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year") \%>\%
-# grapht(layout = "concentric", membership = "gender",
-# node_shape = "gender", node_color = "hair",
-# node_size = "age", edge_color = "links",
-# edge_size = "weekly_meetings")
-#grapht(play_diffusion(ison_adolescents, seeds = 5))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_configuration.Rd b/man/map_layout_configuration.Rd
deleted file mode 100644
index 8f8e0f33..00000000
--- a/man/map_layout_configuration.Rd
+++ /dev/null
@@ -1,54 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_configuration}
-\alias{map_layout_configuration}
-\alias{layout_tbl_graph_configuration}
-\alias{layout_tbl_graph_dyad}
-\alias{layout_tbl_graph_triad}
-\alias{layout_tbl_graph_tetrad}
-\alias{layout_tbl_graph_pentad}
-\alias{layout_tbl_graph_hexad}
-\title{Layout algorithms based on configurational positions}
-\usage{
-layout_tbl_graph_configuration(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_dyad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_triad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_tetrad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_pentad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_hexad(.data, circular = FALSE, times = 1000)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-}
-\description{
-Configurational layouts locate nodes at symmetric coordinates
-to help illustrate particular configurations.
-Currently "triad" and "quad" layouts are available.
-The "configuration" layout will choose the appropriate configurational
-layout automatically.
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_partition.Rd b/man/map_layout_partition.Rd
deleted file mode 100644
index c30b378a..00000000
--- a/man/map_layout_partition.Rd
+++ /dev/null
@@ -1,130 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_partition}
-\alias{map_layout_partition}
-\alias{layout_tbl_graph_hierarchy}
-\alias{layout_tbl_graph_alluvial}
-\alias{layout_tbl_graph_railway}
-\alias{layout_tbl_graph_ladder}
-\alias{layout_tbl_graph_concentric}
-\alias{layout_tbl_graph_multilevel}
-\alias{layout_tbl_graph_lineage}
-\title{Layout algorithms based on bi- or other partitions}
-\source{
-Diego Diez, Andrew P. Hutchins and Diego Miranda-Saavedra. 2014.
-"Systematic identification of transcriptional regulatory modules from
-protein-protein interaction networks".
-\emph{Nucleic Acids Research}, 42 (1) e6.
-}
-\usage{
-layout_tbl_graph_hierarchy(
- .data,
- center = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_alluvial(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_railway(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_ladder(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_concentric(
- .data,
- membership,
- radius = NULL,
- order.by = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_multilevel(.data, level, circular = FALSE)
-
-layout_tbl_graph_lineage(.data, rank, circular = FALSE)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{center}{Further split "hierarchical" layouts by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name in hierarchy layout.
-Defaults to NULL.}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-
-\item{membership}{A node attribute or a vector to draw concentric circles
-for "concentric" layout.}
-
-\item{radius}{A vector of radii at which the concentric circles
-should be located for "concentric" layout.
-By default this is equal placement around an empty centre,
-unless one (the core) is a single node,
-in which case this node occupies the centre of the graph.}
-
-\item{order.by}{An attribute label indicating the (decreasing) order
-for the nodes around the circles for "concentric" layout.
-By default ordering is given by a bipartite placement that reduces
-the number of edge crossings.}
-
-\item{level}{A node attribute or a vector to hierarchically order levels for
-"multilevel" layout.}
-
-\item{rank}{A numerical node attribute to place nodes in Y axis
-according to values for "lineage" layout.}
-}
-\description{
-These algorithms layout networks based on two or more partitions,
-and are recommended for use with \code{graphr()} or \code{{ggraph}}.
-Note that these layout algorithms use \code{{Rgraphviz}},
-a package that is only available on Bioconductor.
-It will first need to be downloaded using \code{BiocManager::install("Rgraphviz")}.
-If it has not already been installed, there is a prompt the first time
-these functions are used though.
-
-The "hierarchy" layout layers the first node set along the bottom,
-and the second node set along the top,
-sequenced and spaced as necessary to minimise edge overlap.
-The "alluvial" layout is similar to "hierarchy",
-but places successive layers horizontally rather than vertically.
-The "railway" layout is similar to "hierarchy",
-but nodes are aligned across the layers.
-The "ladder" layout is similar to "railway",
-but places successive layers horizontally rather than vertically.
-The "concentric" layout places a "hierarchy" layout
-around a circle, with successive layers appearing as concentric circles.
-The "multilevel" layout places successive layers as multiple levels.
-The "lineage" layout ranks nodes in Y axis according to values.
-}
-\examples{
-#graphr(ison_southern_women, layout = "hierarchy", center = "events",
-# node_color = "type", node_size = 3)
-#graphr(ison_southern_women, layout = "alluvial")
-#graphr(ison_southern_women, layout = "concentric", membership = "type",
-# node_color = "type", node_size = 3)
-#graphr(ison_lotr, layout = "multilevel",
-# node_color = "Race", level = "Race", node_size = 3)
-# ison_adolescents \%>\%
-# mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2),
-# cut = node_is_cutpoint(ison_adolescents)) \%>\%
-# graphr(layout = "lineage", rank = "year", node_color = "cut",
-# node_size = migraph::node_degree(ison_adolescents)*10)
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}}
-}
-\concept{mapping}
diff --git a/man/map_palettes.Rd b/man/map_palettes.Rd
deleted file mode 100644
index 8c086ea9..00000000
--- a/man/map_palettes.Rd
+++ /dev/null
@@ -1,32 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_palettes.R
-\name{map_palettes}
-\alias{map_palettes}
-\alias{many_palettes}
-\title{Many palettes generator}
-\source{
-Adapted from \url{https://github.com/karthik/wesanderson/blob/master/R/colors.R}
-}
-\usage{
-many_palettes(palette, n, type = c("discrete", "continuous"))
-}
-\arguments{
-\item{palette}{Name of desired palette. Current choices are:
-\code{IHEID}, \code{Centres}, \code{SDGs}, \code{ETHZ}, \code{RUG},
-and \code{UZH}.}
-
-\item{n}{Number of colors desired. If omitted, uses all colours.}
-
-\item{type}{Either "continuous" or "discrete". Use continuous if you want
-to automatically interpolate between colours.}
-}
-\value{
-A graphic display of colours in palette.
-}
-\description{
-Many palettes generator
-}
-\examples{
-many_palettes()
-#many_palettes("IHEID")
-}
diff --git a/man/map_scales.Rd b/man/map_scales.Rd
deleted file mode 100644
index bdee1bc6..00000000
--- a/man/map_scales.Rd
+++ /dev/null
@@ -1,112 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_scales}
-\alias{map_scales}
-\alias{scale_fill_iheid}
-\alias{scale_colour_iheid}
-\alias{scale_color_iheid}
-\alias{scale_edge_colour_iheid}
-\alias{scale_edge_color_iheid}
-\alias{scale_fill_centres}
-\alias{scale_colour_centres}
-\alias{scale_color_centres}
-\alias{scale_edge_colour_centres}
-\alias{scale_edge_color_centres}
-\alias{scale_fill_sdgs}
-\alias{scale_colour_sdgs}
-\alias{scale_color_sdgs}
-\alias{scale_edge_colour_sdgs}
-\alias{scale_edge_color_sdgs}
-\alias{scale_fill_ethz}
-\alias{scale_colour_ethz}
-\alias{scale_color_ethz}
-\alias{scale_edge_colour_ethz}
-\alias{scale_edge_color_ethz}
-\alias{scale_fill_uzh}
-\alias{scale_colour_uzh}
-\alias{scale_color_uzh}
-\alias{scale_edge_colour_uzh}
-\alias{scale_edge_color_uzh}
-\alias{scale_fill_rug}
-\alias{scale_colour_rug}
-\alias{scale_color_rug}
-\alias{scale_edge_colour_rug}
-\alias{scale_edge_color_rug}
-\title{Many scales}
-\usage{
-scale_fill_iheid(direction = 1, ...)
-
-scale_colour_iheid(direction = 1, ...)
-
-scale_color_iheid(direction = 1, ...)
-
-scale_edge_colour_iheid(direction = 1, ...)
-
-scale_edge_color_iheid(direction = 1, ...)
-
-scale_fill_centres(direction = 1, ...)
-
-scale_colour_centres(direction = 1, ...)
-
-scale_color_centres(direction = 1, ...)
-
-scale_edge_colour_centres(direction = 1, ...)
-
-scale_edge_color_centres(direction = 1, ...)
-
-scale_fill_sdgs(direction = 1, ...)
-
-scale_colour_sdgs(direction = 1, ...)
-
-scale_color_sdgs(direction = 1, ...)
-
-scale_edge_colour_sdgs(direction = 1, ...)
-
-scale_edge_color_sdgs(direction = 1, ...)
-
-scale_fill_ethz(direction = 1, ...)
-
-scale_colour_ethz(direction = 1, ...)
-
-scale_color_ethz(direction = 1, ...)
-
-scale_edge_colour_ethz(direction = 1, ...)
-
-scale_edge_color_ethz(direction = 1, ...)
-
-scale_fill_uzh(direction = 1, ...)
-
-scale_colour_uzh(direction = 1, ...)
-
-scale_color_uzh(direction = 1, ...)
-
-scale_edge_colour_uzh(direction = 1, ...)
-
-scale_edge_color_uzh(direction = 1, ...)
-
-scale_fill_rug(direction = 1, ...)
-
-scale_colour_rug(direction = 1, ...)
-
-scale_color_rug(direction = 1, ...)
-
-scale_edge_colour_rug(direction = 1, ...)
-
-scale_edge_color_rug(direction = 1, ...)
-}
-\arguments{
-\item{direction}{Direction for using palette colors.}
-
-\item{...}{Extra arguments passed to \code{ggplot2::discrete_scale()}.}
-}
-\description{
-These functions enable to add color scales to be graphs.
-}
-\examples{
-#ison_brandes \%>\%
-#mutate(core = migraph::node_is_core(ison_brandes)) \%>\%
-#graphr(node_color = "core") +
-#scale_color_iheid()
-#graphr(ison_physicians[[1]], edge_color = "type") +
-#scale_edge_color_ethz()
-}
diff --git a/man/map_themes.Rd b/man/map_themes.Rd
deleted file mode 100644
index 288176ee..00000000
--- a/man/map_themes.Rd
+++ /dev/null
@@ -1,41 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_themes}
-\alias{map_themes}
-\alias{set_manynet_theme}
-\alias{theme_iheid}
-\alias{theme_ethz}
-\alias{theme_uzh}
-\alias{theme_rug}
-\title{Many themes}
-\usage{
-set_manynet_theme(theme = "default")
-
-theme_iheid(base_size = 12, base_family = "serif")
-
-theme_ethz(base_size = 12, base_family = "sans")
-
-theme_uzh(base_size = 12, base_family = "sans")
-
-theme_rug(base_size = 12, base_family = "mono")
-}
-\arguments{
-\item{theme}{String naming a theme.
