-
Notifications
You must be signed in to change notification settings - Fork 1
Update documentation #80
New issue
Have a question about this project? Sign up for a free GitHub account to open an issue and contact its maintainers and the community.
By clicking “Sign up for GitHub”, you agree to our terms of service and privacy statement. We’ll occasionally send you account related emails.
Already on GitHub? Sign in to your account
Merged
Merged
Changes from 11 commits
Commits
Show all changes
14 commits
Select commit
Hold shift + click to select a range
f8f566a
Update contributing guide to use conda instead of poetry
ataha24 cdc037a
Revise installation guide with improved structure and comparison table
ataha24 dbd0171
Add detailed Conda-based installation and usage instructions
ataha24 d018c6d
Document Apptainer usage for running AutoAFIDs on HPC systems
ataha24 62cfcbc
Add step-by-step Docker instructions for Windows users
ataha24 ef8fe6a
update readme
ataha24 90b9cbe
add small toc
ataha24 9d565bc
remove emojis for hyperlinking in RTD
ataha24 ef4e457
update readme w/ more accurate content
ataha24 edbb78e
add templates for issues and PRs
Dhananjhay 5ce5f49
Merge pull request #81 from afids/djay/update-documentation
Dhananjhay 8cc47ae
fix synthsr comment
ataha24 b8463a9
remove synthsr repo mention and fix contributing page
Dhananjhay ec41a4d
remove git comment
Dhananjhay File filter
Filter by extension
Conversations
Failed to load comments.
Loading
Jump to
Jump to file
Failed to load files.
Loading
Diff view
Diff view
There are no files selected for viewing
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,8 @@ | ||
| ## The problem | ||
|
|
||
| Briefly describe the issue you are experiencing (or feature you want to see added). Tell us what you were trying to do and what happened instead. If you have any helpful screenshots or a file you tried to upload, please include them! Please also select an appropriate label (to the right). | ||
|
|
||
| ## Environment | ||
|
|
||
| - Operating System | ||
| - Autoafids version |
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -0,0 +1,26 @@ | ||
| ## Proposed changes | ||
|
|
||
| Describe the changes implemented in this pull request. If the changes fix a bug or resolves a feature request, be sure to link the issue. Please explain the issue and how the changes address the issue. | ||
|
|
||
| ## Types of changes | ||
|
|
||
| What types of changes does your code introduce? Put an `x` in the boxes that apply | ||
|
|
||
| - [ ] Bugfix (non-breaking change which fixes an issue) | ||
| - [ ] New feature (non-breaking change which adds functionalitiy) | ||
| - [ ] Breaking change (fix or feature that would cause existing functionality to not work as expected) | ||
| - [ ] Other (if none of the other choices apply) | ||