-By default "default".}
-
-\item{base_size}{Font size, by default 12.}
-
-\item{base_family}{Font family, by default "sans".}
-}
-\description{
-These functions enable graphs to be easily and quickly themed,
-e.g. changing the default colour of the graph's vertices and edges.
-}
-\examples{
-to_mentoring(ison_brandes) \%>\%
- mutate(color = c(rep(c(1,2,3), 3), 3)) \%>\%
- graphr(node_color = "color") +
- labs(title = "Who leads and who follows?") +
- scale_color_iheid() +
- theme_iheid()
-}
diff --git a/man/mark_core.Rd b/man/mark_core.Rd
index 6143dee3..7f3c805e 100644
--- a/man/mark_core.Rd
+++ b/man/mark_core.Rd
@@ -4,6 +4,7 @@
\alias{mark_core}
\alias{node_is_universal}
\alias{node_is_core}
+\alias{node_kcoreness}
\alias{node_coreness}
\title{Core-periphery clustering algorithms}
\usage{
@@ -11,6 +12,8 @@ node_is_universal(.data)
node_is_core(.data, method = c("degree", "eigenvector"))
+node_kcoreness(.data)
+
node_coreness(.data)
}
\arguments{
@@ -33,8 +36,13 @@ can be added if there is interest.}
\description{
These functions identify nodes belonging to (some level of) the core of a network:
\itemize{
-\item \code{node_is_core()} assigns nodes to either the core or periphery.
-\item \code{node_coreness()} assigns nodes to their level of k-coreness.
+\item \code{node_is_universal()} identifies whether nodes are adjacent to all other
+nodes in the network.
+\item \code{node_is_core()} identifies whether nodes belong to the core of the
+network, as opposed to the periphery.
+\item \code{node_coreness()} returns a continuous measure of how closely each node
+resembles a typical core node.
+\item \code{node_kcoreness()} assigns nodes to their level of k-coreness.
}
}
\section{Universal/dominating node}{
@@ -68,6 +76,7 @@ node_is_core(ison_adolescents)
#ison_adolescents \%>\%
# mutate(corep = node_is_core()) \%>\%
# graphr(node_color = "corep")
+node_kcoreness(ison_adolescents)
node_coreness(ison_adolescents)
}
\references{
diff --git a/man/mark_ties.Rd b/man/mark_ties.Rd
index 8979a1aa..180bee9f 100644
--- a/man/mark_ties.Rd
+++ b/man/mark_ties.Rd
@@ -61,8 +61,8 @@ tie_is_reciprocated(ison_algebra)
tie_is_feedback(ison_algebra)
tie_is_bridge(ison_brandes)
ison_adolescents \%>\%
- mutate_ties(route = tie_is_path(from = "Jane", to = 7)) \%>\%
- graphr(edge_colour = "route")
+ mutate_ties(route = tie_is_path(from = "Jane", to = 7))
+ #graphr(edge_colour = "route")
}
\seealso{
Other marks:
diff --git a/man/mark_triangles.Rd b/man/mark_triangles.Rd
index aeb301cb..edb37409 100644
--- a/man/mark_triangles.Rd
+++ b/man/mark_triangles.Rd
@@ -52,23 +52,23 @@ are cohesively connected.
}
\examples{
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(tri = tie_is_triangular()) \%>\%
- graphr(edge_color = "tri")
+ mutate_ties(tri = tie_is_triangular())
+ #graphr(edge_color = "tri")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trans = tie_is_transitive()) \%>\%
- graphr(edge_color = "trans")
+ mutate_ties(trans = tie_is_transitive())
+ #graphr(edge_color = "trans")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trip = tie_is_triplet()) \%>\%
- graphr(edge_color = "trip")
+ mutate_ties(trip = tie_is_triplet())
+ #graphr(edge_color = "trip")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(cyc = tie_is_cyclical()) \%>\%
- graphr(edge_color = "cyc")
+ mutate_ties(cyc = tie_is_cyclical())
+ #graphr(edge_color = "cyc")
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(simmel = tie_is_simmelian()) \%>\%
- graphr(edge_color = "simmel")
+ mutate_ties(simmel = tie_is_simmelian())
+ #graphr(edge_color = "simmel")
generate_random(8, directed = TRUE) \%>\%
- mutate_ties(forbid = tie_is_forbidden()) \%>\%
- graphr(edge_color = "forbid")
+ mutate_ties(forbid = tie_is_forbidden())
+ #graphr(edge_color = "forbid")
tie_is_imbalanced(ison_marvel_relationships)
}
\seealso{
diff --git a/man/measure_central_between.Rd b/man/measure_central_between.Rd
index b50e1b57..35a86c42 100644
--- a/man/measure_central_between.Rd
+++ b/man/measure_central_between.Rd
@@ -107,9 +107,7 @@ nodes, and thus associated with bridging or spanning boundaries.
node_betweenness(ison_southern_women)
node_induced(ison_adolescents)
(tb <- tie_betweenness(ison_adolescents))
-plot(tb)
-ison_adolescents \%>\% mutate_ties(weight = tb) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = tb)
net_betweenness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_central_close.Rd b/man/measure_central_close.Rd
index ac2a6e3a..1992e688 100644
--- a/man/measure_central_close.Rd
+++ b/man/measure_central_close.Rd
@@ -176,9 +176,7 @@ removing that node would disconnect the network.
node_closeness(ison_southern_women)
node_reach(ison_adolescents)
(ec <- tie_closeness(ison_adolescents))
-plot(ec)
-ison_adolescents \%>\% mutate_ties(weight = ec) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = ec)
net_closeness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_features.Rd b/man/measure_features.Rd
index f119f997..05666148 100644
--- a/man/measure_features.Rd
+++ b/man/measure_features.Rd
@@ -14,7 +14,11 @@
\code{{signnet}} by David Schoch
}
\usage{
-net_core(.data, mark = NULL)
+net_core(
+ .data,
+ mark = NULL,
+ method = c("correlation", "ident", "ndiff", "diff")
+)
net_richclub(.data)
@@ -41,13 +45,6 @@ net_balance(.data)
\item{mark}{A logical vector the length of the nodes in the network.
This can be created by, among other things, any \verb{node_is_*()} function.}
-\item{membership}{A vector of partition membership.}
-
-\item{resolution}{A proportion indicating the resolution scale.
-By default 1, which returns the original definition of modularity.
-The higher this parameter, the more smaller communities will be privileged.
-The lower this parameter, the fewer larger communities are likely to be found.}
-
\item{method}{There are three small-world measures implemented:
\itemize{
\item "sigma" is the original equation from Watts and Strogatz (1998),
@@ -72,6 +69,13 @@ with the same dimensions.
but where there may not be a network for which \eqn{SWI = 1}.
}}
+\item{membership}{A vector of partition membership.}
+
+\item{resolution}{A proportion indicating the resolution scale.
+By default 1, which returns the original definition of modularity.
+The higher this parameter, the more smaller communities will be privileged.
+The lower this parameter, the fewer larger communities are likely to be found.}
+
\item{times}{Integer of number of simulations.}
}
\description{
@@ -100,6 +104,14 @@ ranging between \code{0} if all triangles are imbalanced and
These \verb{net_*()} functions return a single numeric scalar or value.
}
+\section{Core-Periphery}{
+
+\code{net_core()} calculates the Pearson correlation between the given network,
+where the nodes in the core are assigned by some given mark, and an ideal
+typical core-periphery network with the same number of nodes in the core
+and the periphery.
+}
+
\section{Modularity}{
Modularity measures the difference between the number of ties within each community
diff --git a/man/reexports.Rd b/man/reexports.Rd
index 8efbe88d..7cc10505 100644
--- a/man/reexports.Rd
+++ b/man/reexports.Rd
@@ -1,5 +1,5 @@
% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/reexports_classes.R, R/reexports_ggplot2.R
+% Please edit documentation in R/reexports_classes.R
\docType{import}
\name{reexports}
\alias{reexports}
@@ -10,15 +10,6 @@
\alias{.G}
\alias{.N}
\alias{.E}
-\alias{ggplot}
-\alias{ggtitle}
-\alias{guides}
-\alias{labs}
-\alias{xlab}
-\alias{ylab}
-\alias{aes}
-\alias{ggsave}
-\alias{scale_size}
\title{Objects exported from other packages}
\keyword{internal}
\description{
@@ -28,8 +19,6 @@ below to see their documentation.
\describe{
\item{dplyr}{\code{\link[dplyr:reexports]{\%>\%}}}
- \item{ggplot2}{\code{\link[ggplot2]{aes}}, \code{\link[ggplot2]{ggplot}}, \code{\link[ggplot2]{ggsave}}, \code{\link[ggplot2:labs]{ggtitle}}, \code{\link[ggplot2]{guides}}, \code{\link[ggplot2]{labs}}, \code{\link[ggplot2]{scale_size}}, \code{\link[ggplot2:labs]{xlab}}, \code{\link[ggplot2:labs]{ylab}}}
-
\item{igraph}{\code{\link[igraph]{is_igraph}}}
\item{tidygraph}{\code{\link[tidygraph:context_accessors]{.E}}, \code{\link[tidygraph:context_accessors]{.G}}, \code{\link[tidygraph:context_accessors]{.N}}, \code{\link[tidygraph:tbl_graph]{is.tbl_graph}}, \code{\link[tidygraph]{with_graph}}}
diff --git a/tests/testthat/test-manip_reformed.R b/tests/testthat/test-manip_reformed.R
index 5afe68dd..d0f77e1e 100644
--- a/tests/testthat/test-manip_reformed.R
+++ b/tests/testthat/test-manip_reformed.R
@@ -1,3 +1,15 @@
+match_net <- to_matching(ison_southern_women)
+
+test_that("to_matching works with two-mode networks", {
+ expect_true(net_ties(match_net) == min(net_dims(ison_southern_women)))
+})
+
+test_that("to_matching is idempotent", {
+ match_twice <- to_matching(match_net)
+ expect_equal(as_edgelist(match_net),
+ as_edgelist(match_twice))
+})
+
test_that("to_no_missing.tbl_graph removes nodes with missing values", {
# Create a tbl_graph with some missing values
graph <- tbl_graph(
diff --git a/tests/testthat/test-map_autograph.R b/tests/testthat/test-map_autograph.R
deleted file mode 100644
index e44590be..00000000
--- a/tests/testthat/test-map_autograph.R
+++ /dev/null
@@ -1,187 +0,0 @@
-test_that("unweighted, unsigned, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, undirected network
- test_brandes <- graphr(ison_brandes)
- # Node position
- expect_equal(round(test_brandes[["data"]][["x"]][[1]]), 3)
- expect_equal(round(test_brandes[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- # Node parameters
- expect_equal(round(test_brandes[["layers"]][[2]][["aes_params"]][["size"]]), 11)
- expect_equal(as.character(test_brandes[["layers"]][[2]][["aes_params"]][["shape"]]), "circle")
-})
-
-test_that("unweighted, signed, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, signed, undirected network
- test_marvel <- graphr(to_giant(ison_marvel_relationships))
- # Node position
- expect_equal(round(test_marvel[["data"]][["x"]][[1]]), -1)
- expect_equal(round(test_marvel[["data"]][["y"]][[1]]), 1)
- # Edge parameters
- expect_equal(test_marvel[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- # Node parameters
- expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["size"]], 3)
- #expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("unweighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, directed network
- test_algebra <- graphr(ison_algebra)
- # Node position
- expect_equal(round(test_algebra[["data"]][["x"]][[1]]), 0)
- expect_equal(round(test_algebra[["data"]][["y"]][[1]]), 0)
- # Edge parameters
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_algebra[["layers"]][[1]][["mapping"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_algebra[["layers"]][[2]][["aes_params"]][["size"]]), 8)
- expect_equal(test_algebra[["layers"]][[2]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("weighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # Weighted, unsigned, directed network
- test_networkers <- graphr(ison_networkers)
- # Node position
- expect_equal(round(test_networkers[["data"]][["x"]][[1]]), 9)
- expect_equal(round(test_networkers[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_networkers[["layers"]][[3]][["aes_params"]][["size"]]), 3)
- #expect_equal(test_networkers[["layers"]][[3]][["aes_params"]][["shape"]], "circle")
-})
-
-# Testing the node_color, node_size, and node_shape args by specifying a node attribute
-test_that("fancy node mods graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # one-mode network
- ison_marvel_relationships <- dplyr::mutate(ison_marvel_relationships,
- nodesize = Appearances/1000)
- testcolnodes <- graphr(ison_marvel_relationships, node_color = "Gender",
- node_size = "nodesize", node_shape = "Attractive")
- expect_s3_class(testcolnodes, c("ggraph","gg","ggplot"))
- expect_equal(round(testcolnodes$data$x[1]), 4)
- expect_equal(round(testcolnodes$data$y[1]), 3)
- expect_equal(nrow(testcolnodes[["plot_env"]][["lo"]]),
- c(net_nodes(ison_marvel_relationships)))
- # two-mode network
- ison_southern_women <- add_node_attribute(ison_southern_women, "group",
- c(sample(c("a", "b"),
- length(ison_southern_women),
- replace = TRUE)))
- test2 <- graphr(ison_southern_women, node_color = "type")
- expect_s3_class(test2, c("ggraph","gg","ggplot"))
- expect_equal(round(test2$data$x[1]), 0)
- expect_equal(round(test2$data$y[1]), 0)
- expect_equal(nrow(test2[["plot_env"]][["lo"]]),
- c(net_nodes(ison_southern_women)))
-})
-
-test_that("edge colours and edge size graph correctly", {
- skip_on_cran()
- ison_brandes2 <- ison_brandes %>%
- add_tie_attribute("tiecolour",
- c("A", "B", "A", "B", "B", "B", "B", "B", "B", "B", "B", "B")) %>%
- add_tie_attribute("weight", c(rep(1:6, 2)))
- test_brandes2 <- graphr(ison_brandes2, edge_color = "tiecolour", edge_size = "weight")
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_colour))
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_width))
-})
-
-# Named networks
-test_that("named networks plot correctly", {
- skip_on_cran()
- skip_on_ci()
- onemode <- graphr(ison_adolescents)
- twomode <- graphr(ison_southern_women)
- expect_equal(onemode[["data"]][["name"]], node_names(ison_adolescents))
- expect_equal(twomode[["data"]][["name"]], node_names(ison_southern_women))
-})
-
-# Test that autographr() works with arguments without quotes
-test_that("node_group works correctly", {
- skip_on_cran()
- testthat::skip_if_not_installed("concaveman")
- expect_equal(graphr(ison_lawfirm, node_group = gender),
- graphr(ison_lawfirm, node_group = "gender"))
-})
-
-test_that("unquoted arguments plot correctly", {
- skip_on_cran()
- expect_equal(graphr(ison_lawfirm, node_color = "gender"),
- graphr(ison_lawfirm, node_color = gender))
-})
-
-# Layouts
-test_that("concentric and circular layouts graph correctly", {
- skip_on_cran()
- test_circle <- graphr(to_giant(ison_marvel_relationships),
- layout = "circle")
- test_conc <- graphr(to_giant(ison_marvel_relationships),
- layout = "concentric", membership = "Gender")
- expect_equal(test_circle$plot_env$layout, "circle")
- expect_equal(test_conc$plot_env$layout, "concentric")
- expect_equal(eval(quote(pairlist(...)),
- envir = test_conc$plot_env)$membership,
- "Gender")
-})
-
-test_that("hierarchy and lineage layouts graph correctly", {
- skip_on_cran()
- skip_on_ci()
- test_lin <- ison_adolescents %>%
- mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2)) %>%
- graphr(layout = "lineage", rank = "year")
- test_hie <- graphr(ison_southern_women,
- layout = "hierarchy", center = "events")
- expect_equal(test_lin$plot_env$layout, "lineage")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_lin[["plot_env"]])[["rank"]]),
- "year")
- expect_equal(test_hie$plot_env$layout, "hierarchy")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_hie[["plot_env"]])[["center"]]),
- "events")
-})
-
-# test_that("graphr works for diff_model objects", {
-# skip_on_cran()
-# skip_on_ci()
-# test_diff <- graphr(play_diffusion(ison_brandes, old_version = TRUE))
-# if (inherits(test_diff$guides, "Guides")) {
-# expect_s3_class(test_diff[["guides"]][["guides"]][["shape"]], "GuideLegend")
-# expect_s3_class(test_diff[["guides"]][["guides"]][["colour"]], "GuideColourbar")
-# } else {
-# expect_equal(test_diff[["guides"]][["shape"]][["name"]], "legend")
-# expect_equal(test_diff[["guides"]][["colour"]][["name"]], "colorbar")
-# }
-# })
-
-test_that("concentric layout works when node names are missing", {
- skip_on_cran()
- skip_on_ci()
- llabel <- ison_southern_women %>%
- mutate(name = ifelse(type == TRUE, "", name)) %>%
- graphr(layout = "concentric")
- expect_true(any(llabel$data$name == ""))
-})
-
-test_that("hierarchy layout works for two mode networks", {
- skip_on_cran()
- skip_on_ci()
- tm <- ison_brandes %>%
- mutate(type = twomode_type, name = LETTERS[1:11]) %>%
- graphr()
- expect_length(unique(tm$data[tm$data$type == TRUE, "y"]), 1)
- expect_length(unique(tm$data[tm$data$type == FALSE, "y"]), 1)
-})
diff --git a/tests/testthat/test-map_theme.R b/tests/testthat/test-map_theme.R
deleted file mode 100644
index 6e6ffe6f..00000000
--- a/tests/testthat/test-map_theme.R
+++ /dev/null
@@ -1,58 +0,0 @@
-##### test themes
-test_that("themes graph correctly", {
- test_iheid <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_iheid()
- test_ethz <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_ethz()
- test_uzh <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_uzh()
- expect_equal(names(test_iheid[["theme"]][["title"]][["colour"]]), "IHEIDRed")
- expect_null(names(test_ethz[["theme"]][["title"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["title"]][["colour"]]), "UZH_Orange")
- expect_equal(names(test_iheid[["theme"]][["plot.subtitle"]][["colour"]]),
- "IHEIDGrey")
- expect_null(names(test_ethz[["theme"]][["plot.subtitle"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["plot.subtitle"]][["colour"]]),
- "UZH_Blue")
- expect_equal(test_iheid[["theme"]][["plot.caption"]][["family"]], "serif")
- expect_equal(test_ethz[["theme"]][["plot.caption"]][["family"]], "sans")
- expect_equal(test_uzh[["theme"]][["plot.caption"]][["family"]], "sans")
-})
-
-##### test scales
-test_that("scales graph correctly", {
- test_sdg <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3,4,5), 2), 1)) %>%
- graphr(node_color = color) +
- scale_color_sdgs()
- test_iheid <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 5), 3)) %>%
- graphr(node_color = color) +
- scale_color_iheid()
- test_ethz <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 4, 5)) %>%
- graphr(node_color = color) +
- scale_color_ethz()
- test_uzh <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 1, 2)) %>%
- graphr(node_color = color) +
- scale_color_uzh()
- test_rug <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 4), 1, 2, 1)) %>%
- graphr(node_color = color) +
- scale_color_rug()
- expect_equal(as.character(test_sdg[["scales"]][["scales"]][[2]][["call"]]), "scale_color_sdgs")
- expect_equal(as.character(test_iheid[["scales"]][["scales"]][[2]][["call"]]), "scale_color_iheid")
- expect_equal(as.character(test_ethz[["scales"]][["scales"]][[2]][["call"]]), "scale_color_ethz")
- expect_equal(as.character(test_uzh[["scales"]][["scales"]][[2]][["call"]]), "scale_color_uzh")
- expect_equal(as.character(test_rug[["scales"]][["scales"]][[2]][["call"]]), "scale_color_rug")
-})
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
-
## Marking
`{manynet}` includes four special groups of functions, each with their
diff --git a/cran-comments.md b/cran-comments.md
index 2827d039..634afb8c 100644
--- a/cran-comments.md
+++ b/cran-comments.md
@@ -9,4 +9,4 @@
0 errors | 0 warnings | 0 notes
-- Removed failing URLs from previous submission (v1.5.0)
\ No newline at end of file
+- Required for ggplot2 v4.0.0
\ No newline at end of file
diff --git a/man/defunct.Rd b/man/defunct.Rd
index 7258166b..1a5285e1 100644
--- a/man/defunct.Rd
+++ b/man/defunct.Rd
@@ -4,9 +4,6 @@
\alias{defunct}
\alias{pkg_data}
\alias{node_mode}
-\alias{autographr}
-\alias{autographs}
-\alias{autographd}
\alias{node_optimal}
\alias{node_kernighanlin}
\alias{node_edge_betweenness}
@@ -89,7 +86,6 @@
\alias{node_core}
\alias{node_by_quad}
\alias{net_by_quad}
-\alias{layout_tbl_graph_quad}
\alias{net_hazard}
\title{Functions that have been renamed, superseded, or are no longer working}
\usage{
@@ -97,34 +93,6 @@ pkg_data(pkg = "manynet")
node_mode(.data)
-autographr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- ...