|
|
||
| ## Checklist | ||
|
|
||
| Put an `x` in the boxes that apply. You can also fill these out after creating the PR. If you are unsure about any of the choices, don't hesitate to ask! | ||
|
|
||
| - [ ] Changes have been tested to ensure that fix is effective or that a feature works. | ||
| - [ ] Changes pass the unit tests | ||
| - [ ] Changes pass the wet run tests | ||
| - [ ] Code has been run through the `poe quality` task | ||
| - [ ] I have included necessary documentation or comments (as necessary) | ||
| - [ ] Any dependent changes have been merged and published | ||
|
|
||
| ## Notes | ||
| All PRs will undergo unit and wet run testing before being reviewed. You may be requested to explain or make additional changes before the PR is accepted. |
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -1,36 +1,123 @@ | ||
| # Automatic Anatomical Fiducials (AutoAFIDs) | ||
|
|
||
| [](https://autoafids.readthedocs.io/en/stable/?badge=stable) | ||
|  | ||
|  | ||
| [](https://github.com/afids/autoafids/actions/workflows/lint-and-dryrun-testing.yml?query=branch%3Amain) | ||
|  | ||
|  | ||
|
|
||
| Developed by the AIMS Lab at the Robarts Research Institute | ||
| *2023–2025* | ||
|
|
||
| > ⚠️ This package is under active development. While stable and reproducible, users are encouraged to report any bugs or unexpected behavior to the development team. | ||
|
|
||
| --- | ||
|
|
||
| ## 📑 Table of Contents | ||
|
|
||
| - [🧠 What is AutoAFIDs?](#what-is-autoafids) | ||
| - [⚙️ Workflow Overview](#workflow-overview) | ||
| - [🔍 Known Issues](#known-issues) | ||
| - [📖 Full Documentation](#full-documentation) | ||
| - [💬 Questions, Issues, and Feedback](#questions-issues-and-feedback) | ||
| - [📚 Relevant Papers](#relevant-papers) | ||
|
|
||
| --- | ||
| ## What is AutoAFIDs? | ||
|
|
||
| **AutoAFIDs** is a BIDS App for automatic detection of anatomical fiducials (AFIDs) on MRI scans. We make use of these AFIDs in various neuroimaging applications such as image registration, quality control, and neurosurgical targeting. | ||
|
|
||
| AutoAFIDs leverages: | ||
|
|
||
| AIMS Lab Research Team at the Robarts Research Institute - 2023-2025 | ||
| - A 3D [U-Net architecture](https://arxiv.org/abs/1505.04597) for landmark localization | ||
| - The [Snakemake](https://snakemake.readthedocs.io/) and [SnakeBIDS](https://snakebids.readthedocs.io/en/stable/) workflow frameworks for reproducible processing | ||
| - The AFIDs protocol developed by the [AFIDs community](https://github.com/afids) | ||
|
|
||
| *This package is under active development. It should be stable and reproducible, but please let any of the active contributing members know if there are any bugs or unusual behaviour.* | ||