-)
-
-autographs(netlist, waves, based_on = c("first", "last", "both"), ...)
-
-autographd(
- tlist,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- keep_isolates = TRUE,
- ...
-)
-
node_optimal(.data)
node_kernighanlin(.data)
@@ -295,8 +263,6 @@ node_by_quad(.data)
net_by_quad(.data)
-layout_tbl_graph_quad(.data, circular = FALSE, times = 1000)
-
net_hazard(.data)
}
\description{
@@ -314,12 +280,6 @@ wherever possible and update your scripts accordingly.
\item \code{node_mode()}: Deprecated on 2024-06-17.
-\item \code{autographr()}: Deprecated on 2024-06-17.
-
-\item \code{autographs()}: Deprecated on 2024-06-17.
-
-\item \code{autographd()}: Deprecated on 2024-06-17.
-
\item \code{node_optimal()}: Deprecated on 2024-06-14.
\item \code{node_kernighanlin()}: Deprecated on 2024-06-14.
@@ -486,8 +446,6 @@ wherever possible and update your scripts accordingly.
\item \code{net_by_quad()}: Deprecated on 2024-10-10.
-\item \code{layout_tbl_graph_quad()}: Deprecated on 2024-10-10.
-
\item \code{net_hazard()}: Deprecated on 2024-12-30.
}}
diff --git a/man/depth_first_recursive_search.Rd b/man/depth_first_recursive_search.Rd
deleted file mode 100644
index 9642a068..00000000
--- a/man/depth_first_recursive_search.Rd
+++ /dev/null
@@ -1,20 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{depth_first_recursive_search}
-\alias{depth_first_recursive_search}
-\title{Layouts for snapping layouts to a grid}
-\usage{
-depth_first_recursive_search(layout)
-}
-\description{
-The function uses approximate pattern matching
-to redistribute coarse layouts on square grid points, while
-preserving the topological relationships among the nodes (see Inoue et al. 2012).
-}
-\references{
-Inoue, Kentaro, Shinichi Shimozono, Hideaki Yoshida, and Hiroyuki Kurata. 2012.
-“Application of Approximate Pattern Matching in Two Dimensional Spaces to Grid Layout for Biochemical Network Maps” edited by J. Bourdon.
-\emph{PLoS ONE} 7(6):e37739.
-\doi{https://doi.org/10.1371/journal.pone.0037739}.
-}
-\keyword{internal}
diff --git a/man/make_cran.Rd b/man/make_cran.Rd
index b49d70b3..16557e6d 100644
--- a/man/make_cran.Rd
+++ b/man/make_cran.Rd
@@ -37,8 +37,6 @@ by \href{https://github.com/stocnet/manynet/issues}{raising an issue on Github}.
\examples{
# mnet <- read_cran()
# mnet <- to_ego(mnet, "manynet", max_dist = 2)
-# graphr(mnet, layout = "hierarchy",
-# edge_color = "type", node_color = "Compilation")
}
\seealso{
\link{as}
diff --git a/man/make_learning.Rd b/man/make_learning.Rd
index b06808d4..6f08083a 100644
--- a/man/make_learning.Rd
+++ b/man/make_learning.Rd
@@ -87,9 +87,6 @@ from other nodes whose beliefs are not too dissimilar from their own.
latticeEg <- add_node_attribute(latticeEg, "startValues", startValues)
latticeEg
play_segregation(latticeEg, "startValues", 0.5)
- # graphr(latticeEg, node_color = "startValues", node_size = 5) +
- # graphr(play_segregation(latticeEg, "startValues", 0.2),
- # node_color = "startValues", node_size = 5)
}
\references{
DeGroot, Morris H. 1974.
diff --git a/man/make_play.Rd b/man/make_play.Rd
index fe1b0621..7d1aa8c7 100644
--- a/man/make_play.Rd
+++ b/man/make_play.Rd
@@ -185,8 +185,6 @@ This can be used in in SEI, SEIS, SEIR, and SEIRS models.
\examples{
smeg <- generate_smallworld(15, 0.025)
- plot(play_diffusion(smeg, recovery = 0.4))
- #graphr(play_diffusion(ison_karateka))
}
\seealso{
Other makes:
diff --git a/man/make_random.Rd b/man/make_random.Rd
index 0146a541..e8471363 100644
--- a/man/make_random.Rd
+++ b/man/make_random.Rd
@@ -105,8 +105,8 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_random(12, 0.4))
-# graphr(generate_random(c(6, 6), 0.4))
+generate_random(12, 0.4)
+# generate_random(c(6, 6), 0.4)
}
\references{
\subsection{On random networks}{
diff --git a/man/make_stochastic.Rd b/man/make_stochastic.Rd
index f2930b77..0a977e6b 100644
--- a/man/make_stochastic.Rd
+++ b/man/make_stochastic.Rd
@@ -104,10 +104,10 @@ As an alternative, an existing network can be provided to \code{n}
and the number of modes, nodes, and directedness will be inferred.
}
\examples{
-graphr(generate_smallworld(12, 0.025))
-graphr(generate_smallworld(12, 0.25))
-graphr(generate_scalefree(12, 0.25))
-graphr(generate_scalefree(12, 1.25))
+generate_smallworld(12, 0.025)
+generate_smallworld(12, 0.25)
+generate_scalefree(12, 0.25)
+generate_scalefree(12, 1.25)
generate_fire(10)
generate_islands(10)
generate_citations(10)
diff --git a/man/manip_paths.Rd b/man/manip_paths.Rd
index f62c282e..ca35715c 100644
--- a/man/manip_paths.Rd
+++ b/man/manip_paths.Rd
@@ -109,10 +109,7 @@ This is, however, computationally slower.
\examples{
to_matching(ison_southern_women)
-#graphr(to_matching(ison_southern_women))
-graphr(to_mentoring(ison_adolescents))
to_eulerian(delete_nodes(ison_koenigsberg, "Lomse"))
- #graphr(to_eulerian(delete_nodes(ison_koenigsberg, "Lomse")))
}
\references{
\subsection{On matching}{
@@ -124,7 +121,7 @@ Gale, David, and Lloyd Stowell Shapley. 1962.
Goldberg, Andrew V., and Robert E. Tarjan. 1986.
"A new approach to the maximum flow problem".
-\emph{Proceedings of the eighteenth annual ACM symposium on Theory of computing – STOC '86}.
+\emph{Proceedings of the 18th Annual ACM Symposium on Theory of Computing}.
136-146.
\doi{10.1145/12130.12144}
}
diff --git a/man/manip_permutation.Rd b/man/manip_permutation.Rd
index 0578c343..32a6df8b 100644
--- a/man/manip_permutation.Rd
+++ b/man/manip_permutation.Rd
@@ -26,10 +26,6 @@ By default TRUE.}
on both the rows and columns (for a one-mode network)
or on each of the rows and columns (for a two-mode network).
}
-\examples{
-graphr(ison_adolescents, node_size = 4)
-graphr(to_permuted(ison_adolescents), node_size = 4)
-}
\seealso{
Other modifications:
\code{\link{manip_as}},
diff --git a/man/manip_project.Rd b/man/manip_project.Rd
index 0b59bb28..04f6e5ec 100644
--- a/man/manip_project.Rd
+++ b/man/manip_project.Rd
@@ -71,10 +71,7 @@ Below are the currently implemented S3 methods:\tabular{lrrrrr}{
\examples{
to_mode1(ison_southern_women)
to_mode2(ison_southern_women)
-#graphr(to_mode1(ison_southern_women))
-#graphr(to_mode2(ison_southern_women))
to_ties(ison_adolescents)
-#graphr(to_ties(ison_adolescents))
}
\seealso{
Other modifications:
diff --git a/man/manip_reformat.Rd b/man/manip_reformat.Rd
index 464f1716..17185089 100644
--- a/man/manip_reformat.Rd
+++ b/man/manip_reformat.Rd
@@ -70,7 +70,6 @@ This essentially has no effect on undirected networks or reciprocated ties.
}}
\examples{
to_anti(ison_southern_women)
-#graphr(to_anti(ison_southern_women))
}
\seealso{
Other modifications:
diff --git a/man/manip_ties.Rd b/man/manip_ties.Rd
index 531efcc0..c53695bf 100644
--- a/man/manip_ties.Rd
+++ b/man/manip_ties.Rd
@@ -83,7 +83,7 @@ Note that while \verb{add_*()}/\verb{delete_*()} functions operate similarly as
other <- create_filled(4) \%>\% mutate(name = c("A", "B", "C", "D"))
mutate_ties(other, form = 1:6) \%>\% filter_ties(form < 4)
add_tie_attribute(other, "weight", c(1, 2, 2, 2, 1, 2))
-ison_adolescents \%>\% add_ties(c("Betty","Tina")) \%>\% graphr()
+ison_adolescents \%>\% add_ties(c("Betty","Tina"))
delete_ties(ison_adolescents, 3)
delete_ties(ison_adolescents, "Alice|Sue")
}
diff --git a/man/map_graphr.Rd b/man/map_graphr.Rd
deleted file mode 100644
index 474f3d4b..00000000
--- a/man/map_graphr.Rd
+++ /dev/null
@@ -1,141 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphr}
-\alias{map_graphr}
-\alias{graphr}
-\title{Easily graph networks with sensible defaults}
-\usage{
-graphr(
- .data,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- node_group,
- edge_color,
- edge_size,
- snap = FALSE,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{.data}{A manynet-consistent object.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{node_group}{Node variable to be used for grouping the nodes.
-It is easiest if this is added as a hull over
-groups before plotting.