| The software is modality-aware, but best supports T1-weighted MRI scans. It also includes QC visualizations for quality assurance. | ||
|
|
||
| This Python package is a standard 3D [U-Net](https://arxiv.org/abs/1505.04597) (Ronneberger et al. 2015) machine learning model based on Snakemake and SnakeBIDS workflow management tools that leverages the recent release of the anatomical fiducial framework to solve the landmark regression problem on 3D MRI images. It is currently in development phase and contains tunable parameters that are not normally exposed in most other machine learning models; the user is highly advised to get familiar with the above mentioned workflow managaments tools and read docstrings and relevant documentation before using this software. Please see the [changelog](CHANGELOG.md) for more details. | ||
| --- | ||
|
|
||
| ## Workflow | ||
| ## Workflow Overview | ||
|
|
||
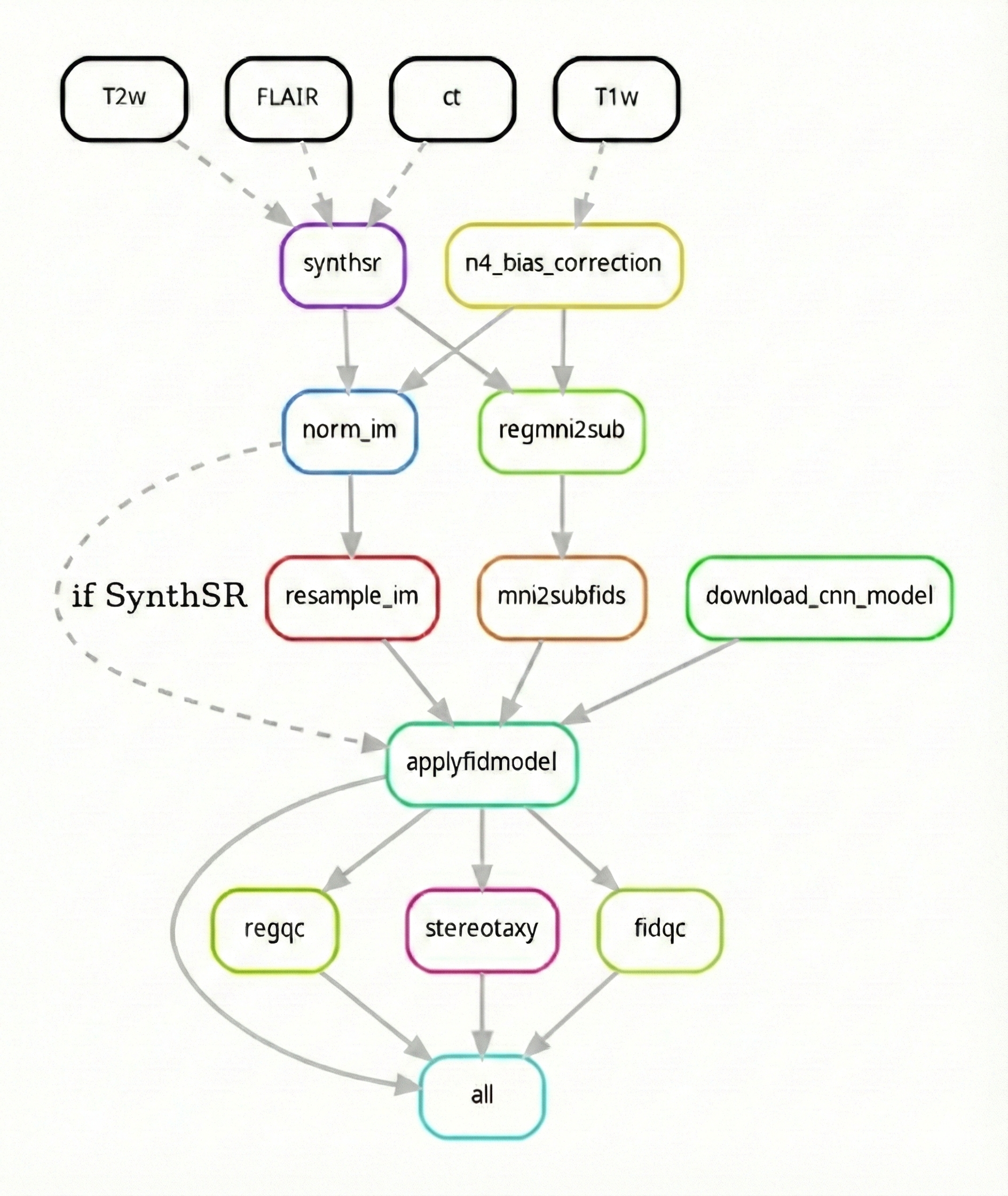
| A brief summary of the workflow can be found below along with its Directed Acyclic Graph (DAG) (see documentation for a detailed summary): | ||
| Below is a high-level summary of the AutoAFIDs processing pipeline: | ||
|
|
||
|  | ||
|
|
||
| 1. Preprocess input NIfTI files based on image modality | ||
| 2. Download and apply the fiducial model | ||
| 1. Preprocess BIDS input files based on image modality (T1w, T2w) | ||
| 2. Load trained fiducial models (32 AFID points) | ||
| 3. Run patch-based U-Net inference per AFID | ||
| 4. Generate predictions | ||
|
|
||
| ## Processing landmark data (AFIDs) | ||
| 1. Extract fiducial points from the landmark files (.fcsv is supported) | ||
| 2. Generate a landmark Euclidean distance/probability map with each voxel communicating distance to an AFID of interest | ||
| --- | ||
|
|
||
| ## Known Issues | ||
| - Factorize apply workflow to run per landmark of interest | ||
|
|
||
| ### **Full documentation:** [here](https://autoafids.readthedocs.io/en/) | ||
| - `gen_fcsv` rule is currently sequential and may benefit from AFID-level parallelization | ||
| - T1w-like scan synthesis is available (via SynthSR [Iglesias et al., 2023](https://www-science-org.proxy1.lib.uwo.ca/doi/10.1126/sciadv.add3607)) but requires millimetric validation and may not work for all operating systems | ||
|
|
||
| --- | ||
|
|
||
| ## Full Documentation | ||
|
|
||
| 👉 [autoafids.readthedocs.io](https://autoafids.readthedocs.io/en/) | ||
|
|
||
| Includes installation instructions, usage examples, and advanced configuration. | ||
|
|
||
| --- | ||
|
|
||
| ## Questions, Issues, and Feedback | ||
|
|
||
| - Open an issue on the [GitHub issues page](https://github.com/afids/autoafids/issues) | ||
| - Start a conversation on the [Discussions board](https://github.com/afids/autoafids/discussions) | ||