-Group variables should have a minimum of 3 nodes,
-if less, number groups will be reduced by
-merging categories with lower counts into one called "other".}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{snap}{Logical scalar, whether the layout should be snapped to a grid.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-A \code{ggplot2::ggplot()} object.
-The last plot can be saved to the file system using \code{ggplot2::ggsave()}.
-}
-\description{
-This function provides users with an easy way to graph
-(m)any network data for exploration, investigation, inspiration,
-and communication.
-
-It builds upon \code{{ggplot2}} and \code{{ggraph}} to offer
-pretty and extensible graphing solutions.
-However, compared to those solutions,
-\code{graphr()} contains various algorithms to provide better looking
-graphs by default.
-This means that just passing the function some network data
-will often be sufficient to return a reasonable-looking graph.
-
-The function also makes it easy to modify many of the most
-commonly adapted aspects of a graph,
-including node and edge size, colour, and shape,
-as arguments rather than additional functions that you need to remember.
-These can be defined outright, e.g. \code{node_size = 8}, or
-in reference to an attribute of the network, e.g. \code{node_size = "wealth"}.
-
-Lastly, \code{graphr()} uses \code{{ggplot2}}-related theme information, so
-it is easy to make colour palette and fonts institution-specific and consistent.
-See e.g. \code{theme_iheid()} for more.
-
-To learn more about what can be done visually,
-try \code{run_tute("Visualisation")}.
-}
-\examples{
-graphr(ison_adolescents)
-ison_adolescents \%>\%
- mutate(color = rep(c("introvert","extrovert"), times = 4),
- size = ifelse(node_is_cutpoint(ison_adolescents), 6, 3)) \%>\%
- mutate_ties(ecolor = rep(c("friends", "acquaintances"), times = 5)) \%>\%
- graphr(node_color = "color", node_size = "size",
- edge_size = 1.5, edge_color = "ecolor")
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_graphs.Rd b/man/map_graphs.Rd
deleted file mode 100644
index 549d1128..00000000
--- a/man/map_graphs.Rd
+++ /dev/null
@@ -1,60 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_graphs}
-\alias{map_graphs}
-\alias{graphs}
-\title{Easily graph a set of networks with sensible defaults}
-\usage{
-graphs(netlist, waves, based_on = c("first", "last", "both"), ...)
-}
-\arguments{
-\item{netlist}{A list of manynet-compatible networks.}
-
-\item{waves}{Numeric, the number of plots to be displayed side-by-side.
-If missing, the number of plots will be reduced to the first and last
-when there are more than four plots.
-This argument can also be passed a vector selecting the waves to plot.}
-
-\item{based_on}{Whether the layout of the joint plots should
-be based on the "first" or the "last" network, or "both".}
-
-\item{...}{Additional arguments passed to \code{graphr()}.}
-}
-\value{
-Multiple \code{ggplot2::ggplot()} objects displayed side-by-side.
-}
-\description{
-This function provides users with an easy way to graph
-lists of network data for comparison.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{patchwork}} package to plot the graphs
-side by side and, if necessary, in successive rows.
-This is useful for lists of networks that represent, for example,
-ego or component subgraphs of a network,
-or a list of a network's different types of tie or across time.
-By default just the first and last network will be plotted,
-but this can be overridden by the "waves" parameter.
-
-Where the graphs are of the same network (same nodes),
-the graphs may share a layout to facilitate comparison.
-By default, successive graphs will use the layout calculated for
-the "first" network, but other options include the "last" layout,
-or a mix, "both", of them.
-}
-\examples{
-#graphs(to_egos(ison_adolescents))
-#graphs(to_egos(ison_adolescents), waves = 8)
-#graphs(to_egos(ison_adolescents), waves = c(2, 4, 6))
-#graphs(play_diffusion(ison_adolescents))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_grapht.Rd b/man/map_grapht.Rd
deleted file mode 100644
index ff0cce5a..00000000
--- a/man/map_grapht.Rd
+++ /dev/null
@@ -1,137 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_autograph.R
-\name{map_grapht}
-\alias{map_grapht}
-\alias{grapht}
-\title{Easily animate dynamic networks with sensible defaults}
-\source{
-https://blog.schochastics.net/posts/2021-09-15_animating-network-evolutions-with-gganimate/
-}
-\usage{
-grapht(
- tlist,
- keep_isolates = TRUE,
- layout,
- labels = TRUE,
- node_color,
- node_shape,
- node_size,
- edge_color,
- edge_size,
- ...,
- node_colour,
- edge_colour
-)
-}
-\arguments{
-\item{tlist}{The same migraph-compatible network listed according to
-a time attribute, waves, or slices.}
-
-\item{keep_isolates}{Logical, whether to keep isolate nodes in the graph.
-TRUE by default.
-If FALSE, removes nodes from each frame they are isolated in.}
-
-\item{layout}{An igraph, ggraph, or manynet layout algorithm.
-If not declared, defaults to "triad" for networks with 3 nodes,
-"quad" for networks with 4 nodes,
-"stress" for all other one mode networks,
-or "hierarchy" for two mode networks.
-For "hierarchy" layout, one can further split graph by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name.
-For "concentric" layout algorithm please declare the "membership" as an
-extra argument.
-The "membership" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to draw concentric circles.
-For "multilevel" layout algorithm please declare the "level"
-as extra argument.
-The "level" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes to hierarchically
-order categories.
-If "level" is missing, function will look for 'lvl' node attribute in data.
-The "lineage" layout ranks nodes in Y axis according to values.
-For "lineage" layout algorithm please declare the "rank"
-as extra argument.
-The "rank" argument expects either a quoted node attribute present
-in data or vector with the same length as nodes.}
-
-\item{labels}{Logical, whether to print node names
-as labels if present.}
-
-\item{node_color, node_colour}{Node variable to be used for coloring the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be colored by declaring a color instead.}
-
-\item{node_shape}{Node variable to be used for shaping the nodes.
-It is easiest if this is added as a node attribute to
-the graph before plotting.
-Nodes can also be shaped by declaring a shape instead.}
-
-\item{node_size}{Node variable to be used for sizing the nodes.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all node-related statistics prior
-to using this function.
-Nodes can also be sized by declaring a numeric size or vector instead.}
-
-\item{edge_color, edge_colour}{Tie variable to be used for coloring the nodes.
-It is easiest if this is added as an edge or tie attribute
-to the graph before plotting.
-Edges can also be colored by declaring a color instead.}
-
-\item{edge_size}{Tie variable to be used for sizing the edges.
-This can be any continuous variable on the nodes of the network.
-Since this function expects this to be an existing variable,
-it is recommended to calculate all edge-related statistics prior
-to using this function.
-Edges can also be sized by declaring a numeric size or vector instead.}
-
-\item{...}{Extra arguments to pass on to the layout algorithm, if necessary.}
-}
-\value{
-Shows a .gif image. Assigning the result of the function
-saves the gif to a temporary folder and the object holds the path to this file.
-}
-\description{
-This function provides users with an easy way to graph
-dynamic network data for exploration and presentation.
-
-It builds upon this package's \code{graphr()} function,
-and inherits all the same features and arguments.
-See \code{graphr()} for more.
-However, it uses the \code{{gganimate}} package to animate the changes
-between successive iterations of a network.
-This is useful for networks in which the ties and/or the node or tie
-attributes are changing.
-
-A progress bar is shown if it takes some time to encoding all the
-.png files into a .gif.
-}
-\examples{
-#ison_adolescents \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year", cumulative = TRUE) \%>\%
-# grapht()
-#ison_adolescents \%>\%
-# mutate(gender = rep(c("male", "female"), times = 4),
-# hair = rep(c("black", "brown"), times = 4),
-# age = sample(11:16, 8, replace = TRUE)) \%>\%
-# mutate_ties(year = sample(1995:1998, 10, replace = TRUE),
-# links = sample(c("friends", "not_friends"), 10, replace = TRUE),
-# weekly_meetings = sample(c(3, 5, 7), 10, replace = TRUE)) \%>\%
-# to_waves(attribute = "year") \%>\%
-# grapht(layout = "concentric", membership = "gender",
-# node_shape = "gender", node_color = "hair",
-# node_size = "age", edge_color = "links",
-# edge_size = "weekly_meetings")
-#grapht(play_diffusion(ison_adolescents, seeds = 5))
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_layout_configuration}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_configuration.Rd b/man/map_layout_configuration.Rd
deleted file mode 100644
index 8f8e0f33..00000000
--- a/man/map_layout_configuration.Rd
+++ /dev/null
@@ -1,54 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_configuration}
-\alias{map_layout_configuration}
-\alias{layout_tbl_graph_configuration}
-\alias{layout_tbl_graph_dyad}
-\alias{layout_tbl_graph_triad}
-\alias{layout_tbl_graph_tetrad}
-\alias{layout_tbl_graph_pentad}
-\alias{layout_tbl_graph_hexad}
-\title{Layout algorithms based on configurational positions}
-\usage{
-layout_tbl_graph_configuration(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_dyad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_triad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_tetrad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_pentad(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_hexad(.data, circular = FALSE, times = 1000)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-}
-\description{
-Configurational layouts locate nodes at symmetric coordinates
-to help illustrate particular configurations.
-Currently "triad" and "quad" layouts are available.
-The "configuration" layout will choose the appropriate configurational
-layout automatically.
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_partition}}
-}
-\concept{mapping}
diff --git a/man/map_layout_partition.Rd b/man/map_layout_partition.Rd
deleted file mode 100644
index c30b378a..00000000
--- a/man/map_layout_partition.Rd
+++ /dev/null
@@ -1,130 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_layouts.R
-\name{map_layout_partition}
-\alias{map_layout_partition}
-\alias{layout_tbl_graph_hierarchy}
-\alias{layout_tbl_graph_alluvial}
-\alias{layout_tbl_graph_railway}
-\alias{layout_tbl_graph_ladder}
-\alias{layout_tbl_graph_concentric}
-\alias{layout_tbl_graph_multilevel}
-\alias{layout_tbl_graph_lineage}
-\title{Layout algorithms based on bi- or other partitions}
-\source{
-Diego Diez, Andrew P. Hutchins and Diego Miranda-Saavedra. 2014.
-"Systematic identification of transcriptional regulatory modules from
-protein-protein interaction networks".
-\emph{Nucleic Acids Research}, 42 (1) e6.