| - Contact: [[email protected]](mailto:[email protected]) or [[email protected]](mailto:[email protected]) | ||
|
|
||
| We welcome feedback, contributions, and collaboration! | ||
|
|
||
|
|
||
| ## Relevant Papers | ||
|
|
||
| AutoAFIDs builds upon a series of foundational works that introduce, validate, and apply the anatomical fiducials (AFIDs) framework for neuroimaging quality control, registration evaluation, and surgical planning. | ||
|
|
||
| --- | ||
|
|
||
| ### Methodological Foundations | ||
|
|
||
| - **Lau et al., 2019** | ||
| *A framework for evaluating correspondence between brain images using anatomical fiducials.* | ||
| *Human Brain Mapping*, 40(14), 4163–4179. | ||
| [DOI: 10.1002/hbm.24695](https://doi.org/10.1002/hbm.24695) | ||
|
|
||
| --- | ||
|
|
||
| ### Clinical Applications | ||
|
|
||
| - **Abbass et al., 2022** | ||
| *Application of the anatomical fiducials framework to a clinical dataset of patients with Parkinson’s disease.* | ||
| *Brain Structure & Function*, 227(1), 393–405. | ||
| [DOI: 10.1007/s00429-021-02408-3](https://doi-org.proxy1.lib.uwo.ca/10.1007/s00429-021-02408-3) | ||
|
|
||
| - **Taha et al., 2022** | ||
| *An indirect deep brain stimulation targeting tool using salient anatomical fiducials.* | ||
| *Neuromodulation: Journal of the International Neuromodulation Society*, 25(8), S6–S7. | ||
|
|
||
| --- | ||
|
|
||
| ### Dataset & Resource Publication | ||
|
|
||
| - **Taha et al., 2023** | ||
| *Magnetic resonance imaging datasets with anatomical fiducials for quality control and registration.* | ||
| *Scientific Data*, 10(1), 449. | ||
| [DOI: 10.1038/s41597-023-02330-9](https://doi.org/10.1038/s41597-023-02330-9) | ||
|
|
||
| --- | ||
|
|
||
| ### Registration and Localization Accuracy | ||
|
|
||
| - **Abbass et al., 2025** | ||
| *The impact of localization and registration accuracy on estimates of deep brain stimulation electrode position in stereotactic space.* | ||
| *Imaging Neuroscience.* | ||
| [DOI: 10.1162/imag_a_00579](https://doi.org/10.1162/imag_a_00579) | ||
|
|
||
| ## Revalent Papers: | ||
| * Link relavent papers to autoafids here | ||
| --- | ||
|
|
||
| ## Questions, Issues, Suggestions, and Other Feedback | ||
| Please reach out if you have any questions, suggestions, or other feedback related to this software—either through email ([email protected]) or the discussions page. Larger issues or feature requests can be posted and tracked via the issues page. Finally, you can also reach out to Alaa Taha, the Science Lead. | ||
| > 📌 If you use AutoAFIDs or AFIDs data in your research, please cite the relevant papers above. |
This file contains hidden or bidirectional Unicode text that may be interpreted or compiled differently than what appears below. To review, open the file in an editor that reveals hidden Unicode characters.
Learn more about bidirectional Unicode characters
| Original file line number | Diff line number | Diff line change |
|---|---|---|
| @@ -1,79 +1,103 @@ | ||
| # Contributing to Autoafids | ||
| # Contributing to AutoAFIDs | ||
|
|
||
| Autoafids python package uses Poetry pacakge manager to manage its dependencies. You’ll need it installed on your machine before contributing to the software. Installation instructions can be found on the | ||
| [Poetry website](https://python-poetry.org/docs/master/#installation). | ||
| AutoAFIDs is a Python-based BIDS App for automatic anatomical fiducial detection. Development is managed using **Conda** for dependency and environment management. | ||
|
|
||
| Autoafids primarily caters to T1w modality images, in which case it doesn't need to depend on pacakges outside of python but for other modalities it has a single dependency outside of python, i.e., `synthsr`. | ||
| > 🛠️ These instructions are intended for contributors making code changes to the AutoAFIDs codebase or using advanced features such as Snakemake cluster execution profiles. | ||
|
|
||
| Note: These instructions are only recommended if you are making changes to the Autoafids codebase and committing these back to the repository or if you are using Snakemake’s cluster execution profiles. | ||
| --- | ||
|
|
||
| ## Setup the development environment | ||
| ## 📦 Prerequisites | ||
|
|
||
| Once Poetry is available, clone this repository and install all dependencies (including `dev`): | ||
| Ensure **Conda** is installed on your system. You can install Miniconda or Anaconda by following the official guide: | ||
| 👉 [https://docs.conda.io/projects/conda/en/latest/user-guide/install/index.html](https://docs.conda.io/projects/conda/en/latest/user-guide/install/index.html) | ||
|
|
||
| ``` | ||
| Note: AutoAFIDs primarily supports T1-weighted images. For additional modalities (e.g., T2w), an external dependency (`SynthSR`) is required and must be installed separately. | ||
|
|
||
| --- | ||
|
|
||
| ## 🧪 Setting Up the Development Environment | ||
|
|
||
| 1. Clone the repository: | ||
|
|
||
| ```bash | ||
| git clone https://github.com/afids/autoafids.git | ||
| cd autoafids | ||
| poetry install --with dev | ||
| ``` | ||
|
|
||
| Poetry will automatically create a virtual environment. To customize where | ||
| these virtual environments are stored, see the poetry docs | ||
| [here](https://python-poetry.org/docs/configuration/). | ||
| 2. Create and activate the development environment: | ||
|
|
||
| Then, you can run autoafids: | ||
| ```bash | ||
| conda create -n autoafids-dev python=3.10 | ||
| conda activate autoafids-dev | ||
| ``` | ||
|
|
||
| 3. Install AutoAFIDs with development dependencies: | ||
|
|
||
| ```bash | ||
| mamba install -c conda-forge -c bioconda -c khanlab autoafids | ||
| pip install -r requirements-dev.txt | ||
| ``` | ||
|
|
||
| --- | ||
|
|
||
| ## ▶️ Running AutoAFIDs (Dev Mode) | ||
Dhananjhay marked this conversation as resolved.
Outdated
Show resolved
Hide resolved
|
||
|
|
||
| Once installed, you can run AutoAFIDs directly from the command line: | ||
|
|
||
| ```bash | ||
| autoafids -h | ||
| ``` | ||
|
|
||
| or you can activate a virtual environment shell and run autoafids directly: | ||
| To modify the CLI or core modules, edit files in the `autoafids/` directory and re-run commands in your active Conda environment. | ||
|
|
||
| ``` | ||
| poetry shell | ||
| autoafids | ||
| ``` | ||
| --- | ||
|
|
||
| You can exit the poetry shell with `exit` | ||
| ## 🧹 Code Quality and Formatting | ||
|
|
||
| ## Running and fixing code format quality | ||
| AutoAFIDs uses several tools to ensure clean and consistent code: | ||
|
|
||
| Autoafids uses [poethepoet](https://github.com/nat-n/poethepoet) as a task runner. | ||
| You can see what commands are available by running: | ||
| - [`ruff`](https://github.com/charliermarsh/ruff) – for linting and formatting | ||
| - [`snakefmt`](https://github.com/snakemake/snakefmt) – for formatting Snakemake files | ||