-}
-\usage{
-layout_tbl_graph_hierarchy(
- .data,
- center = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_alluvial(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_railway(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_ladder(.data, circular = FALSE, times = 1000)
-
-layout_tbl_graph_concentric(
- .data,
- membership,
- radius = NULL,
- order.by = NULL,
- circular = FALSE,
- times = 1000
-)
-
-layout_tbl_graph_multilevel(.data, level, circular = FALSE)
-
-layout_tbl_graph_lineage(.data, rank, circular = FALSE)
-}
-\arguments{
-\item{.data}{An object of a manynet-consistent class:
-\itemize{
-\item matrix (adjacency or incidence) from \code{{base}} R
-\item edgelist, a data frame from \code{{base}} R or tibble from \code{{tibble}}
-\item igraph, from the \code{{igraph}} package
-\item network, from the \code{{network}} package
-\item tbl_graph, from the \code{{tidygraph}} package
-}}
-
-\item{center}{Further split "hierarchical" layouts by
-declaring the "center" argument as the "events", "actors",
-or by declaring a node name in hierarchy layout.
-Defaults to NULL.}
-
-\item{circular}{Should the layout be transformed into a radial representation.
-Only possible for some layouts. Defaults to FALSE.}
-
-\item{times}{Maximum number of iterations, where appropriate}
-
-\item{membership}{A node attribute or a vector to draw concentric circles
-for "concentric" layout.}
-
-\item{radius}{A vector of radii at which the concentric circles
-should be located for "concentric" layout.
-By default this is equal placement around an empty centre,
-unless one (the core) is a single node,
-in which case this node occupies the centre of the graph.}
-
-\item{order.by}{An attribute label indicating the (decreasing) order
-for the nodes around the circles for "concentric" layout.
-By default ordering is given by a bipartite placement that reduces
-the number of edge crossings.}
-
-\item{level}{A node attribute or a vector to hierarchically order levels for
-"multilevel" layout.}
-
-\item{rank}{A numerical node attribute to place nodes in Y axis
-according to values for "lineage" layout.}
-}
-\description{
-These algorithms layout networks based on two or more partitions,
-and are recommended for use with \code{graphr()} or \code{{ggraph}}.
-Note that these layout algorithms use \code{{Rgraphviz}},
-a package that is only available on Bioconductor.
-It will first need to be downloaded using \code{BiocManager::install("Rgraphviz")}.
-If it has not already been installed, there is a prompt the first time
-these functions are used though.
-
-The "hierarchy" layout layers the first node set along the bottom,
-and the second node set along the top,
-sequenced and spaced as necessary to minimise edge overlap.
-The "alluvial" layout is similar to "hierarchy",
-but places successive layers horizontally rather than vertically.
-The "railway" layout is similar to "hierarchy",
-but nodes are aligned across the layers.
-The "ladder" layout is similar to "railway",
-but places successive layers horizontally rather than vertically.
-The "concentric" layout places a "hierarchy" layout
-around a circle, with successive layers appearing as concentric circles.
-The "multilevel" layout places successive layers as multiple levels.
-The "lineage" layout ranks nodes in Y axis according to values.
-}
-\examples{
-#graphr(ison_southern_women, layout = "hierarchy", center = "events",
-# node_color = "type", node_size = 3)
-#graphr(ison_southern_women, layout = "alluvial")
-#graphr(ison_southern_women, layout = "concentric", membership = "type",
-# node_color = "type", node_size = 3)
-#graphr(ison_lotr, layout = "multilevel",
-# node_color = "Race", level = "Race", node_size = 3)
-# ison_adolescents \%>\%
-# mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2),
-# cut = node_is_cutpoint(ison_adolescents)) \%>\%
-# graphr(layout = "lineage", rank = "year", node_color = "cut",
-# node_size = migraph::node_degree(ison_adolescents)*10)
-}
-\seealso{
-Other mapping:
-\code{\link{map_graphr}},
-\code{\link{map_graphs}},
-\code{\link{map_grapht}},
-\code{\link{map_layout_configuration}}
-}
-\concept{mapping}
diff --git a/man/map_palettes.Rd b/man/map_palettes.Rd
deleted file mode 100644
index 8c086ea9..00000000
--- a/man/map_palettes.Rd
+++ /dev/null
@@ -1,32 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_palettes.R
-\name{map_palettes}
-\alias{map_palettes}
-\alias{many_palettes}
-\title{Many palettes generator}
-\source{
-Adapted from \url{https://github.com/karthik/wesanderson/blob/master/R/colors.R}
-}
-\usage{
-many_palettes(palette, n, type = c("discrete", "continuous"))
-}
-\arguments{
-\item{palette}{Name of desired palette. Current choices are:
-\code{IHEID}, \code{Centres}, \code{SDGs}, \code{ETHZ}, \code{RUG},
-and \code{UZH}.}
-
-\item{n}{Number of colors desired. If omitted, uses all colours.}
-
-\item{type}{Either "continuous" or "discrete". Use continuous if you want
-to automatically interpolate between colours.}
-}
-\value{
-A graphic display of colours in palette.
-}
-\description{
-Many palettes generator
-}
-\examples{
-many_palettes()
-#many_palettes("IHEID")
-}
diff --git a/man/map_scales.Rd b/man/map_scales.Rd
deleted file mode 100644
index bdee1bc6..00000000
--- a/man/map_scales.Rd
+++ /dev/null
@@ -1,112 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_scales}
-\alias{map_scales}
-\alias{scale_fill_iheid}
-\alias{scale_colour_iheid}
-\alias{scale_color_iheid}
-\alias{scale_edge_colour_iheid}
-\alias{scale_edge_color_iheid}
-\alias{scale_fill_centres}
-\alias{scale_colour_centres}
-\alias{scale_color_centres}
-\alias{scale_edge_colour_centres}
-\alias{scale_edge_color_centres}
-\alias{scale_fill_sdgs}
-\alias{scale_colour_sdgs}
-\alias{scale_color_sdgs}
-\alias{scale_edge_colour_sdgs}
-\alias{scale_edge_color_sdgs}
-\alias{scale_fill_ethz}
-\alias{scale_colour_ethz}
-\alias{scale_color_ethz}
-\alias{scale_edge_colour_ethz}
-\alias{scale_edge_color_ethz}
-\alias{scale_fill_uzh}
-\alias{scale_colour_uzh}
-\alias{scale_color_uzh}
-\alias{scale_edge_colour_uzh}
-\alias{scale_edge_color_uzh}
-\alias{scale_fill_rug}
-\alias{scale_colour_rug}
-\alias{scale_color_rug}
-\alias{scale_edge_colour_rug}
-\alias{scale_edge_color_rug}
-\title{Many scales}
-\usage{
-scale_fill_iheid(direction = 1, ...)
-
-scale_colour_iheid(direction = 1, ...)
-
-scale_color_iheid(direction = 1, ...)
-
-scale_edge_colour_iheid(direction = 1, ...)
-
-scale_edge_color_iheid(direction = 1, ...)
-
-scale_fill_centres(direction = 1, ...)
-
-scale_colour_centres(direction = 1, ...)
-
-scale_color_centres(direction = 1, ...)
-
-scale_edge_colour_centres(direction = 1, ...)
-
-scale_edge_color_centres(direction = 1, ...)
-
-scale_fill_sdgs(direction = 1, ...)
-
-scale_colour_sdgs(direction = 1, ...)
-
-scale_color_sdgs(direction = 1, ...)
-
-scale_edge_colour_sdgs(direction = 1, ...)
-
-scale_edge_color_sdgs(direction = 1, ...)
-
-scale_fill_ethz(direction = 1, ...)
-
-scale_colour_ethz(direction = 1, ...)
-
-scale_color_ethz(direction = 1, ...)
-
-scale_edge_colour_ethz(direction = 1, ...)
-
-scale_edge_color_ethz(direction = 1, ...)
-
-scale_fill_uzh(direction = 1, ...)
-
-scale_colour_uzh(direction = 1, ...)
-
-scale_color_uzh(direction = 1, ...)
-
-scale_edge_colour_uzh(direction = 1, ...)
-
-scale_edge_color_uzh(direction = 1, ...)
-
-scale_fill_rug(direction = 1, ...)
-
-scale_colour_rug(direction = 1, ...)
-
-scale_color_rug(direction = 1, ...)
-
-scale_edge_colour_rug(direction = 1, ...)
-
-scale_edge_color_rug(direction = 1, ...)
-}
-\arguments{
-\item{direction}{Direction for using palette colors.}
-
-\item{...}{Extra arguments passed to \code{ggplot2::discrete_scale()}.}
-}
-\description{
-These functions enable to add color scales to be graphs.
-}
-\examples{
-#ison_brandes \%>\%
-#mutate(core = migraph::node_is_core(ison_brandes)) \%>\%
-#graphr(node_color = "core") +
-#scale_color_iheid()
-#graphr(ison_physicians[[1]], edge_color = "type") +
-#scale_edge_color_ethz()
-}
diff --git a/man/map_themes.Rd b/man/map_themes.Rd
deleted file mode 100644
index 288176ee..00000000
--- a/man/map_themes.Rd
+++ /dev/null
@@ -1,41 +0,0 @@
-% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/map_theme.R
-\name{map_themes}
-\alias{map_themes}
-\alias{set_manynet_theme}
-\alias{theme_iheid}
-\alias{theme_ethz}
-\alias{theme_uzh}
-\alias{theme_rug}
-\title{Many themes}
-\usage{
-set_manynet_theme(theme = "default")
-
-theme_iheid(base_size = 12, base_family = "serif")
-
-theme_ethz(base_size = 12, base_family = "sans")
-
-theme_uzh(base_size = 12, base_family = "sans")
-
-theme_rug(base_size = 12, base_family = "mono")
-}
-\arguments{
-\item{theme}{String naming a theme.
-By default "default".}
-
-\item{base_size}{Font size, by default 12.}
-
-\item{base_family}{Font family, by default "sans".}
-}
-\description{
-These functions enable graphs to be easily and quickly themed,
-e.g. changing the default colour of the graph's vertices and edges.
-}
-\examples{
-to_mentoring(ison_brandes) \%>\%
- mutate(color = c(rep(c(1,2,3), 3), 3)) \%>\%
- graphr(node_color = "color") +
- labs(title = "Who leads and who follows?") +
- scale_color_iheid() +
- theme_iheid()
-}
diff --git a/man/mark_core.Rd b/man/mark_core.Rd
index 6143dee3..7f3c805e 100644
--- a/man/mark_core.Rd
+++ b/man/mark_core.Rd
@@ -4,6 +4,7 @@
\alias{mark_core}
\alias{node_is_universal}
\alias{node_is_core}
+\alias{node_kcoreness}
\alias{node_coreness}
\title{Core-periphery clustering algorithms}
\usage{
@@ -11,6 +12,8 @@ node_is_universal(.data)
node_is_core(.data, method = c("degree", "eigenvector"))
+node_kcoreness(.data)
+
node_coreness(.data)
}
\arguments{
@@ -33,8 +36,13 @@ can be added if there is interest.}
\description{
These functions identify nodes belonging to (some level of) the core of a network:
\itemize{
-\item \code{node_is_core()} assigns nodes to either the core or periphery.
-\item \code{node_coreness()} assigns nodes to their level of k-coreness.
+\item \code{node_is_universal()} identifies whether nodes are adjacent to all other
+nodes in the network.
+\item \code{node_is_core()} identifies whether nodes belong to the core of the
+network, as opposed to the periphery.
+\item \code{node_coreness()} returns a continuous measure of how closely each node
+resembles a typical core node.
+\item \code{node_kcoreness()} assigns nodes to their level of k-coreness.
}
}
\section{Universal/dominating node}{
@@ -68,6 +76,7 @@ node_is_core(ison_adolescents)
#ison_adolescents \%>\%
# mutate(corep = node_is_core()) \%>\%
# graphr(node_color = "corep")
+node_kcoreness(ison_adolescents)
node_coreness(ison_adolescents)
}
\references{
diff --git a/man/mark_ties.Rd b/man/mark_ties.Rd
index 8979a1aa..180bee9f 100644
--- a/man/mark_ties.Rd
+++ b/man/mark_ties.Rd
@@ -61,8 +61,8 @@ tie_is_reciprocated(ison_algebra)
tie_is_feedback(ison_algebra)
tie_is_bridge(ison_brandes)
ison_adolescents \%>\%
- mutate_ties(route = tie_is_path(from = "Jane", to = 7)) \%>\%
- graphr(edge_colour = "route")
+ mutate_ties(route = tie_is_path(from = "Jane", to = 7))
+ #graphr(edge_colour = "route")
}
\seealso{
Other marks:
diff --git a/man/mark_triangles.Rd b/man/mark_triangles.Rd
index aeb301cb..edb37409 100644
--- a/man/mark_triangles.Rd
+++ b/man/mark_triangles.Rd
@@ -52,23 +52,23 @@ are cohesively connected.
}
\examples{
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(tri = tie_is_triangular()) \%>\%
- graphr(edge_color = "tri")
+ mutate_ties(tri = tie_is_triangular())
+ #graphr(edge_color = "tri")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trans = tie_is_transitive()) \%>\%
- graphr(edge_color = "trans")
+ mutate_ties(trans = tie_is_transitive())
+ #graphr(edge_color = "trans")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(trip = tie_is_triplet()) \%>\%
- graphr(edge_color = "trip")
+ mutate_ties(trip = tie_is_triplet())
+ #graphr(edge_color = "trip")
ison_adolescents \%>\% to_directed() \%>\%
- mutate_ties(cyc = tie_is_cyclical()) \%>\%
- graphr(edge_color = "cyc")
+ mutate_ties(cyc = tie_is_cyclical())
+ #graphr(edge_color = "cyc")
ison_monks \%>\% to_uniplex("like") \%>\%
- mutate_ties(simmel = tie_is_simmelian()) \%>\%
- graphr(edge_color = "simmel")
+ mutate_ties(simmel = tie_is_simmelian())
+ #graphr(edge_color = "simmel")
generate_random(8, directed = TRUE) \%>\%
- mutate_ties(forbid = tie_is_forbidden()) \%>\%
- graphr(edge_color = "forbid")
+ mutate_ties(forbid = tie_is_forbidden())
+ #graphr(edge_color = "forbid")
tie_is_imbalanced(ison_marvel_relationships)
}
\seealso{
diff --git a/man/measure_central_between.Rd b/man/measure_central_between.Rd
index b50e1b57..35a86c42 100644
--- a/man/measure_central_between.Rd
+++ b/man/measure_central_between.Rd
@@ -107,9 +107,7 @@ nodes, and thus associated with bridging or spanning boundaries.
node_betweenness(ison_southern_women)
node_induced(ison_adolescents)
(tb <- tie_betweenness(ison_adolescents))
-plot(tb)
-ison_adolescents \%>\% mutate_ties(weight = tb) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = tb)
net_betweenness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_central_close.Rd b/man/measure_central_close.Rd
index ac2a6e3a..1992e688 100644
--- a/man/measure_central_close.Rd
+++ b/man/measure_central_close.Rd
@@ -176,9 +176,7 @@ removing that node would disconnect the network.
node_closeness(ison_southern_women)
node_reach(ison_adolescents)
(ec <- tie_closeness(ison_adolescents))
-plot(ec)
-ison_adolescents \%>\% mutate_ties(weight = ec) \%>\%
- graphr()
+ison_adolescents \%>\% mutate_ties(weight = ec)
net_closeness(ison_southern_women, direction = "in")
}
\references{
diff --git a/man/measure_features.Rd b/man/measure_features.Rd
index f119f997..05666148 100644
--- a/man/measure_features.Rd
+++ b/man/measure_features.Rd
@@ -14,7 +14,11 @@
\code{{signnet}} by David Schoch
}
\usage{
-net_core(.data, mark = NULL)
+net_core(
+ .data,
+ mark = NULL,
+ method = c("correlation", "ident", "ndiff", "diff")
+)
net_richclub(.data)
@@ -41,13 +45,6 @@ net_balance(.data)
\item{mark}{A logical vector the length of the nodes in the network.
This can be created by, among other things, any \verb{node_is_*()} function.}
-\item{membership}{A vector of partition membership.}
-
-\item{resolution}{A proportion indicating the resolution scale.
-By default 1, which returns the original definition of modularity.
-The higher this parameter, the more smaller communities will be privileged.
-The lower this parameter, the fewer larger communities are likely to be found.}
-
\item{method}{There are three small-world measures implemented:
\itemize{
\item "sigma" is the original equation from Watts and Strogatz (1998),
@@ -72,6 +69,13 @@ with the same dimensions.
but where there may not be a network for which \eqn{SWI = 1}.
}}
+\item{membership}{A vector of partition membership.}
+
+\item{resolution}{A proportion indicating the resolution scale.
+By default 1, which returns the original definition of modularity.
+The higher this parameter, the more smaller communities will be privileged.
+The lower this parameter, the fewer larger communities are likely to be found.}
+
\item{times}{Integer of number of simulations.}
}
\description{
@@ -100,6 +104,14 @@ ranging between \code{0} if all triangles are imbalanced and
These \verb{net_*()} functions return a single numeric scalar or value.
}
+\section{Core-Periphery}{
+
+\code{net_core()} calculates the Pearson correlation between the given network,
+where the nodes in the core are assigned by some given mark, and an ideal
+typical core-periphery network with the same number of nodes in the core
+and the periphery.
+}
+
\section{Modularity}{
Modularity measures the difference between the number of ties within each community
diff --git a/man/reexports.Rd b/man/reexports.Rd
index 8efbe88d..7cc10505 100644
--- a/man/reexports.Rd
+++ b/man/reexports.Rd
@@ -1,5 +1,5 @@
% Generated by roxygen2: do not edit by hand
-% Please edit documentation in R/reexports_classes.R, R/reexports_ggplot2.R
+% Please edit documentation in R/reexports_classes.R
\docType{import}
\name{reexports}
\alias{reexports}
@@ -10,15 +10,6 @@
\alias{.G}
\alias{.N}
\alias{.E}
-\alias{ggplot}
-\alias{ggtitle}
-\alias{guides}
-\alias{labs}
-\alias{xlab}
-\alias{ylab}
-\alias{aes}
-\alias{ggsave}
-\alias{scale_size}
\title{Objects exported from other packages}
\keyword{internal}
\description{
@@ -28,8 +19,6 @@ below to see their documentation.