| - [`yamlfix`](https://github.com/lyz-code/yamlfix) – for YAML cleanup | ||
|
|
||
| ``` | ||
| poetry run poe | ||
| ### Check formatting: | ||
|
|
||
| ```bash | ||
| ruff check . | ||
| snakefmt --check Snakefile | ||
| yamlfix --check config/ | ||
| ``` | ||
|
|
||
| We use a few tools, including `ruff`, `snakefmt`, and `yamlfix` to ensure | ||
| formatting and style of our codebase is consistent. There are two task runners | ||
| you can use to check and fix your code, which can be invoked with: | ||
| ### Auto-fix formatting: | ||
|
|
||
| ```bash | ||
| ruff check . --fix | ||
| snakefmt Snakefile | ||
| yamlfix config/ | ||
| ``` | ||
| poetry run poe quality-check | ||
| poetry run poe quality | ||
|
|
||
| --- | ||
|
|
||
| ## 🧪 Dry Run / Workflow Testing | ||
|
|
||
| To test the Snakemake workflow without running any jobs, use the built-in **dry-run** feature: | ||
|
|
||
| ```bash | ||
| autoafids tests/data tests/output participant -n | ||
| ``` | ||
|
|
||
| _Note: If you are in a poetry shell, you do not need to prepend `poetry run` to | ||
| the command._ | ||
| The `tests/data` directory contains a lightweight fake BIDS dataset (with zero-byte files) useful for testing the pipeline logic. | ||
|
|
||
| ## Dry-run / testing your workflow | ||
| You can simulate more complex scenarios by modifying the `tests/` configuration and rerunning dry-runs with `--modality` and other options. | ||
|
|
||
| Using Snakemake\'s dry-run option (`--dry-run`/`-n`) is an easy way to verify | ||
| any changes made to the workflow are working direcctly. The `tests/data` folder | ||
| contains a _fake_ BIDS dataset (i.e. dataset with zero-sized files) that is | ||
| useful for verifying different aspects of the workflow. These dry-run tests are | ||
| part of the automated Github actions that are run for every commit. | ||
| --- | ||
|
|
||
| You can invoke the pre-configured task via | ||
| [poethepoet](https://github.com/nat-n/poethepoet) to perform a dry-run: | ||
| ## 🙋 Questions, Issues, and Feedback | ||
|
|
||
| ``` | ||
| poetry run poe test_base | ||
| ``` | ||
| We welcome all forms of feedback and contributions. | ||
|
|
||
| This performs a number of tests, involving different scenarios in which a user | ||
| may use autoafids. | ||
| - 💬 For questions or suggestions, use the [Discussions](https://github.com/afids/autoafids/discussions) page. | ||
| - 🐛 For bugs or feature requests, open an issue on the [Issues](https://github.com/afids/autoafids/issues) page. | ||
| - 📧 You may also contact [[email protected]](mailto:[email protected]) or reach out to **Alaa Taha**, the Science Lead. | ||
|
|
||
| ## Questions, Issues, Suggestions, and Other Feedback | ||
| Please reach out if you have any questions, suggestions, or other feedback related to this software—either through email ([email protected]) or the discussions page. Larger issues or feature requests can be posted and tracked via the issues page. Finally, you can also reach out to Alaa Taha, the Science Lead. | ||
| Thanks for helping improve AutoAFIDs! | ||
Oops, something went wrong.
Oops, something went wrong.
Add this suggestion to a batch that can be applied as a single commit.
This suggestion is invalid because no changes were made to the code.
Suggestions cannot be applied while the pull request is closed.
Suggestions cannot be applied while viewing a subset of changes.
Only one suggestion per line can be applied in a batch.
Add this suggestion to a batch that can be applied as a single commit.
Applying suggestions on deleted lines is not supported.
You must change the existing code in this line in order to create a valid suggestion.
Outdated suggestions cannot be applied.
This suggestion has been applied or marked resolved.
Suggestions cannot be applied from pending reviews.
Suggestions cannot be applied on multi-line comments.
Suggestions cannot be applied while the pull request is queued to merge.
Suggestion cannot be applied right now. Please check back later.
Uh oh!
There was an error while loading. Please reload this page.