\describe{
\item{dplyr}{\code{\link[dplyr:reexports]{\%>\%}}}
- \item{ggplot2}{\code{\link[ggplot2]{aes}}, \code{\link[ggplot2]{ggplot}}, \code{\link[ggplot2]{ggsave}}, \code{\link[ggplot2:labs]{ggtitle}}, \code{\link[ggplot2]{guides}}, \code{\link[ggplot2]{labs}}, \code{\link[ggplot2]{scale_size}}, \code{\link[ggplot2:labs]{xlab}}, \code{\link[ggplot2:labs]{ylab}}}
-
\item{igraph}{\code{\link[igraph]{is_igraph}}}
\item{tidygraph}{\code{\link[tidygraph:context_accessors]{.E}}, \code{\link[tidygraph:context_accessors]{.G}}, \code{\link[tidygraph:context_accessors]{.N}}, \code{\link[tidygraph:tbl_graph]{is.tbl_graph}}, \code{\link[tidygraph]{with_graph}}}
diff --git a/tests/testthat/test-manip_reformed.R b/tests/testthat/test-manip_reformed.R
index 5afe68dd..d0f77e1e 100644
--- a/tests/testthat/test-manip_reformed.R
+++ b/tests/testthat/test-manip_reformed.R
@@ -1,3 +1,15 @@
+match_net <- to_matching(ison_southern_women)
+
+test_that("to_matching works with two-mode networks", {
+ expect_true(net_ties(match_net) == min(net_dims(ison_southern_women)))
+})
+
+test_that("to_matching is idempotent", {
+ match_twice <- to_matching(match_net)
+ expect_equal(as_edgelist(match_net),
+ as_edgelist(match_twice))
+})
+
test_that("to_no_missing.tbl_graph removes nodes with missing values", {
# Create a tbl_graph with some missing values
graph <- tbl_graph(
diff --git a/tests/testthat/test-map_autograph.R b/tests/testthat/test-map_autograph.R
deleted file mode 100644
index e44590be..00000000
--- a/tests/testthat/test-map_autograph.R
+++ /dev/null
@@ -1,187 +0,0 @@
-test_that("unweighted, unsigned, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, undirected network
- test_brandes <- graphr(ison_brandes)
- # Node position
- expect_equal(round(test_brandes[["data"]][["x"]][[1]]), 3)
- expect_equal(round(test_brandes[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_brandes[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- # Node parameters
- expect_equal(round(test_brandes[["layers"]][[2]][["aes_params"]][["size"]]), 11)
- expect_equal(as.character(test_brandes[["layers"]][[2]][["aes_params"]][["shape"]]), "circle")
-})
-
-test_that("unweighted, signed, undirected networks graph correctly", {
- skip_on_cran()
- # Unweighted, signed, undirected network
- test_marvel <- graphr(to_giant(ison_marvel_relationships))
- # Node position
- expect_equal(round(test_marvel[["data"]][["x"]][[1]]), -1)
- expect_equal(round(test_marvel[["data"]][["y"]][[1]]), 1)
- # Edge parameters
- expect_equal(test_marvel[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- # Node parameters
- expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["size"]], 3)
- #expect_equal(test_marvel[["layers"]][[4]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("unweighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- # Unweighted, unsigned, directed network
- test_algebra <- graphr(ison_algebra)
- # Node position
- expect_equal(round(test_algebra[["data"]][["x"]][[1]]), 0)
- expect_equal(round(test_algebra[["data"]][["y"]][[1]]), 0)
- # Edge parameters
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_alpha"]], 0.4)
- expect_equal(test_algebra[["layers"]][[1]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_algebra[["layers"]][[1]][["mapping"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_algebra[["layers"]][[2]][["aes_params"]][["size"]]), 8)
- expect_equal(test_algebra[["layers"]][[2]][["aes_params"]][["shape"]], "circle")
-})
-
-test_that("weighted, unsigned, directed networks graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # Weighted, unsigned, directed network
- test_networkers <- graphr(ison_networkers)
- # Node position
- expect_equal(round(test_networkers[["data"]][["x"]][[1]]), 9)
- expect_equal(round(test_networkers[["data"]][["y"]][[1]]), -1)
- # Edge parameters
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_alpha"]], 0.4)
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_linetype"]], "solid")
- #expect_equal(test_networkers[["layers"]][[2]][["aes_params"]][["edge_colour"]], "black")
- # Node parameters
- expect_equal(round(test_networkers[["layers"]][[3]][["aes_params"]][["size"]]), 3)
- #expect_equal(test_networkers[["layers"]][[3]][["aes_params"]][["shape"]], "circle")
-})
-
-# Testing the node_color, node_size, and node_shape args by specifying a node attribute
-test_that("fancy node mods graph correctly", {
- skip_on_cran()
- skip_on_ci()
- # one-mode network
- ison_marvel_relationships <- dplyr::mutate(ison_marvel_relationships,
- nodesize = Appearances/1000)
- testcolnodes <- graphr(ison_marvel_relationships, node_color = "Gender",
- node_size = "nodesize", node_shape = "Attractive")
- expect_s3_class(testcolnodes, c("ggraph","gg","ggplot"))
- expect_equal(round(testcolnodes$data$x[1]), 4)
- expect_equal(round(testcolnodes$data$y[1]), 3)
- expect_equal(nrow(testcolnodes[["plot_env"]][["lo"]]),
- c(net_nodes(ison_marvel_relationships)))
- # two-mode network
- ison_southern_women <- add_node_attribute(ison_southern_women, "group",
- c(sample(c("a", "b"),
- length(ison_southern_women),
- replace = TRUE)))
- test2 <- graphr(ison_southern_women, node_color = "type")
- expect_s3_class(test2, c("ggraph","gg","ggplot"))
- expect_equal(round(test2$data$x[1]), 0)
- expect_equal(round(test2$data$y[1]), 0)
- expect_equal(nrow(test2[["plot_env"]][["lo"]]),
- c(net_nodes(ison_southern_women)))
-})
-
-test_that("edge colours and edge size graph correctly", {
- skip_on_cran()
- ison_brandes2 <- ison_brandes %>%
- add_tie_attribute("tiecolour",
- c("A", "B", "A", "B", "B", "B", "B", "B", "B", "B", "B", "B")) %>%
- add_tie_attribute("weight", c(rep(1:6, 2)))
- test_brandes2 <- graphr(ison_brandes2, edge_color = "tiecolour", edge_size = "weight")
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_colour))
- expect_false(is.null(test_brandes2$layers[[1]]$mapping$edge_width))
-})
-
-# Named networks
-test_that("named networks plot correctly", {
- skip_on_cran()
- skip_on_ci()
- onemode <- graphr(ison_adolescents)
- twomode <- graphr(ison_southern_women)
- expect_equal(onemode[["data"]][["name"]], node_names(ison_adolescents))
- expect_equal(twomode[["data"]][["name"]], node_names(ison_southern_women))
-})
-
-# Test that autographr() works with arguments without quotes
-test_that("node_group works correctly", {
- skip_on_cran()
- testthat::skip_if_not_installed("concaveman")
- expect_equal(graphr(ison_lawfirm, node_group = gender),
- graphr(ison_lawfirm, node_group = "gender"))
-})
-
-test_that("unquoted arguments plot correctly", {
- skip_on_cran()
- expect_equal(graphr(ison_lawfirm, node_color = "gender"),
- graphr(ison_lawfirm, node_color = gender))
-})
-
-# Layouts
-test_that("concentric and circular layouts graph correctly", {
- skip_on_cran()
- test_circle <- graphr(to_giant(ison_marvel_relationships),
- layout = "circle")
- test_conc <- graphr(to_giant(ison_marvel_relationships),
- layout = "concentric", membership = "Gender")
- expect_equal(test_circle$plot_env$layout, "circle")
- expect_equal(test_conc$plot_env$layout, "concentric")
- expect_equal(eval(quote(pairlist(...)),
- envir = test_conc$plot_env)$membership,
- "Gender")
-})
-
-test_that("hierarchy and lineage layouts graph correctly", {
- skip_on_cran()
- skip_on_ci()
- test_lin <- ison_adolescents %>%
- mutate(year = rep(c(1985, 1990, 1995, 2000), times = 2)) %>%
- graphr(layout = "lineage", rank = "year")
- test_hie <- graphr(ison_southern_women,
- layout = "hierarchy", center = "events")
- expect_equal(test_lin$plot_env$layout, "lineage")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_lin[["plot_env"]])[["rank"]]),
- "year")
- expect_equal(test_hie$plot_env$layout, "hierarchy")
- expect_equal((eval(quote(pairlist(...)),
- envir = test_hie[["plot_env"]])[["center"]]),
- "events")
-})
-
-# test_that("graphr works for diff_model objects", {
-# skip_on_cran()
-# skip_on_ci()
-# test_diff <- graphr(play_diffusion(ison_brandes, old_version = TRUE))
-# if (inherits(test_diff$guides, "Guides")) {
-# expect_s3_class(test_diff[["guides"]][["guides"]][["shape"]], "GuideLegend")
-# expect_s3_class(test_diff[["guides"]][["guides"]][["colour"]], "GuideColourbar")
-# } else {
-# expect_equal(test_diff[["guides"]][["shape"]][["name"]], "legend")
-# expect_equal(test_diff[["guides"]][["colour"]][["name"]], "colorbar")
-# }
-# })
-
-test_that("concentric layout works when node names are missing", {
- skip_on_cran()
- skip_on_ci()
- llabel <- ison_southern_women %>%
- mutate(name = ifelse(type == TRUE, "", name)) %>%
- graphr(layout = "concentric")
- expect_true(any(llabel$data$name == ""))
-})
-
-test_that("hierarchy layout works for two mode networks", {
- skip_on_cran()
- skip_on_ci()
- tm <- ison_brandes %>%
- mutate(type = twomode_type, name = LETTERS[1:11]) %>%
- graphr()
- expect_length(unique(tm$data[tm$data$type == TRUE, "y"]), 1)
- expect_length(unique(tm$data[tm$data$type == FALSE, "y"]), 1)
-})
diff --git a/tests/testthat/test-map_theme.R b/tests/testthat/test-map_theme.R
deleted file mode 100644
index 6e6ffe6f..00000000
--- a/tests/testthat/test-map_theme.R
+++ /dev/null
@@ -1,58 +0,0 @@
-##### test themes
-test_that("themes graph correctly", {
- test_iheid <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_iheid()
- test_ethz <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_ethz()
- test_uzh <- graphr(to_mentoring(ison_brandes)) +
- labs(title = "Who leads and who follows?",
- subtitle = "A mentoring network",
- caption = "ison_brandes network") +
- theme_uzh()
- expect_equal(names(test_iheid[["theme"]][["title"]][["colour"]]), "IHEIDRed")
- expect_null(names(test_ethz[["theme"]][["title"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["title"]][["colour"]]), "UZH_Orange")
- expect_equal(names(test_iheid[["theme"]][["plot.subtitle"]][["colour"]]),
- "IHEIDGrey")
- expect_null(names(test_ethz[["theme"]][["plot.subtitle"]][["colour"]]))
- expect_equal(names(test_uzh[["theme"]][["plot.subtitle"]][["colour"]]),
- "UZH_Blue")
- expect_equal(test_iheid[["theme"]][["plot.caption"]][["family"]], "serif")
- expect_equal(test_ethz[["theme"]][["plot.caption"]][["family"]], "sans")
- expect_equal(test_uzh[["theme"]][["plot.caption"]][["family"]], "sans")
-})
-
-##### test scales
-test_that("scales graph correctly", {
- test_sdg <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3,4,5), 2), 1)) %>%
- graphr(node_color = color) +
- scale_color_sdgs()
- test_iheid <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 5), 3)) %>%
- graphr(node_color = color) +
- scale_color_iheid()
- test_ethz <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 4, 5)) %>%
- graphr(node_color = color) +
- scale_color_ethz()
- test_uzh <- ison_brandes %>%
- mutate(color = c(rep(c(1,2,3), 3), 1, 2)) %>%
- graphr(node_color = color) +
- scale_color_uzh()
- test_rug <- ison_brandes %>%
- mutate(color = c(rep(c(1,2), 4), 1, 2, 1)) %>%
- graphr(node_color = color) +
- scale_color_rug()
- expect_equal(as.character(test_sdg[["scales"]][["scales"]][[2]][["call"]]), "scale_color_sdgs")
- expect_equal(as.character(test_iheid[["scales"]][["scales"]][[2]][["call"]]), "scale_color_iheid")
- expect_equal(as.character(test_ethz[["scales"]][["scales"]][[2]][["call"]]), "scale_color_ethz")
- expect_equal(as.character(test_uzh[["scales"]][["scales"]][[2]][["call"]]), "scale_color_uzh")
- expect_equal(as.character(test_rug[["scales"]][["scales"]][[2]][["call"]]), "scale_color_rug")
-})